Abstract
It is currently unclear whether changes in viral communities will ever be predictable. Here we investigate whether viral communities in wildlife are inherently structured (inferring predictability) by looking at whether communities are assembled through deterministic (often predictable) or stochastic (not predictable) processes. We sample macaque faeces across nine sites in Bangladesh and use consensus PCR and sequencing to discover 184 viruses from 14 viral families. We then use network modelling and statistical null-hypothesis testing to show the presence of non-random deterministic patterns at different scales, between sites and within individuals. We show that the effects of determinism are not absolute however, as stochastic patterns are also observed. In showing that determinism is an important process in viral community assembly we conclude that it should be possible to forecast changes to some portion of a viral community, however there will always be some portion for which prediction will be unlikely.
Similar content being viewed by others
Introduction
The recent Ebola virus outbreak in West Africa1,2 is a timely reminder that we have never successfully predicted the emergence of a new infectious disease in people3. Perhaps precluded from doing so (at least in part) by historical deficiencies in our knowledge of global viral diversity in wildlife4,5,6 or the pathways and mechanisms of spillover and spread7,8,9, the threat that infectious diseases now increasingly pose to public health4,5,6 and economic stability10,11,12 has excited efforts to establish a predictive understanding of emergence3,4,13. One area of ‘prediction’ that would be particularly useful is the ability to forecast how viral diversity might respond to environmental drivers of disease emergence, for example land-use change. This would allow us to test response options designed to mitigate, or adapt to, the impact of those changes and potentially reduce the risk of zoonotic emergence14. Being able to predict such changes however assumes that viral diversity is inherently predictable. It assumes that viral communities are built and controlled through deterministic and inherently ecological processes that can be identified and understood, and we do not yet know whether this is true. If viral diversity in wildlife is inherently random (stochastic), then predicting the outcome of an environmental perturbation would be impossible, as many have long believed15. But if it is not random, if it is deterministically structured (or at least structured to some degree) then predicting changes in viral diversity might indeed be possible.
Here, we apply established ecological theory on macrobial species distributions (for example, plants and animals)16,17 to viral assemblages of the rhesus macaque and look for evidence of deterministic and stochastic effects in the structure of these communities. We adopt the null hypothesis that virodiversity can be readily explained by random processes (chance colonisation or extinction and ecological drift) and look for departure from random via the presence of discernible pattern to identify and subsequently test for the presence of determinism18,19. Our data indicate that viral communities within the macaque are assembled though largely ecological (deterministic) processes and should therefore be inherently predictable. However, we also show that stochastic processes contribute to patterns of viral diversity, suggesting that changes to some portion of the community will never be predictable.
Throughout the paper we use the terms ‘determinism’ to refer to the identification of non-random patterns and ‘stochastic’ to refer to any ecological process that results in patterns of diversity, relative abundance and composition that are indistinguishable from random chance alone18. We clarify that it is not our intention at this time to determine the processes behind non-randomness, as these might involve a variety of either neutral processes assuming ecological equivalence17 or processes based on ecological niche differentiation16.
Results
Virodiversity of the rhesus macaque in Bangladesh
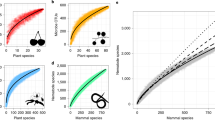
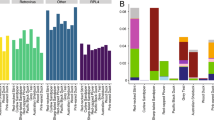
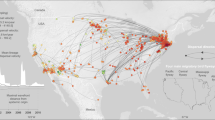
Using a combination of consensus polymerase chain reaction (cPCR) and high-throughput sequencing (HTS), we characterised the faecal virodiversity of 458 rhesus macaques sampled across nine urban sites in Bangladesh (Fig. 1) and identified 184 unique viruses from 14 families (Fig. 2a and Supplementary Table 1). We identified 37/184 viruses by cPCR and 147/184 by HTS — highlighting the usefulness of combining the high sensitivity of PCR with the broad reactivity of HTS. We make particular note of an unprecedented diversity of a small bipartite picobirnaviruses (PbVs), which accounted for 120 of the 184 viruses found in these animals (Fig. 2b). Importantly, we make no assertion that all 184 viruses are singularly associated with macaques, or that true infection has occurred. Indeed, several human viruses were detected during this discovery effort (Supplementary Table 1) suggesting a multihost ecology that would be readily explained by the long and close association between people and macaques at each of our sampling sites21,22. Instead, we use genetic detection to demonstrate inclusion in the viral community to which these macaques are exposed, even if the presence of a virus is the result of dispersal from another host species or contribution to the community is low because of rarity23. For purposes of definition, we consider a ‘unique virus’ to be a monophyletic cluster of sequences that is distinct from its nearest neighbour by non-overlapping genetic identities24.
