Abstract
Increasingly complex metabolic pathways have been engineered by modifying natural pathways and establishing de novo pathways with enzymes from a variety of organisms. Here we apply retro-biosynthetic screening to a modular pathway design to identify a redox neutral, theoretically high yielding route to a branched C6 alcohol. Enzymes capable of converting natural E. coli metabolites into 4-methyl-pentanol (4MP) via coenzyme A (CoA)-dependent chemistry were taken from nine different organisms to form a ten-step de novo pathway. Selectivity for 4MP is enhanced through the use of key enzymes acting on acyl-CoA intermediates, a carboxylic acid reductase from Nocardia iowensis and an alcohol dehydrogenase from Leifsonia sp. strain S749. One implementation of the full pathway from glucose demonstrates selective carbon chain extension and acid reduction with 4MP constituting 81% (90±7 mg l−1) of the observed alcohol products. The highest observed 4MP titre is 192±23 mg l−1. These results demonstrate the ability of modular pathway screening to facilitate de novo pathway engineering.
Similar content being viewed by others
Introduction
Increasing interest in sustainable production of a wide range of chemical products has encouraged development of microbial catalysts for the conversion of renewable feedstocks to specialty and bulk chemicals as well as transportation fuels. Although natural hosts and metabolic pathways have been used for decades in the production of chemical products, the desire to synthesize direct replacements for current petroleum products has led to appropriation of natural pathways for production of noncognate chemicals1,2,3. Early examples of this new paradigm have focused on introducing or combining portions of natural pathways in alternative host organisms or creating new products by altering the termination of natural pathways of a host organism with promiscuous enzymes1,3,4,5,6,7,8,9,10,11. Recent work has been focused on improving modified natural pathways by substituting new enzymes to improve kinetics, improve expression or utilize alternate cofactors12,13,14. In some cases, pathways have been repurposed to synthesize new products by capitalizing on the natural capacity of enzymes to accept closely related substrates or by engineering protein specificity15,16,17. The boldest designs have utilized previously undescribed pathways created by combining the natural chemistry of individual enzymes from multiple hosts18,19,20.
Liquid transportation fuels are one class of chemical targets for which natural pathways to exact replacements have not been discovered. Recent interest in microbial production of renewable fuels has led to successful synthesis of a variety of next-generation biofuels with improved properties over ethanol12,14,15,21,22,23,24,25,26,27. Microbial synthesis of these reduced chemical species by de novo designed pathways can potentially lead to more efficient production strains, which are necessary for next-generation targets to achieve commercial relevance. Predominantly, these pathways have employed reconstitution of natural pathways in new hosts (acetone-butanol-ethanol5,6) or modified termination of natural pathways (fatty acid synthesis (FAS)11,23,24, amino acid synthesis12,28, isoprenoid synthesis22).
Although natural metabolites of lipid and terpene synthesis closely resemble biodiesel, there are few natural metabolites that closely mimic gasoline. Because gasoline constitutes 40% of total US petroleum consumption, a bio-based gasoline alternative would be useful for alleviating petroleum reliance. Two synergistic approaches are required for making a bio-based gasoline more economical: utilization of less expensive feedstocks and implementation of high-efficiency pathways29,30,31,32. The presented work focuses on development of a high-efficiency pathway for a bio-alternative alcohol for spark ignition engines. Such an alcohol would ideally fall in the C6–C7 range25,26,33. These medium-chain length alcohols achieve energy density equal to that of petroleum-derived gasoline (32 MJ l−1; Supplementary Fig. 1a)34,35,36,37. Branched alcohols have the additional desired property of improved octane rating (Supplementary Fig. 1b)33. To date, the best examples of synthesizing such compounds have come from modified termination of branched amino-acid synthesis or from isoprenoid synthesis22,28. The amino acid-based pathway utilizes carbon chain extension by engineering α-ketoacid elongation (αKAE) enzymes and an α-ketoacid decarboxylase to produce a blend of alcohols. The inefficient carbon chain extension mechanism creates a redox imbalance for medium-chain products, which limits maximum pathway efficiency38. The isoprenoid pathway can be used to produce C5 isopentenol, but it is limited to forming carbon chain lengths in multiples of five because it uses isopentenyl diphosphate as a C5 building block, and it is limited by the inherent redox imbalance of the pathway.
