Abstract
Ankylosing spondylitis (AS) is a common, highly heritable, inflammatory arthritis for which HLA-B*27 is the major genetic risk factor, although its role in the aetiology of AS remains elusive. To better understand the genetic basis of the MHC susceptibility loci, we genotyped 7,264 MHC SNPs in 22,647 AS cases and controls of European descent. We impute SNPs, classical HLA alleles and amino-acid residues within HLA proteins, and tested these for association to AS status. Here we show that in addition to effects due to HLA-B*27 alleles, several other HLA-B alleles also affect susceptibility. After controlling for the associated haplotypes in HLA-B, we observe independent associations with variants in the HLA-A, HLA-DPB1 and HLA-DRB1 loci. We also demonstrate that the ERAP1 SNP rs30187 association is not restricted only to carriers of HLA-B*27 but also found in HLA-B*40:01 carriers independently of HLA-B*27 genotype.
Similar content being viewed by others
Introduction
Ankylosing spondylitis (AS) is a common, highly heritable1, inflammatory arthritis for which HLA-B*27 is the major genetic risk factor. To better understand the genetic basis of the major histocompatibility complex (MHC) susceptibility loci, we genotyped 7,264 MHC single-nucleotide polymorphisms (SNPs) in 9,069 AS cases and 13,578 population controls of European descent using the Illumina Immunochip microarray. In addition to extremely strong effects due to HLA-B*27:02 and B*27:05, several other HLA-B alleles (B*07:02, B*13:02, B*40:01, B*40:02, B*47:01, B*51:01 and B*57:01) also affect susceptibility to AS. HLA-B-independent associations were demonstrated with variants in the HLA-A, HLA-DPB1 and HLA-DRB1 loci. We also demonstrate that the ERAP1 SNP rs30187 association is not restricted only to carriers of HLA-B*27 but also found in HLA-B*40:01 carriers independently of the HLA-B*27 genotype. The presence of associations in both HLA class I and II loci might reflect effects on antigen presentation to both CD4+ and CD8+ T lymphocytes in the pathogenesis of AS.
While the classical HLA-B*27 allele is found in over 85% of AS patients2,3,4, it is clearly not sufficient alone to cause disease, with only 1–5% of HLA-B*27 carriers developing the disease. From epidemiological data, it is evident that susceptibility to AS is affected by other genes within and outside the MHC1. Indeed, 26 risk loci outside the MHC have now been identified by genome-wide association studies5,6,7,8.
The biological mechanism(s) by which HLA-B27 confers risk of disease remains elusive. The main hypotheses regarding this mechanism can be divided into canonical mechanisms based on the known function of HLA-B27 within the adaptive immune system, and non-canonical mechanisms related to unusual properties of HLA-B27, notably its propensity to dimerise or misfold. Suggested canonical mechanisms propose either that HLA-B27 is uniquely capable of presenting particular peptide(s) found at sites of inflammation in AS to cytotoxic T lymphocytes (the arthritogenic peptide hypothesis)9 or that HLA-B27 is associated with reduced gut mucosal immunity, leading to migration of enteric bacteria across the intestinal mucosa, driving the production of the pro-inflammatory cytokine interleukin (IL)-23 and development of AS (the mucosal immunodeficiency hypothesis)10,11. Both these theories place antigenic peptide presentation and handling as critical steps in the pathogenesis of AS. One of the first non-MHC susceptibility loci to be identified in AS was endoplasmic reticulum aminopeptidase 1 (ERAP1)5, the main function of which is to trim peptides in the endoplasmic reticulum (ER) to optimal length for binding to MHC class I molecules on antigen-presenting cells for subsequent interaction with CD8+ T cells12,13. Moreover, this association is so far uniquely found in HLA-B*27-positive disease7.
HLA-B27 has an unusual property of forming homodimers through disulphide bonding of the unpaired cysteine residue at position 67 (ref. 14). It has been proposed that these homodimers may cause AS through abnormal presentation of peptides or by facilitating ‘abnormal’ interaction with natural killer cells15. Apart from HLA-B*27, the subtypes of the alleles HLA-B*14, HLA-B*15, HLA-B*38, HLA-B*39 and HLA-B*75 encode a cysteine residue at position 67 but of these there is only evidence that HLA-B*14 may be AS associated16,17. It is also unclear if these other non-HLA-B27 Cys67 variants can form homodimers. In addition, Cys67 is found on all HLA-B27 subtypes, including the subtypes HLA-B*27:06 and HLA-B*27:09, which are not AS associated18,19. A further hypothesis suggests that abnormal folding of the HLA-B27 molecule during assembly results in ER stress and activation of the unfolded protein response20,21. ER stress is evident in the HLA-B*27-transgenic rat model of AS and correlates with production of IL-23 (ref. 21), but has not been demonstrated in HLA-B*27-positive patients22,23,24.