Number of individuals sampled at each site is indicated. All sites are urban/peri-urban, with known contact between macaques and people, livestock and domestic animals (though frequency of contact is not assessed here). Number of samples collected is not consistent across sites, however sampling effort (number of collection days per site) is the same. Further description of each site is provided in Supplementary Table 3.
(a) Diversity of viruses discovered by family. (b) Maximum likelihood phylogeny of picobirnavirus diversity. Although viruses from many families were discovered, we have selected to show the PbV tree because of the substantial diversity observed. One section of the tree is shown as an insert for better visualization. (c) Viral discovery curve to assess saturation and estimate the total richnesss (number of viruses) that exists. Red line is the collector curve. Solid black line is the rarefaction curve. Dotted line indicates Chao2 estimation of asymptotic richness by sample number, and shading indicates 95% confidence intervals. (d) Rank abundance curve of the 184 viruses detected in this study.
Non-parametric viral discovery curves were used to assess the bounds of the viral community (total number of viruses) and assess the completeness of our discovery effort24,25,26. These curves indicated that the community contains a total of 283 viruses (Fig. 2c). We estimate therefore that the 184 viruses detected in our study represent ∼65% of the viruses that exist in these macaques. Plotting the rank abundance of the observed virodiversity showed that only a few of these viruses dominated the community, whereas most occurred only rarely (Fig. 2d). This uneven distribution is a pervasive pattern characterising macrobial communities27, and lends support to the notion of universal or unifying laws of assembly that apply as equally to microbes as they do to communities of plants and animals23. Assuming that the remaining (undiscovered) virodiversity was not detected because of rarity (that is, exists within the long tail of the rank abundance curve) and that rare viruses contribute little to the community27, we suggest that sufficient diversity has been detected with which to explore the structure of this naturally occurring community.
Evidence of determinism driving viral community structure
A two-mode affiliation network28 was used to illustrate the connectivity between viruses and their hosts. This revealed a dominant (though not exclusive) pattern of site-specific diversity consistent with determinism (Fig. 3a). To ensure that the patterns observed here were not simply the result of chance, null models19 were incorporated into an assessment of β-diversity29 (difference in viral composition between sites) and used to confirm that individual macaques mostly shared viruses with other individuals from the same site, and only rarely with those located elsewhere (Fig. 3c and Supplementary Fig. 1). By applying phylogenetic measures of β-diversity (Beta Nearest Taxon Index30,31) to a subset of the community (applied to the 120 PbVs) we were also able to infer that these non-random patterns may be emerging due to dispersal limitation (Supplementary Fig. 2).
Viruses shaded grey. Size of node indicates abundance. Coloured nodes indicate individual macaques, coloured by site. (a) The overall network shows a non-random pattern of site-associated diversity. (b) Insert showing an area of apparently stochastic distributions. (c) Lower corner (blue shading) shows a pairwise assessment of beta (β) diversity between sites. The Jaccard (incidence based) index was used and demonstrates that little similarity exists between sites. Additional metrics shown in Supplementary Fig. 1. Top corner (green shading) presents results of the null model, and shows that the observed distributions are different from chance (dark green=significant P value). (d) Distribution of pairwise genetic identities for PbVs found in the same host, and those found in different hosts. Results presented are for G1 PbVs, but results consistent for G2. P value (Wilcoxon rank-sum test) indicates that the ‘within same host’ values (max=85.5%) are different from chance.