Here we present a pathway design, which combines a portion of a native pathway (valine biosynthesis) with a ten-step de novo pathway to produce 4-methyl-pentanol (4MP). To identify specific pathway variants, we utilize a conceptual modular framework based on general natural chemistries. Biosynthetic routes to alkyl chains most commonly employ a system of precursor generation followed by chain elongation through carbon–carbon bond-forming reactions39,40,41. We structure our conceptual modules to correspond to this pathway structure. We envision precursor generating modules (glycolysis and modules 1 and 2) and carbon chain elongation modules (module 3) coupled to alcohol terminating modules (module 4).
This approach produces a pathway with enzymes selected from nine different metabolic contexts (organisms and/or pathways). We select four enzymes to act on their presumed cognate substrates and apply six to presumed noncognate substrates in the engineered pathway. The core module 3 pathway architecture is based on synthetic coenzyme A (CoA)-dependent chemistry first understood in the acetone-butanol-ethanol pathway of Clostridium acetobutylicum. This CoA-dependent architecture has several advantages compared with the αKAE and isoprenoid pathways described above. Biosynthetic CoA-dependent pathways typically utilize acetyl-CoA building blocks and extend carbon chains through condensation reactions without release of CO2. The potential for generation of two reducing equivalents per acetyl-CoA generated from glycolysis perfectly balances with those consumed for reduction to a primary alcohol product15,42. Indeed, the presented pathway achieves redox neutrality and a maximum theoretical yield of 0.67 mol 4MP per mole glucose (0.38 gg−1, 100% maximum pathway energy efficiency). β-Oxidation chemistry has been used for synthesis of several straight-chain acids and alcohols5,6,16,21,43. Unlike these previous demonstrations of CoA-dependent pathways, here we present the expansion of potential products to branched alcohols of medium chain length using independently selected enzymes chosen to enhance specificity for our desired intermediates.
Results
4MP pathway description
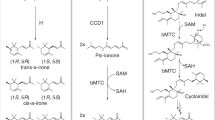
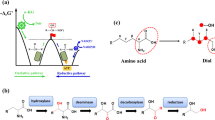
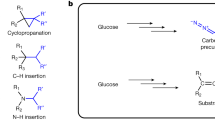
The 4MP pathway design does not rely on the simple transfer of a single recombinant pathway; rather, it relies on a patchwork of enzymes from multiple organisms and multiple natural pathways (Fig. 1). Figure 1 presents the overall pathway as a composite of four modules. Module 1, an adaptation of previously described routes to isobutanol and isobutyrate12,44, converts pyruvate, from glycolysis, to α-keto-isovalerate (α-KIV) via valine biosynthesis using Bacillus subtilis acetolactate synthase AlsSBs, Escherichia coli acetohydroxy acid isomeroreductase IlvCEc and E. coli dihydroxy acid dehydratase IlvDEc. α-KIV is further converted to isobutyrate by the Lactococcus lactis decarboxylase KivDLl and an isobutyraldehyde preferring aldehyde dehydrogenase from Flavobacterium johnsonaie (Fjoh_2967)45. Module 2 consists of the ATP-dependent activator Rhodopseudomonas palustris isobutyryl-CoA ligase IbuARp, which converts isobutyrate to isobutyryl-CoA46. Acetyl-CoA, generated by the endogenous pyruvate decarboxylase complex, condenses with isobutyryl-CoA in the first reaction of module 3, mediated by the Cupriavidus necator thiolase BktBCn. The subsequent reactions of module 3 reduce the branched 3-keto-4-methylvaleryl-CoA intermediate to 4-methyl-valeryl-CoA by C. necator acetoacetyl-CoA reductase PhaBCn, C. necator enoyl-CoA hydratase PhaJ4bCn and Treponema denticola enoyl-CoA reductase TerTd (refs 14, 15, 47, 48, 49, 50, 51). Endogenous thioesterase activity (potentially from TesB and YdiI52) generates free 