While non-B27 HLA associations have been reported, notably with HLA-B40 (refs 25, 26, 27) and HLA-A*02 (ref. 8), most have not been definitive or replicated in independent studies. In this study, we analyse the associations of AS across the MHC aiming to identify functional and potentially causal variants using a large, previously reported, panel of cases and controls of European ancestry8. Here we extend on our primary analysis of this cohort by fine mapping the MHC region with imputation of SNPs, MHC class I and II classical alleles, and amino-acid residues within the classical HLA proteins28. In addition to HLA-B27, we identify further HLA-B and other HLA class I and II alleles associated with AS, and demonstrate that HLA-B40 in addition to HLA-B27 interacts with ERAP1 to cause disease. This implicates both CD4 and CD8 lymphocytes in AS pathogenesis and suggests that HLA-B40 and HLA-B27 operate by similar mechanisms to induce the disease.
Results
HLA-B susceptibility alleles
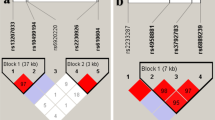
At the HLA-B locus, 38 classical alleles at four-digit resolution were imputed. All SNP, HLA and amino-acid association P-values were determined by logistic regression. As expected, the two common HLA-B*27 alleles in the European population, B*27:02 (odds ratio (OR)=43; P=1.07 × 10−122) and B*27:05 (OR=62; P<10−321), were the most significantly associated with disease risk (Fig. 1a–b; Tables 1 and 2). Controlling for the effect of the two B*27 alleles, we identified the protective alleles HLA-B*07:02 (OR=0.82; P=5.04 × 10−6) and HLA-B*57:01 (OR=0.75; P=5.13 × 10−4; Table 2). Moderate association was also observed, sequentially, with the risk alleles HLA-B*51:01 (OR=1.33; P=2.14 × 10−3), HLA-B*47:01 (OR=2.35; P=2.25 × 10−3), HLA-B*40:02 (OR=1.59; P=4.65 × 10−3), HLA-B*13:02 (OR=1.43; P=4.29 × 10−3) and HLA-B*40:01 (OR=1.22; P=4.93 × 10−3). No evidence of further susceptibility alleles was observed after controlling for the risk and protective alleles identified above (P>0.05; Fig. 1d). The HLA-B associations were similar in both HLA-B*27-positive and HLA-B*27-negative restricted analyses (Supplementary Tables 1–6).
Omnibus SNP and amino-acid association tests are shown in a, c, e, g and i, and classical allele association tests with two- and four-digit resolution in b, d, f, h and j. The strongest association was found with positions in the polymorphic nucleotide rs41558317 and in the polymorphic amino acid 97 of HLA-B (a), and with the HLA-B*27 allele (b). Controlling for the effect of HLA-B susceptibility alleles, an independent association was observed with SNPs and amino-acid position in the HLA-A locus (c) corresponding to the HLA-A*02:01 allele (d). Further conditioning on HLA-A and HLA-B loci an independent association with SNPs and amino-acid positions in the HLA-DPB1 locus was evident (e); no HLA-DPB1 classical allele was significant at the same magnitude as the SNPs and amino-acid positions (f). Further controlling for the effect of variation in the HLA-DPB1 locus association was observed with SNPs in the HLA-DRB1 locus (g, h). SNP association tests are shown in blue circles, colour coded by linkage disequilibrium from the SNP with the strongest association. Amino-acid position tests are shown as red triangles. Classical allele tests are shown as bars for two- and four-digit imputation resolution.
Non-HLA-B susceptibility loci in the MHC
To assess whether other MHC loci affect disease susceptibility independently from the HLA-B locus, we performed additional conditional analyses. Adjusting for the HLA-B susceptibility alleles identified, we observed an association signal with SNPs in the HLA-A locus (rs2975033; OR=1.22; P=6.16 × 10−10) and with the classical allele HLA-A*02:01 (OR=1.22; P=1.41 × 10−9; Fig. 1c–d). The risk allele ‘A’ of rs2975033 was in near perfect linkage disequilibrium with the risk allele HLA-A*02:01 (r2=0.97).
Further controlling for the effect of the susceptibility SNP rs2975033 in HLA-A revealed an independent signal with SNPs (rs1126513; OR=1.21; P=2.46 × 10−7) in the class II locus HLA-DPB1 (Fig. 1e); no association of similar strength to those seen with SNPs (P>10−5) were observed with classical HLA-DPB1 alleles (Fig. 1f). After controlling for the effect of the SNP rs1126513 in HLA-DPB1, we observed an association with the SNP rs17885388 (OR=1.16; P=1.27 × 10−5) in the HLA-DRB1 locus, and a similar level of significance was also observed with the class II allele HLA-DRB1*01:03 (P=3.78 × 10−5). No further associations were observed after controlling for all identified effects (P>5 × 10−5; Fig. 1i–j).