To verify that dispersal (a largely stochastic process) was not responsible for the observed distributions, we correlated β-diversity (Jaccard index) with distance between sites to look further at the potential influence of dispersal limitation, and found no significant association (Mantel test: P=0.807; Principle Coordinates of Neighbour Matrices (PCNM)31,32,33: −0.352, P=0.134). We also tested for dispersal limitation by looking at whether PbV sequences from the same site were more related to each other than to viruses from other sites and whether this relatedness decreased with increasing site distance. When ‘same-site’ (distance=0 km) was included in the analysis, the association was shown to be significant for both genotype 1 (G1) and genotype 2 (G2) PbVs (Spearman’s rank correlation test; G1: ρ=−0.034; P<0.001; G2: ρ=−0.186; P<0.001). However when removed to test the strength of the effect, the significance of the correlation was lost (G1: ρ=−0.209; P=0.222; G2: ρ=−0.149; P=0.448). These results confirm that while there is substantial dispersal among macaques within a population, there is very limited dispersal among populations — regardless of geographic distance separating them. Evidence of multiple recombination events between viruses detected at different sites (Supplementary Table 2) and the natal migrations of male macaques seeking new groups22,34 both demonstrate connectivity between these populations, and suggest that these viruses are not (completely) limited in their ability to disperse. However, the frequency at which viruses become established in new populations via dispersal is seemingly low. Although we interpret these patterns of β-diversity as the result of deterministic processes based on our definition (that is, non-random), we also acknowledge that very low, or very high, rates of dispersal can lead to non-random patterns.
Determinism was also observed on more local scales, within sites and individuals. Using the PbV data (again, because of its presence at all sites) we looked to see whether there was a limit to how genetically similar two co-occurring viruses could be. The maximum observed identity between any two PbV sequences found in the same individual was 85.8% for G1 viruses, and 88.7% for G2 viruses (Fig. 3d). In contrast, the maximum identity for any two non-identical sequences found in different individuals at the same site was 99.8% (for both G1 and G2). This pattern was shown to be significantly different from chance (Wilcoxon rank-sum test; P<0.001) based on 1,000 random selections of γ-diversity (restricting α-diversity to the richness observed), and was consistent when stratified by site. It strongly suggests deterministic mechanisms do exist to limit the co-occurrence of closely related viruses in the same animal, and while the specific mechanisms are unknown we postulate they could well include virus:virus interactions such as competitive exclusion (analogous to the theory of limiting similarity16) or virus:host interactions like immune recognition. We qualify that this conclusion is dependent on the assumption that a correlation exists between phylogenetic relatedness and ecological similarity30 (for a competitive process) or host response (for immune recognition), and while we see no reason to doubt the validity of this assumption we acknowledge that little is currently known about picobirnavirus ecology and host interactions. We therefore suggest that additional data exploring whether ecological similarity increases with genetic similarity will now be required to confirm this relationship.
The potential for virus:virus interactions was investigated further using a one-mode network that showed the connectivity of viruses based on the relative frequency of host sharing (Fig. 4). Several biological associations were apparent, including that of adenovirus MmAdV-5 with dependoviruses MmAaV-1, 2 and 3. Named dependoviruses (or adeno-associated viruses) because of their requirement for a ‘helper virus’, these small DNA viruses are well known to use adenovirus to satisfy their replicative deficiencies35. The strength of this association was tested using PAIRS24,36, and the frequency of their co-occurrence shown to be significantly greater than expected by chance (C-score; P<0.001). The network also identified significant co-occurrence between MmAaV-1 and the herpesvirus MmHV-1 (P=0.002). Herpesviruses are also known to satisfy the helper requirements of dependoviruses35. Together these results demonstrate (and to some degree, validate) the usefulness of networks in understanding biological relationships in viral communities. In total, 35/184 viruses showed statistically supported (P=<0.05) positive co-occurrence with another virus, while 12/184 had negative associations. These results demonstrate that deterministic mechanisms exist to both promote and prevent the co-occurrence of viruses in the community.
Stochastic distributions
Not all distributions could be attributed to determinism. MmAdV-5 was shown to significantly co-occur with various dependoviruses, but no discernible pattern could be identified and tested to explain its own distribution (that is, the presence of MmAdV-5 would explain the presence of MmAaV-1, 2 and 3, but perhaps not vice versa). The same is true for simian foamy virus (MmSFV) and the two HVs (MmHV-1 and 2), which like MmAdV-5 were detected at multiple sites without any apparently deterministic signature (Fig. 3b). We therefore attribute the distribution of these viruses to stochastic processes but acknowledge that scale or incomplete sampling might be obscuring determinism18.