4-methyl-valerate (4MV). Module 4 reduces the free 4MV to 4MP by the Nocardia iowensis carboxylic acid reductase CarNi and either Saccharomyces cerevisiae alcohol dehydrogenase Adh6pSc or Leifsonia sp. strain S749 alcohol dehydrogenase Lsadh53,54,55,56,57 (see Supplementary Fig. 2 for enzyme cofactor requirments and by-product reactions). Throughout this manuscript, strain names indicate the modules present in the strain (that is, M1F2I34 includes ‘M’ for modules, ‘1F’ for module 1 with feaBEc, ‘2I’ for module 2 with ibuARp, ‘3’ for module 3 and ‘4’ for module 4). Strain names with ‘( )’ contain abbreviations for operon structure indicating the order of alsSBs and ilvCEc (that is, M1F(IA)2I34 includes (IA) indicating an ilvCEc-alsSBs operon order). See Table 1 and Supplementary Table 1 for descriptions of all strains used and Supplementary Table 2 for relevant descriptions of enzymes used in all pathway variants.
(a) The α-ketoacid elongation (αKAE) pathway was previously used to synthesize 4MP among other products. The αKAE route utilizes relatively inefficient single-carbon extension and non-specific decarboxylation and reduction of upstream precursors resulting in a redox imbalance and a mix of products, three of which are shown. (b) The presented CoA-dependent pathway to 4MP is assembled with genes from nine organisms taken from ten different pathway contexts. Pathway genes are shown with known native pathways or putative metabolic roles. Selectivity for 4MP was achieved while requiring enzymes for modules 3 and 4 to act on noncognate substrates. (c) The 4MP pathway can be organized into four modules, which were used to identify better performing enzymes for individual steps and validate portions of the overall pathway independently in vivo: Module 1, modified valine biosynthesis to isobutyrate; module 2, isobutyrate activation to isobutyryl-CoA; module 3, CoA-dependent condensation and reduction of isobutyryl-CoA and acetyl-CoA to 4-methyl-valerate (4MV); module 4, reduction of 4MV to 4MP. Genes in italics were overexpressed from plasmid sets. Modules were constructed working backwards from the 4MP product. A potential by-product shunt to butyrate and butanol was monitored during pathway construction.
Identification of acid and aldehyde reductases (module 4)
Adoption of a CoA-dependent synthesis route required identification of pathways to link a saturated CoA thioester to the final alcohol product. The CoA thioester could be reduced by either a CoA-dependent aldehyde dehydrogenase or thioesterase/carboxylic acid reductase pairing. The resulting aldehyde could then be reduced by an alcohol dehydrogenase to generate the primary alcohol product. The wide array of identified alcohol dehydrogenases created a high probability that an alcohol dehydrogenase could be found for conversion to the final alcohol13,58,59. S. cerevisiae Adh6pSc was initially selected because it was previously found to be a broad specificity alcohol dehydrogenase with high activity on medium- and branched-chain aliphatic aldehydes53.
A smaller number of acid or CoA-thioester reductases have been characterized in the literature. From these previously identified enzymes, we looked to identify candidates with the potential to selectively convert 4MV to 4-methyl-valeraldehyde. Recently, a carboxylic acid reductase (Car) from Mycobacterium marinum was shown to convert a range of straight-chain fatty acids to fatty aldehydes, but with increasing catalytic efficiency (kcat/KM) for longer chain lengths60. A previously studied homologue, Car from N. iowensis, was found to have activity on a broad array of acids, but its activity on a range of aliphatic acids was not examined61. CarNi from N. iowensis was selected for further study to determine whether it had specificity for targeted medium-chain branched acids.