Association signals and amino-acid positions in HLA proteins
We observed disease-associated alleles at MHC class I and II loci. Classical alleles at these loci determine the amino-acid sequence of the respective HLA proteins, which could in turn influence the specificity of the peptides presented to CD8+ and CD4+ T lymphocytes. We, therefore, analysed the polymorphic amino-acid residues at these proteins to assess their effect in disease susceptibility. In this analysis, the strongest association was observed for amino-acid position 97 in HLA-B (omnibus P<10−3221; Table 1; Fig. 1a–b). In addition, through conditional analysis, we found that the association at the HLA-B*27 allele, and other HLA-B*27-associated polymorphisms, was explained by position 97 while the reverse was not true (Supplementary Table 7). This polymorphic position carries as many as six different amino-acid residues in the population (Fig. 2), each conferring a different degree of risk (or protection) to disease, consistent with the analysis of HLA-B alleles mentioned above (Table 2). Position 97 lies in the floor of the HLA-B peptide-binding groove (Fig. 3), located in the C/F pocket, also referred as the C-terminal pocket, which anchors the side chain of the C-terminal peptide residue29. Asparagine at position 97 is uniquely observed in HLA-B*27 alleles. Threonine at position 97 (predominantly found in HLA-B*51 alleles) was also found to increase disease risk (OR=1.12; P=4.50 × 10−3); serine (found in HLA-B*07 and *08 alleles) decreased risk of disease (OR=0.86; P=5.2 × 10−8); and valine (found in HLA-B*57 alleles) was also protective (OR=0.68; P=1.4 × 10−8; Table 3).
Strong associations were also observed with the amino-acid positions 70, 114, 77 and 67 of HLA-B but these signals were strongly attenuated after conditioning on amino-acid position 97. In contrast, none of these positions could explain the association at position 97. In particular, there was little evidence of association at position 67 (that is, the position where disulphide bonding of unlinked cysteine residues might occur) after conditioning on position 97 (P-value=0.04; Supplementary Table 8).
The most strongly associated amino acid of the HLA-A molecule, after conditioning on associated HLA-B alleles, was amino acid valine at position 95 (P=3.70 × 10−9). The association with this amino acid was statistically equivalent with that observed with the SNP rs2975033 and with the classical allele HLA-A*02:01. This amino acid is positioned within the binding site of HLA-A (Fig. 3).
Independent associations were observed at the two class II loci HLA-DPB1 and HLA-DRB1, and these were highly correlated with polymorphic amino acids in the peptide-binding site of these molecules (Fig. 3). At the HLA-DPB1 locus, rs1126513 showed the strongest association and the risk allele for rs1126513 was perfectly correlated with the presence of leucine at position 11 of the HLA-DPB1 molecule (position 11; OR=1.21; P-value=2.46 × 10−7). At the HLA-DRB1 locus, the strongest association with an amino acid was observed with aspartic acid at position 70 (OR=1.16; P-value=3.44 × 10−5); due to linkage disequilibrium this association was statistical equivalent to the one observed with the SNP rs17885388.
Gene–gene interactions and susceptibility loci
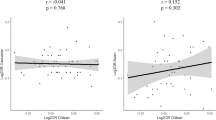
We have previously observed that the association with the variant rs30187 in the ERAP1 locus is restricted to HLA-B*27-positive subjects, consistent with an epistatic interaction between these two loci7. Here we investigated the possibility of interaction between the other HLA-B susceptibility alleles and the variant rs30187. When testing for interaction with the HLA-B*40 alleles, we found that rs30187-T increased the risk of disease in the strata where HLA-B*27 was present, as previously shown, or when HLA-B*40:01 was present in the absence of HLA-B*27 (OR=1.41; P=5.81 × 10−3); rs30187 had no effect on disease susceptibility when both HLA-B*27 and HLA-B*40 alleles were absent or in the non-HLA-B*27/HLA-B*40:02 stratum (Fig. 4). No evidence of interaction was observed between rs30187 and the other HLA-B susceptibility alleles. This suggests that the rs30187 variant interacts with the HLA-B*40:01 allele; although no evidence to support an interaction was observed with HLA-B*40:02, the study had low power to detect such an effect. There was no evidence of interaction between either of the HLA-B*40 alleles and any of the other independently associated susceptibility SNPs in the loci encoding the aminopeptidases ERAP1, ERAP2 and NPEPPS (P>0.1).