Discussion
Our results suggest that viral communities in the rhesus macaque are heavily influenced by deterministic factors, and therefore likely to be inherently structured. The effects of determinism were not absolute however, as stochastic processes also appeared to contribute to virodiversity. As such, we conclude that it should be possible to forecast changes to a significant portion of the viral community in a given location, but suggest there will also be some portion for which prediction will always be unlikely. We qualify that our study only demonstrates that changes in viral diversity should eventually be predictable, based on the assumption that non-random patterns in biological systems infer inherent predictability16,18,19,23,37, and based on the assumption that our data is representative of the entire community (that is, including those viruses that were not discovered, and assumed to be rare). It does not, and is not intended to, present a framework for how this prediction might be achieved. Instead, this study contributes to the theory that will support the future development of these probabilistic models, describing how the distribution of viruses is likely to change in response to different environmental or host factors. To achieve this, we advocate investigating the specific mechanisms associated with determinism (for example, the host:host; host:virus or virus:virus interactions) as well as the continued and systematic description of wildlife virodiversity through time and space. We also acknowledge that this finding will not lead directly to predictions of disease emergence, however we suggest it does provide the basis on which to test the hypothesis that drivers such as land-use change or climate (among others) promote disease emergence through their effect on the structure of the zoonotic pool.
Methods
Sample collection
Faecal samples (n=458) were collected non-invasively from free-ranging rhesus macaques (Macaca mulatta) over a 2-month period (February/March-2013) under ethical approval from the International Centre for Diarrhoeal Disease Research, Bangladesh (icddr, b; protocol: 2008-074) and UC Davis (protocol: 16048). Location of each site and a description of the macaque populations are provided in Supplementary Table 3. Sampling effort was consistent for all sites (3 days per site). All samples were collected immediately after defecation and stored in liquid nitrogen within 10 min of collection, until transfer to −80 °C for storage.
Sample processing and viral discovery
Samples were viral particle enriched through (i) filtration to remove cellular debris and bacteria and (ii) nuclease treatment to remove unencapsulated RNA/DNA. For this, samples were thawed on ice and 500 μl of viral transport medium (Viral Transport Medium (VTM); BD Universal Viral Transport System) added, vortexed to homogenise, and centrifuged for 5 min at 8,000g. Supernatant was transferred to an Ultrafree-MC HV Centrifugal Filter 0.45 μM (Milipore Cat. No. UFC30HVNB) and centrifuged for 3 min at 12,000g. The flow (∼130–150 μl) was collected and 1 μl RNase A (Ribonuclease protection assay Grade, 1 mg ml−1, Life Technologies Cat. No. AM2272) added and incubated at room temperature for 15 min. If the flow volume was close to 200 μl then 2 μl of RNase A was used. Following RNase treatment, 1.5 μl of MgCl2 (1M), 4 μl of Turbo DNase (2 U ml−1, Ambion Cat. No. AM2238) and 1 μl of Benzonase. (Novagen, Cat. No. 70664-3), were added, mixed gently and incubated at room temperature for 45 min. Roche MagNa Pure lysis solution was added immediately to inactivate nucleases and lyse viral particles, and total nucleic acids extracted using the Roche MagNA Pure 96 platform according to the manufacturer’s instructions.
Samples were processed for viral detection and discovery using both consensus PCR and next-generation sequencing. cPCR, allows the ‘universal’ amplification of sequences from viruses within a given family or genus, and the subsequent discernment of viral strains within. Total nucleic acids was reverse transcribed into cDNA using SuperScript III (Invitrogen) according to the manufacturer’s instructions, and a total of 41 assays representing 27 viral families or genera used for the detection of viral sequences. Two synthetic plasmids were constructed for use as ‘universal controls’ to confirm successful execution of each assay and check for contamination (Supplementary Fig. 3). Detailed protocols for all cPCR assays used are provided in the Supplementary Methods and Supplementary Table 4. Bands of the expected size were excised from 1% agarose, cloned into Strataclone PCR cloning vector, and 24 white colonies sequenced to confirm detection and look for co-occurring viruses.