Assays were devised to confirm activity on desired substrates in vitro and in vivo. First, N-terminal His-tagged CarNi was purified and assayed for relative activity on 13 straight and branched acid substrates from C2 to C8. CarNi showed a peak in activity for acids with a primary chain-length of five or six carbons (Fig. 2a). The highest CarNi activity was found for the branched species 4MV and 4-methyl-hexanoate. Given the need to reduce flux of precursors to undesired by-product alcohols, CarNi was a logical selection because of its preference for 4MV over the short-chain acids acetate, isobutyrate and butyrate. A complementary in vivo assay was designed to examine the effectiveness of the module 4 pairing. Butyrate, valerate, 3-methyl-valerate, 4MV and hexanoate were fed to E. coli cultures expressing CarNi/Adh6pSc and conversion was monitored by sampling the culture media. OD600-normalized conversions followed a trend similar to observed in vitro results with maximal conversion for substrates with C5 primary chain length, as desired (Supplementary Fig. 3).
(a) In vitro analysis of His-purified CarNi reveals a dependence on acid primary-chain length with maximum activity at a chain length of five and six carbons. Branching at the C4 position is preferred significantly over straight acid species. The potential substrates for by-product formation, butyrate and isobutyrate, are seen to have 56% and 25% of the observed activity on 4-methyl-valerate (4MV), respectively. (b) Michaelis–Menten kinetics for isobutyrate and 4MV reveal that CarNi has a strong preference for the latter intermediate. (c) CarNi substrate preference influences product selectivity in vivo generating 1.9 times as much 4-methyl-pentanol (4MP; 272 mg l−1, 2.7 mM) as butanol (142 mg l−1, 1.9 mM) in strain M2P34 (expressing modules 2(pct), 3 and 4 genes) supplied with both glucose and isobutyrate. Even though 10 mM isobutyrate is supplied to the cultures of strain M2P34 only 111 mg l−1 (1.5 mM) isobutanol is observed. All data are presented as the mean±s.d. (n=3) with in vivo data generated using biological triplicates.
Strong specificity of CarNi for 4MV over isobutyrate was desired because isobutyrate is produced as an upstream intermediate. The Michaelis–Menten kinetic parameters of CarNi were found using these two key intermediates to confirm the desired specificity (Fig. 2b). The kcat/Km ratio with 4MV was found to be 450 times higher than with isobutyrate, indicating the significant preference of CarNi for 4MV over other acid substrates generated by the pathway. With a Km of 78±9 mM with isobutyrate, CarNi is expected to convert isobutyrate poorly under physiologically relevant concentrations, which limits shunting of the precursor to isobutyraldehyde.
Identification of dehydratases and reductases (module 3)
Enzymes of the Clostridium acetobutylicum butanol pathway and enzymes from polyhydroxyalkanoate pathways have previously been used to synthesize straight chain butanol and pentanol14,15,21. Previous work from our group found the bktBCn/phaBCn combination is capable of synthesizing 3-hydroxy-4-methylvaleryl-CoA from isobutyryl-CoA and acetyl-CoA50. To the best of our knowledge, 3-hydroxy-acyl-CoA dehydratases and trans-enoyl reductases with activity on the subsequent branched intermediates have not been identified. From enzymes documented to have activity on straight medium-chain CoA substrates, 4 phaJ and 6 ter homologues were selected for further screening48,49 (Supplementary Table 1). An assay was developed to screen for enzymes with the desired activity by isolating modules 2 and 3 of our pathway in vivo with different combinations of dehydratases and reductases. Isobutyrate (10 mM) and glucose (1%) were supplied in lysogeny broth (LB) medium, and active gene combinations were identified by detecting 4MV secretion. The propionyl-CoA transferase from Megasphera elsdenii, pctMe, was used to activate isobutyrate62. Of the 24 combinations tested, phaJ4 homologues from Pseudomonas syringae, Pseudomonas aeruginosa and C. necator in combination with ter homologues from Vibrio parahaemolyticus and T. denticola produced 4MV (Supplementary Fig. 4A). The high producer, C. necator phaJ4bCn/T. denticola terTd (strain M3Sc-TdCn), yielded 297±45 mg l−1 4MV and was selected for module 3 moving forward. The previously used dehydratase hbdCa and reductase crtCa from the C. acetobutylicum butanol pathway were also tested in place of phaBCn and phaJ4bCn (strain M3Sc-Ca; Supplementary Fig. 4B). Although some butyrate was produced by strain M3Sc-Ca, 4MV was not detected.