We then examined whether our data supported a model where the HLA-B*27 and HLA-B*40 alleles increased disease susceptibility beyond their inferred independent effects, as previously reported30. No support for an interaction between these alleles was observed in this data set (Supplementary Table 1).
Discussion
Independently of the expected HLA-B associations, this study demonstrates that both HLA-B*40:01 and -B*40:02 are disease associated alleles, and identified three further HLA-B risk alleles, HLA-B*51:01, B*47:01 and B*13:02. The allele HLA-B*51:01 is also the major genetic risk factor for Behçet’s disease31, a seronegative disease complicated by sacroiliitis resembling AS in up to 10% of cases32. In addition to the seven HLA-B risk alleles, we identified two protective alleles at this locus, HLA-B*07:02 and HLA-B*57:01. Interestingly, in the HLA-B*27-transgenic rat model of AS, the HLA-B*27-negative control carries the HLA-B*07 allele, and does not develop disease, consistent with the protective effect of this allele in humans33. It has recently been shown in HLA-B7/B27 co-expressing mice that there is partial negative selection of HLA-B27+ T cells in the course of defining the immunodominant response to influenza infection34. Further, in Erap-deficient, influenza-infected HLA-B27-positive mice, there was a marked reduction in presentation of the HLA-B27 immunodominant epitope, and T-cell immunity to that epitope, presumed to be because the HLA-B27-related immunodominant flu epitope requires cleavage by Erap to be presented by HLA-B27. In contrast, in HLA-B7-transgenic mice, Erap deficiency had no effect on presentation of the HLA-B7 immunodominant epitope or the corresponding T-cell response to it, suggesting that it does not require Erap cleavage for presentation35. This provides a potential mechanism to explain the genetic effects observed in humans with AS, with ERAP1 loss of function protecting against HLA-B27-associated AS, but having no effect in HLA-B7 carriers, where an HLA-B7 protective association is observed.
Outside the HLA-B locus, we identified three independent significant signals associated with AS; one was in the HLA class I locus HLA-A, and one each in the HLA class II loci HLA-DPB1 and HLA-DRB1. The association in the HLA-A locus corresponded to the classical allele HLA-A*02:01, which has also been implicated in multiple sclerosis36; however, while this allele is protective in multiple sclerosis, it increases the risk of AS. Previous studies have hinted at HLA-DPB1 associations with AS, which we have confirmed here. HLA-DPB1, in conjunction with HLA-DPA1, forms the HLA-DP heterodimer, which typically plays a role in the presentation of exogenously derived peptides, such as microbial peptides, to CD4+ T lymphocytes. The strongest association was found with an amino-acid position located in the base of the peptide-binding groove of HLA-DP, suggesting that this polymorphism might impact on the peptide repertoire presented by HLA-DP.
Previous findings that ERAP1 variants influence risk of disease in HLA-B*27 positive, but not negative individuals, strongly support the notion that both these molecules act in the same biological pathway to affect disease susceptibility7. We have now shown that HLA-B*40:01 interacts with ERAP1 variants in the same manner. Similar genetic interactions involving ERAP1 have been observed in two other immune-mediated disorders—psoriasis with HLA-Cw6 (ref. 37) and Behçet’s syndrome with HLA-B*51 (ref. 38), two disorders that are already known to share genetic susceptibility factors with AS. It is likely that the similar molecular mechanisms are involved in these disorders, and that these include the pathways of MHC class I antigen presentation. To our knowledge, there is no evidence that HLA-B40, HLA-B51 or HLA-Cw6 have non-canonical disease-related properties such as those by which HLA-B27 is proposed to function in the pathogenesis of AS.
Analysis of polymorphic amino-acid positions in these AS-associated HLA molecules showed that the SNPs with the strongest evidence of association at each of these three loci were highly associated with amino-acid positions located in the peptide-binding groove of these proteins. From these results, we infer that antigen presentation to both CD4+ and CD8+ T lymphocytes is likely to be important in the pathogenesis of AS and/or its tissue specificity, although other mechanisms underlying the associations cannot formally be excluded.