To guard against the potential of cPCR to miss viruses that are divergent or not among the targeted viral families, HTS was also applied to all samples. Although generally less sensitive than PCR, HTS allows for the capture of a very broad diversity of viruses because it amplifies all viral nucleic acids present. Samples were processed in pools of eight, and libraries prepared for both the Ion Torrent (PGM; 1 million reads per pool) and Illumina (High-Seq; 10 million reads per pool) platforms, according to each of the manufacturer’s instructions. Sequence reads were aligned against host reference databases to remove host background using bowtie2 mapper, and host-subtracted reads primer trimmed and filtered based on quality, GC content and sequence complexity. The remaining reads were de novo assembled using Newbler (v2.6) for PGM data and MIRA (v4.0) for Illumina. Contigs and unique singletons were subjected to homology search using MegaBlast against the GenBank nucleotide database. Sequences that showed poor or no homology at the nucleotide level were blasted using BLASTx against the viral GenBank protein database. Viral sequences from the BLASTx analysis were subjected to a homology search against the GenBank protein database to correct for biased e-values. Sequences of plant viruses or insect viruses from viral families that have (to date) never been associated with infection of any vertebrate species were not considered in this study. All other viral sequences identified by HTS were subsequently confirmed by PCR and all samples re-screened individually to assess sequence distribution in all macaques (as we assume the lower sensitivity of HTS may produce false negatives). Where substantial diversity was observed (for example, PbVs), new cPCR assays were designed based on the HTS data, and used to re-screen all pooled samples individually to detect the full diversity present.
Network, phylogenetic and statistical analyses
A presence/absence matrix was constructed to show the distribution of viruses across all samples, and used for network and statistical analyses as described in the main text. In summary, A bipartite (two-mode) affiliation network was generated for virus–macaque host matrix data, stratified by site name, and a unipartite (one-mode) virus:virus network was generated to display the connections between viruses. Network analyses and visualisation were conducted in the network analysis platform Gephi, using the force-directed algorithm ForceAtlas2 (ref. 28). Significance of pairwise associations was determined using PARS24,36. All other statistical analyses were performed using MATLAB (Mathworks, Natick USA) version R2013a. Discovery curves were generated in R package iNEXT. Phylogenetic analyses of sequence data were performed using MUSCLE38 for initial alignments, followed by manual refinement in Se-Al v2.0a11 (ref. 39). Maximum Likelihood trees were reconstructed using PAUP* (ref. 40) and best fitting models selected using jModeltest v2.1.5 (ref. 41). Trees were annotated using iTOL (v2.1)42. Measures of phylogenetic β-diversity were performed by first calculating the genetic similarity between every two different PbV sequences. This similarity matrix was then transformed into a distance matrix using the method proposed by Dray et al.31, which was then used to calculate the βMNTD (Beta Mean Nearest Taxon Distance)30. Results were compared with a null distribution of βMNTD, where PbV taxa were randomised across sites with a fixed relative abundance and recalculated 999 times. Beta Nearest Taxon Index values were then calculated as the number of s.d. that the observed βMNTD is from the mean of the null distribution.
Additional information
Accession codes: The sequence data have been deposited in the GenBank nucleotide database under accession codes KT599642 to KT599859, KT334810 to KT335259, and KT599483 to KT599639.
How to cite this article: Anthony, S. J. et al. Non-random patterns in viral diversity. Nat. Commun. 6:8147 doi: 10.1038/ncomms9147 (2015).
References
Baize, S. et al. Emergence of Zaire Ebola virus disease in Guinea. N. Engl. J. Med. 371, 1418–1425 (2014).
WHO Ebola Response Team. Ebola virus disease in West Africa – the first 9 months of the epidemic and forward projections. N. Engl. J. Med. 371, 1481–1495 (2014).
Morse, S. S. et al. Prediction and prevention of the next pandemic zoonosis. Lancet 380, 1956–1965 (2012).
Jones, K. E. et al. Global trends in emerging infectious diseases. Nature 451, 990–993 (2008).
Karesh, W. B. et al. Ecology of zoonoses: natural and unnatural histories. Lancet 380, 1936–1945 (2012).