With module 4 in vivo activity previously confirmed, 4MP production from glucose and isobutyrate was tested with a strain expressing modules 2, 3 and 4 genes (strain M2P34). As predicted by observed activities for CarNi and Adh6pSc, strain M2P34 preferentially produced 4MP (272±7 mg l−1) over isobutanol (111±7 mg l−1) and butanol (142±9 mg l−1) even while feeding 10 mM (870 mg l−1) isobutyrate (Fig. 2c).
Glucose to isobutyryl-CoA (modules 1 and 2)
Combining isobutyrate synthesis with CoA activation supplies the necessary isobutyryl-CoA precursor for 4MP synthesis from glucose or other simple carbon sources. The module 1 pathway was identified by building from the previous work of Zhang et al., which synthesized isobutyrate in E. coli by combining valine biosynthesis with the L. lactis kivDLl decarboxylase and various native aldehyde dehydrogenases, including PuuC and FeaB44. In previous work, our group has utilized the M. elsdenii transferase PctMe for CoA activation of carboxylic acids including isobutyrate50. The ATP-dependent isobutyryl-CoA ligase (IbuARp) from R. palustris provides an alternative activation mechanism for module 2, which creates a redox neutral overall pathway46.
Multiple module combinations were used to explore the activities of the two E. coli aldehyde dehydrogenases selected for the final oxidation of isobutyraldehyde to isobutyrate. Module 1 expression with modules 2 (pctMe) and 3 led to the production of 4MV from glucose with titres up to 111±11 mg l−1 (puuCEc) and 90±9 mg l−1 (feaBEc; Supplementary Fig. 5A). Module 1 with puuCEc led to higher 4MV titres when coupled to modules 2 and 3, but it was possible that the E. coli aldehyde dehydrogenases had activity on the module 4 intermediate 4-methyl-valeraldehyde, which could regenerate 4MV and decrease reduction to 4MP when module 4 is added. Both aldehyde dehydrogenases were used for alternate versions of the full pathway, and only the feaBEc strain (strain M1F2P34) produced 4MP (67±13 mg l−1), whereas the puuCEc strain (strain M1P2P34) produced 4MV (67±11 mg l−1; Supplementary Fig. 5B). In addition, the puuCEc strain produced more butyrate (156±4 mg l−1) and less butanol (15±6 mg l−1) than the feaBEc strain (62±15 mg l−1 butyrate, 49±17 mg l−1 butanol).