MHC class I molecules contain six specificity pockets in the peptide-binding groove, alphabetically named A to F, which serve to anchor particular side chains of the bound peptide39. Position 97 of HLA-B is located in the C/F pocket, also referred to as the C-terminal pocket, which anchors the side chain of the C-terminal peptide residue29. Experimental evidence suggests that this position is important for protein function and shaping the peptide repertoire presented by HLA-B. Mutagenesis experiments have shown that Asn97 is important for HLA-B*27:04 surface expression; mutating this residue from Asn97 to Asp97 results in reduced surface expression and increased accumulation of unfolded protein in the ER, as well as reduced homodimers formation40; thus, Asn97 relative to Asp97 reduces ER stress and B27 homodimer formation, yet is associated with AS risk. Moreover, work in the mouse homologue has shown that changing residue 97 (W97R) results in altered peptide specificity and affinity for β2-microglobulin41, and previous crystallographic studies of viral peptides bound to HLA-B27 have shown that this position influences the location of the peptide in the binding groove of the molecule42. Last, this position was also found to be associated with HIV-1 viral control, where Val97 was found to provide the strongest protective effect to progression to AIDS, hypothesized to be through a mechanism of peptide presentation43. Asp97 is not shared by the AS-associated subtype HLA-B*27:07, where it is substituted by serine. Serine is also a polar amino acid and the substitution would be expected to have only minor effects on the protein structure. While AS is known to occur in individuals carrying HLA-B*27:07, its relative strength of association compared with other AS-associated HLA-B*27 subtypes is unknown.
In summary, with high-density genotyping of the MHC, we have demonstrated independent association signals located in HLA class I and II loci. Imputation of amino-acid residues in the classical HLA class I and II proteins resolved the peak of association at each of these loci to an amino-acid residue located in the peptide-binding groove of these proteins. Refining this analysis by imputation of classical HLA alleles showed that there are multiple risk and protective haplotypes in the HLA-B locus. Further, epistatic interaction was demonstrated between ERAP1 and the HLA class I alleles HLA-B*27 and HLA-B*40:01.
Methods
Sample collection
All cases met the modified New York classification criteria for AS44. Nine thousand sixty-nine cases and 13,578 controls were recruited through a multi-center study coordinated by the International Genetics of Ankylosing Spondylitis Consortium8, and all samples were unrelated and met European ancestry criteria as detailed therein. All subjects provided written informed consent and the study was approved by the Princess Alexandra Hospital Research Ethics Committee (reference HREC/05/QPAH/221) and University of Queensland Research Ethics Committee (Project Clearance No: 2006000102). All samples were genotyped with the Illumina (San Diego, CA, USA) Infinium platform Immunochip45, and the current study was restricted to 7,264 markers in the MHC (chromosome 6, bps 29,602,876–33,268,403, NCBI Build 36 human genome coordinates).
Imputation
We imputed SNPs across the MHC, and classical HLA class I and II alleles (HLA-A, HLA-C, HLA-B, HLA-DRBI, HLA-DQA1, HLA-DQB1, HLA-DPA1 and HLA-DPB1) and their corresponding amino acids determinants with SNP2HLA28. Samples with cumulative dosage above 2.5, across all four-digit alleles for any one of the HLA loci, were removed from the analysis. SNPs, alleles or amino-acid residues were excluded from the analysis if the r2 imputation quality score was below 0.2.
Classical allele imputation at the HLA-B locus resulted in high-quality data, with a median sensitivity and specificity for imputed HLA-B alleles of 0.958 and 0.998, respectively (Supplementary Fig. 1). With our imputation strategy, similar imputation performance has previously been shown for the other HLA class I and II loci (HLA-A, HLA-C, HLA-DRB1, HLA-DQA1, HLA-DQB1, HLA-DPA1 and HLA-DPB1), suggesting that imputation performance for these loci was also accurate in our study28,43,46,47.
Statistical framework for association analysis
Associations of SNPs, HLA protein amino-acid positions and non-HLA-B alleles across the MHC locus were assessed with logistic regression, assuming an additive risk effect on the log-odds scale. To account for population stratification, we included as covariates 10 principal components for each individual, computed with 16,145 unlinked autosomal, non-MHC, SNPs with the tool shellfish ( http://www.stats.ox.ac.uk/~davison/software/shellfish/shellfish.php). The omnibus association test compares, via likelihood ratio test, the null model H0, where there is no risk effect at the position tested, against the alternative model H1, where the risk effect at the position is included in the model as a fixed effect:


where yi denotes the binary phenotype code for individual i (0=control and 1=case). The πk parameter is the effect associated with each of the principal components and pi,k is the value of the kth principal component for individual i . The θ parameter represents the sampling fraction (that is, the logistic regression intercept). In the alternative model, a indicates the specific allele being tested and ga,i is the dosage (imputed or genotyped) of allele a in individual i. The βa parameter represents the effect on the log odds of disease per allele. For testing a multi-allelic locus, nucleic or amino-acid positions, with m possible alleles we included m-1 β parameters, one for each allele, where the most common allele was selected as the reference allele. The likelihood ratio test that compares model H0 with H1 results in a test statistic that is χ2 distributed with m-1 degrees of freedom.