Woolhouse, M. E. & Gowtage-Sequeria, S. Host range and emerging and reemerging pathogens. Emerg. Infect. Dis. 11, 1842–1847 (2005).
Keesing, F. et al. Impacts of biodiversity on the emergence and transmission of infectious diseases. Nature 468, 647–652 (2010).
Smolinski, M. S., Hamburg, M. A. & Lederberg, J. in Microbial Threats to Health: Emergence, Detection and Response Institute of Medicine; National Academies Press (2003).
Weiss, R. A. & McMichael, A. J. Social and environmental risk factors in the emergence of infectious diseases. Nat. Med. 10, S70–S76 (2004).
The World Bank. People, Pathogens and Our Planet: The Economics of One Health. vol. 2. Report #69145-GLB (2012).
The World Bank. The Economic Impact of the 2014 Ebola Epidemic: Short and Medium Term Estimates For West Africa The World Bank (2014).
Pike, J., Bogich, T., Elwood, S., Finnoff, D. C. & Daszak, P. Economic optimization of a global strategy to address the pandemic threat. Proc. Natl Acad. Sci. USA 111, 18519–18523 (2014).
Han, B. A., Schmidt, J. P., Bowden, S. E. & Drake, J. M. Rodent reservoirs of future zoonotic diseases. Proc. Natl Acad. Sci. USA 112, 7039–7044 (2015).
Evans, M. R. et al. Predictive systems ecology. Proc. Biol. Sci. 280, http://dx.doi.org/10.1098/rspb.2013.1452 (2013).
Murphy, F. A. Emerging zoonoses. Emerg. Infect. Dis. 4, 429–435 (1998).
Chase, J. M. & Leibold, M. A. Ecological Niches: Linking Classical and Contemporary Approaches University of Chicago Press (2003).
Hubbell, S. P. The Unified Neutral Theory of Biodiversity and Biogeography Princeton University Press (2001).
Chase, J. M. & Myers, J. A. Disentangling the importance of ecological niches from stochastic processes across scales. Philos. Trans. R. Soc. Lond. B Biol. Sci. 366, 2351–2363 (2011).
Chase, J. M., Kraft, N. J. B., Smith, K. G., Vellend, M. & Inouye, B. D. Using null models to disentangle variation in community dissimilarity from variation in alpha-diversity. Ecosphere 2, doi 10.1890/Es10-00117.1 (2011).
Li, C. X. et al. Unprecedented genomic diversity of RNA viruses in arthropods reveals the ancestry of negative-sense RNA viruses. eLife 4, doi: 10.7554/eLife.05378 (2015).
Lloyd-Smith, J. O. et al. Epidemic dynamics at the human-animal interface. Science 326, 1362–1367 (2009).
Hasan, M. K. et al. Distribution of Rhesus Macaques (Macaca mulatta) in Bangladesh: inter-population variation in group size and composition. Primate Conserv. 26, 125–132 (2013).
Fuhrman, J. A. Microbial community structure and its functional implications. Nature 459, 193–199 (2009).
Anthony, S. J. et al. A Strategy to estimate unknown viral diversity in mammals. MBio 4, e00598–13 (2013).
Chao, A. in Encyclopedia of Statistical Sciences Vol. 12, eds Balakrishnan N., Read C. B., Vidakovic B. Wiley (2005).
Chao, A., Colwell, R. K., Lin, C. W. & Gotelli, N. J. Sufficient sampling for asymptotic minimum species richness estimators. Ecology 90, 1125–1133 (2009).
Magurran, A. E. & Henderson, P. A. in Biological Diversity: Frontiers in Measurement and Assessment eds Magurran A. E., McGill B. J. Oxford University Press (2011).
Jacomy, M., Venturini, T., Heymann, S. & Bastian, M. ForceAtlas2, a continuous graph layout algorithm for handy network visualization designed for the Gephi software. Plos One 9, e98679 (2014).
Jost, L., Chao, A. & Chazdon, R. L. in Biological Diversity: Frontiers in Measurement and Assessment eds Magurran A. E., McGill B. J. Oxford University Press (2011).
Stegen, J. C., Lin, X. J., Konopka, A. E. & Fredrickson, J. K. Stochastic and deterministic assembly processes in subsurface microbial communities. ISME J. 6, 1653–1664 (2012).