When testing for association with imputed classical HLA alleles, we defined a series of binary markers coding the presence or absence of the allele being tested, and each different allele was tested as a biallelic position as described above.
To identify independent effects. we performed conditional logistic regression by including the most strongly associated position/polymorphism as a fixed effect in both the null model H0 and the alternative model H1. We then analysed all positions as described above. Conditional analysis was repeated in an iterative fashion by sequentially adding the most significant positions as fixed effects until no significant position or polymorphism was observed. Allelic associations were deemed significant with P<10−5, this statistical significance threshold accounted for 5,000 independent tests using Bonferroni correction. Two tests were considered independent if the two SNPs had a pairwise correlation (r2)<0.90, which resulted in 3,252 SNPs independent tests. For the special case of HLA-B alleles where we had a higher prior probability of association, we defined significance as P<10−3 as only 38 alleles were tested.
Additional information
How to cite this article: Cortes, A. et al. Major histocompatibility complex associations of ankylosing spondylitis are complex and involve further epistasis with ERAP1. Nat. Commun. 6:7146 doi: 10.1038/ncomms8146 (2015).
References
Brown, M. A. et al. Susceptibility to ankylosing spondylitis in twins: the role of genes, HLA, and the environment. Arthritis Rheum. 40, 1823–1828 (1997).
Brewerton, D. A. et al. Ankylosing spondylitis and HL-A 27. Lancet 1, 904–907 (1973).
Caffrey, M. F. & James, D. C. Human lymphocyte antigen association in ankylosing spondylitis. Nature 242, 121 (1973).
Schlosstein, L., Terasaki, P. I., Bluestone, R. & Pearson, C. M. High association of an HL-A antigen, W27, with ankylosing spondylitis. N. Engl. J. Med. 288, 704–706 (1973).
Burton, P. R. et al. Association scan of 14,500 nonsynonymous SNPs in four diseases identifies autoimmunity variants. Nat. Genet. 39, 1329–1337 (2007).
Reveille, J. D. et al. Genome-wide association study of ankylosing spondylitis identifies non-MHC susceptibility loci. Nat. Genet. 42, 123–127 (2010).
Evans, D. M. et al. Interaction between ERAP1 and HLA-B27 in ankylosing spondylitis implicates peptide handling in the mechanism for HLA-B27 in disease susceptibility. Nat. Genet. 43, 761–767 (2011).
International Genetics of Ankylosing Spondylitis Consortium (IGAS). et al. Identification of multiple risk variants for ankylosing spondylitis through high-density genotyping of immune-related loci. Nat. Genet. 45, 730–738 (2013).
Benjamin, R. & Parham, P. HLA-B27 and disease: a consequence of inadvertent antigen presentation? Rheum. Dis. Clin. North Am. 18, 11–21 (1992).
Kenna, T. J. & Brown, M. A. Immunopathogenesis of ankylosing spondylitis. Int. J. Clin. Rheumatol. 8, 265–274 (2013).
Sherlock, J. P., Buckley, C. D. & Cua, D. J. The critical role of interleukin-23 in spondyloarthropathy. Mol. Immunol. 57, 38–43 (2014).
Chang, S. C., Momburg, F., Bhutani, N. & Goldberg, A. L. The ER aminopeptidase, ERAP1, trims precursors to lengths of MHC class I peptides by a ’molecular ruler’ mechanism. Proc. Natl Acad. Sci. USA 102, 17107–17112 (2005).
Saveanu, L. et al. Concerted peptide trimming by human ERAP1 and ERAP2 aminopeptidase complexes in the endoplasmic reticulum. Nat. Immunol. 6, 689–697 (2005).
Allen, R. L., O'Callaghan, C. A., McMichael, A. J. & Bowness, P. Cutting edge: HLA-B27 can form a novel beta 2-microglobulin-free heavy chain homodimer structure. J. Immunol. 162, 5045–5048 (1999).
Chan, A. T., Kollnberger, S. D., Wedderburn, L. R. & Bowness, P. Expansion and enhanced survival of natural killer cells expressing the killer immunoglobulin-like receptor KIR3DL2 in spondylarthritis. Arthritis Rheum. 52, 3586–3595 (2005).
Diaz-Pena, R. et al. Influence of HLA-B*5703 and HLA-B*1403 on susceptibility to spondyloarthropathies in the Zambian population. J. Rheumatol. 35, 2236–2240 (2008).
Lopez-Larrea, C. et al. Association of ankylosing spondylitis with HLA-B*1403 in a West African population. Arthritis Rheum. 46, 2968–2971 (2002).
D'Amato, M. et al. Relevance of residue 116 of HLA-B27 in determining susceptibility to ankylosing spondylitis. Eur. J. Immunol. 25, 3199–3201 (1995).
Nasution, A. R. et al. HLA-B27 subtypes positively and negatively associated with spondyloarthropathy. J. Rheumatol. 24, 1111–1114 (1997).
Turner, M. J. et al. HLA-B27 misfolding in transgenic rats is associated with activation of the unfolded protein response. J. Immunol. 175, 2438–2448 (2005).
DeLay, M. L. et al. HLA-B27 misfolding and the unfolded protein response augment interleukin-23 production and are associated with Th17 activation in transgenic rats. Arthritis Rheum. 60, 2633–2643 (2009).
Campbell, E. C., Fettke, F., Bhat, S., Morley, K. D. & Powis, S. J. Expression of MHC class I dimers and ERAP1 in an ankylosing spondylitis patient cohort. Immunology 133, 379–385 (2011).
Ciccia, F. et al. Evidence that autophagy, but not the unfolded protein response, regulates the expression of IL-23 in the gut of patients with ankylosing spondylitis and subclinical gut inflammation. Ann. Rheum. Dis. 73, 1566–1574 (2014).
Kenna, T. J. et al. Disease-associated polymorphisms in ERAP1 do not alter endoplasmic reticulum stress in patients with ankylosing spondylitis. Genes Immun. 16, 35–42 (2015).
Robinson, W. P. et al. HLA-Bw60 increases susceptibility to ankylosing spondylitis in HLA-B27+ patients. Arthritis Rheum. 32, 1135–1141 (1989).
Brown, M. A. et al. HLA class I associations of ankylosing spondylitis in the white population in the United Kingdom. Ann. Rheum. Dis. 55, 268–270 (1996).
Wei, J. C., Tsai, W. C., Lin, H. S., Tsai, C. Y. & Chou, C. T. HLA-B60 and B61 are strongly associated with ankylosing spondylitis in HLA-B27-negative Taiwan Chinese patients. Rheumatology (Oxford) 43, 839–842 (2004).
Jia, X. et al. Imputing amino Acid polymorphisms in human leukocyte antigens. PLoS One 8, e64683 (2013).
Madden, D. R., Gorga, J. C., Strominger, J. L. & Wiley, D. C. The three-dimensional structure of HLA-B27 at 2.1 A resolution suggests a general mechanism for tight peptide binding to MHC. Cell 70, 1035–1048 (1992).
van Gaalen, F. A. et al. Epistasis between two HLA antigens defines a subset of individuals at a very high risk for ankylosing spondylitis. Ann. Rheum. Dis. 72, 974–978 (2012).
Takano, M., Miyajima, T., Kiuchi, M., Ohmori, K. & Amemiya, H. Behcet disease and the HLA system. Tissue Antigens 8, 95–99 (1976).
Ait Badi, M. A. et al. [Skeletal manifestations in Behcet's disease. A report of 79 cases]. Rev. Med. Interne 29, 277–282 (2008).
Taurog, J. D. & Hammer, R. E. Experimental spondyloarthropathy in HLA-B27 transgenic rats. Clin. Rheumatol. 15, (Suppl 1): 22–27 (1996).
Akram, A. & Inman, R. D. Co-expression of HLA-B7 and HLA-B27 alleles is associated with B7-restricted immunodominant responses following influenza infection. Eur. J. Immunol. 43, 3254–3267 (2013).
Akram, A., Lin, A., Gracey, E., Streutker, C. J. & Inman, R. D. HLA-B27, but not HLA-B7, immunodominance to influenza is ERAP dependent. J. Immunol. 192, 5520–5528 (2014).
Sawcer, S. et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476, 214–219 (2011).
Strange, A. et al. A genome-wide association study identifies new psoriasis susceptibility loci and an interaction between HLA-C and ERAP1. Nat. Genet. 42, 985–990 (2010).
Kirino, Y. et al. Genome-wide association analysis identifies new susceptibility loci for Behcet's disease and epistasis between HLA-B*51 and ERAP1. Nat. Genet. 45, 202–207 (2013).
Saper, M. A., Bjorkman, P. J. & Wiley, D. C. Refined structure of the human histocompatibility antigen HLA-A2 at 2.6 A resolution. J. Mol. Biol. 219, 277–319 (1991).
Blanco-Gelaz, M. A. et al. The amino acid at position 97 is involved in folding and surface expression of HLA-B27. Int. Immunol. 18, 211–220 (2006).
Smith, R. A., Myers, N. B., Robinson, M., Hansen, T. H. & Lee, D. R. Polymorphism at position 97 in MHC class I molecules affects peptide specificity, cell surface stability, and affinity for beta2-microglobulin. J. Immunol. 169, 3105–3111 (2002).