Dray, S., Legendre, P. & Peres-Neto, P. R. Spatial modelling: a comprehensive framework for principal coordinate analysis of neighbour matrices (PCNM). Ecol. Model. 196, 483–493 (2006).
Borcard, D. & Legendre, P. All-scale spatial analysis of ecological data by means of principal coordinates of neighbour matrices. Ecol. Model. 153, 51–68 (2002).
Legendre, P., Lapointe, F. J. & Casgrain, P. Modeling brain evolution from behavior – a permutational regression approach. Evolution 48, 1487–1499 (1994).
Drickamer, L. C. & Vessey, S. H. Group changing in free-ranging male rhesus monkeys. Primates 14, 359–368 (1973).
Geoffroy, M. C. & Salvetti, A. Helper functions required for wild type and recombinant adeno-associated virus growth. Curr. Gene Ther. 5, 265–271 (2005).
Pairs – a FORTRAN program for studying pair-wise species associations in ecological matrices http://www.keib.umk.pl/pairs . 1.0 (Department of Animal Ecology, Toruń, Poland, (2008).
Fuhrman, J. A. & Steele, J. A. Community structure of marine bacterioplankton: patterns, networks and relationship to function. Aquat. Microb. Ecol. 53, 69–81 (2008).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Se-Al: Sequence Alignment Editor, version 2.0a11 http://tree.bio.ed.ac.uk/software/seal/ (2002).
PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods), version 4.0b10.Sinauer Associates., Sunderland, MA (2003).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: more models, new heuristics and parallel computing. Nat. Methods 9, 772 (2012).
Letunic, I. & Bork, P. Interactive Tree Of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res. 39, W475–W478 (2011).
Acknowledgements
This study was made possible by the support of the American people through the United States Agency for International Development (USAID) Emerging Pandemic Threats PREDICT project (cooperative agreement number GHN-A-OO-09-00010-00). We thank the Bangladesh Forest Department and the Ministry of Environment and Forest for permission to conduct this study. We are thankful to icddr, b and its core donor the Governments of Australia, Bangladesh, Canada, Sweden and the UK for providing core/unrestricted support to icddr,b. We thank Jens H Kuhn, Bohyun Lee, Angelica de Almeida Campos, Kawthar Muhammad, Oscar Rico, Amy Wray, Nicole Arrigo, Emily. S. Gurley, Ausraful Islam, Tapan Kumar Dey, Shafiqul Islam, Abdul Hai, Pitu Biswas and Gafur Sheikh for their contributions to this study.
Author information
Authors and Affiliations
Contributions
S.J.A. conceived the study. A.I. and N.H. collected the samples, with contributions from M.K.R., and J.H.E. S.J.A., I.N.M., E.L., A.P., J.G. and T.G. developed laboratory methods and conducted experiments for viral discovery and characterisation. S.J.A., C.J., K.J., P.L.H., X.C., A.S., A.L.H., R.O.F., W.U., C.Z.T., T.G. and W.I.L. performed data analysis and provided interpretations. S.J.A., N.W., S.S.M., M.R., J.K.M., P.D. and W.I.L. wrote the paper. The contents are the responsibility of the authors and do not necessarily reflect the views of USAID or the United States Government.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-3, Supplementary Tables 1-4, Supplementary Methods and Supplementary References (PDF 1206 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Anthony, S., Islam, A., Johnson, C. et al. Non-random patterns in viral diversity. Nat Commun 6, 8147 (2015). https://doi.org/10.1038/ncomms9147
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms9147
This article is cited by
-
Human Exposure to Bats, Rodents and Monkeys in Bangladesh
EcoHealth (2023)
-
Coronavirus and Paramyxovirus Shedding by Bats in a Cave and Buildings in Ethiopia
EcoHealth (2022)
-
Classification of new morbillivirus and jeilongvirus sequences from bats sampled in Brazil and Malaysia
Archives of Virology (2022)
-
Potential zoonotic pathogens hosted by endangered bonobos
Scientific Reports (2021)
-
Ecological and Conservation Significance of Herpesvirus Infection in Neotropical Bats
EcoHealth (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.