Kumar, P. et al. Structural basis for T cell alloreactivity among three HLA-B14 and HLA-B27 antigens. J. Biol. Chem. 284, 29784–29797 (2009).
Pereyra, F. et al. The major genetic determinants of HIV-1 control affect HLA class I peptide presentation. Science 330, 1551–1557 (2010).
van der Linden, S., Valkenburg, H. A. & Cats, A. Evaluation of diagnostic criteria for ankylosing spondylitis. A proposal for modification of the New York criteria. Arthritis Rheum. 27, 361–368 (1984).
Cortes, A. & Brown, M. A. Promise and pitfalls of the Immunochip. Arthritis Res. Ther. 13, 101 (2011).
McLaren, P. J. et al. Fine-mapping classical HLA variation associated with durable host control of HIV-1 infection in African Americans. Hum. Mol. Genet. 21, 4334–4347 (2012).
Raychaudhuri, S. et al. Five amino acids in three HLA proteins explain most of the association between MHC and seropositive rheumatoid arthritis. Nat. Genet. 44, 291–296 (2012).
Acknowledgements
We thank all participating subjects with AS and healthy individuals who provided the DNA and clinical information necessary for this study. This work was in part funded by grants from Arthritis Research UK (19536 and 18797), the NIHR Oxford comprehensive Biomedical Research Centre (immunity and inflammation theme A93081) and NIHR Thames Valley collaborative research network and National Ankylosing Spondylitis Society (UK). SPARCC was established through the support of the Arthritis Society of Canada. Support was received from National Institutes of Health/National Institute of Allergy and Infectious Diseases grant 1U01AI09090-01. This work was supported in part by grant PI12/02587 (Inst. Carlos III, Spain) and by European Union ‘Fondos FEDER’. Support was received from Agence Nationale de la Recherche (grant ANR 2010 GEMISA and Investissements d’Avenir programme ANR-11-IDEX-0005-02), the Société Française de Rhumatologie (SFR) and the Arthritis Foundation. M.W. is funded by the Intramural Research Program, National Institute of Arthritis and Musculoskeletal and Skin Diseases, National Institutes of Health. M.A.B. is funded by a National Health and Medical Research Council (Australia) Senior Principal Research Fellowship. D.M.E. is funded by an Australian Research Council Future Fellowship (FT130101709). P.I.W.d.B. is funded in part by the Netherlands Organization for Scientific Research (VIDI Vernieuwingsimpuls project 016.126.354) and by the National Institutes of Health (1R01AR062886-1).
Author information
Authors and Affiliations
Contributions
All authors contributed to the manuscript preparation and approved the final manuscript. Case and control recruitment was performed by P.L.C., M.H.H., M.W., L.S.G., J.H., O.F., J.T., K.L., L.A.B., D.E., R.B-V., S.S., C.F., N.H., J.M., F.J.B., M.A.G-G., C.L-L, P.B., K.G., H.G., D.D.G., P.R., W.P.M., I.V.H-B., R.V-O., C.R-S., I.M.H., F.M.P-S., R.D.I., M.B., B.P.W., J.D.R. and M.A.B. Study design was contributed by A.C., P.J.L., R.D.I., M.B., B.P.W., J.D.R., D.M.E., P.I.W.d.B. and M.A.B. DNA preparation and genotyping were performed by J.J.P., X.Z., H-J.G., G.C., B.A.L., L.A., J.L., J.B.A.C., F.M.P-S. and M.A.B.. Analysis and interpretation were performed by A.C., S.L.P., P.J.L., P.C.R., D.M.E., P.I.W.d.B. and M.A.B. The manuscript preparation was performed by A.C., P.J.L., D.M.E., P.I.W.d.B. and M.A.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figure 1 and Supplementary Tables 1-8 (PDF 450 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Cortes, A., Pulit, S., Leo, P. et al. Major histocompatibility complex associations of ankylosing spondylitis are complex and involve further epistasis with ERAP1. Nat Commun 6, 7146 (2015). https://doi.org/10.1038/ncomms8146
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms8146
This article is cited by
-
Zur Rolle von HLA-B27 in der Pathogenese und Diagnostik der axialen Spondyloarthritis
Zeitschrift für Rheumatologie (2024)
-
Predominant ligament-centric soft-tissue involvement differentiates axial psoriatic arthritis from ankylosing spondylitis
Nature Reviews Rheumatology (2023)
-
How Has Molecular Biology Enhanced Our Undertaking of axSpA and Its Management
Current Rheumatology Reports (2023)
-
A common human MLKL polymorphism confers resistance to negative regulation by phosphorylation
Nature Communications (2023)
-
Markers of immune dysregulation in response to the ageing gut: insights from aged murine gut microbiota transplants
BMC Gastroenterology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.