Although a demonstration of 4MP synthesis from glucose was made, relatively low 4MP titres and high isobutyrate (1,113±34 mg l−1) and isobutanol (2,205±225 mg l−1) titres suggested there were possible bottlenecks in the initial strain, strain M1F2P34. The ATP-dependent isobutyrate activator, ibuARp, was used in place of the CoA transferase pctMe in order to relieve acetyl-CoA requirements and create redox neutrality for the pathway. New plasmid constructs were made to reorganize genes onto plasmids by module (Supplementary Tables 1–3). It was anticipated that operon construction would reduce enzyme expression, especially for genes in the second position, but it was unknown if the effect would be detrimental to overall production without knowledge of the rate-limiting enzyme63. Two module 3 plasmid variants were tested to explore whether PhaBRe activity could become limiting when expressed from an operon used in the new constructs (Supplementary Fig. 6A). The variant with phaBRe in the first position of a two-gene operon generated higher titres supporting the theory that phaBRe could be the limiting activity within module 3. Comparison of phaBCn expression from the two operon variants by SDS–polyacrylamide gel electrophoresis (SDS–PAGE) confirmed higher phaBCn expression when placed in the first postion (Supplementary Fig. 6B). In addition, operon variants for alsSBs and ilvCEc expression were tested to examine if better balancing of flux between module 1 and the native acetyl-CoA pathway could improve 4MP production (Supplementary Fig. 6A). Placing alsSBs in the second position while using the new plasmid constructs (strain M1F(IA)2I34) increased 4MP titres (168±31 mg l−1), whereas isobutyrate (290±24 mg l−1) and isobutanol (1,046±45 mg l−1) titres were reduced (see Supplementary Note 1 for further details).
Based on available in vitro data and the presence of 4MV (42±7 mg l−1) even for improved strain M1F(IA)2I34, it was possible that FeaBEc could be oxidizing 4-methyl-valeraldehyde into 4MV creating a futile cycle with CarNi (Fig. 3a, Supplementary Fig. 6C). An aldehyde dehydrogenase Fjoh2967Fj from F. johnsonaie had been found to prefer an isobutyraldehyde substrate over other aldehyde substrates when tested in vitro45. Replacing feaBEc with Fjoh2967Fj in strain M1Fj(IA)2I34 led to increased 4MP (193±23 mg l−1, 0.033±0.005 mol mol−1 glucose) and elimination of detectable 4MV (Fig. 3b). Isobutyrate titres (424±9 mg l−1, 0.084±0.004 mol mol−1 glucose) were increased and isobutanol titres (797±7 mg l−1, 0.187±0.006 mol mol−1 glucose) were reduced (Fig. 3c).
(a) Key reactions involving aldehydes can generate futile cycles (aldehyde dehydrogenase feaBEc with the carboxylic acid reductase carNi) or by-product shunts (alcohol dehydrogenase ADH6Sc). The desired pathway route is shown in bold arrows within the shaded box. Undesired reactions are shown with dashed arrows. Enzymes with insufficient selectivity have dashed outlines. (b) When feaBEc (Strain M1F(IA)2I34) was replaced with the isobutyraldehyde-specific aldehyde dehydrogenase Fjoh2967Fj in strain M1Fj(IA)2I34 4MV titres were reduced and 4MP titres were increased. (c) Complementarily, strain M1Fj(IA)2I34 (Fjoh2967Fj) produced lower isobutanol and higher isobutyrate titres relative to strain M1F(IA)2I34 (feaBEc). All data are presented as the mean±s.d. (n=3) with in vivo data generated using biological triplicates.
To examine if alcohol toxicity could be limiting product titres, toxicities of the dominant by-product isobutanol and desired product 4MP were assayed through exogenous addition of alcohols to the growth medium at concentrations from 1 to 10 mM. Isobutanol and 4MP concentrations up to 5 mM did not inhibit the exponential growth rate (Supplementary Fig. 7). A combination of 10 mM (741 mg l−1) isobutanol and 2 mM (204 mg l−1) 4MP (comparable to titres observed for Strain M1Fj(IA)2I34) only reduced the exponential growth rate by 10%, the same reduction observed with 10 mM isobutanol alone. Although endogenously produced alcohols may be involved in alternative toxicity mechanisms, this result suggests that current titres are likely not limited by product toxicity.
Pathway selectivity through enzyme selection (module 4)
With knowledge of the specificity of CarNi for 4MV over isobutyrate, the continued high isobutanol titres suggested Adh6pSc was converting the isobutyraldehyde intermediate to isobutanol (Fig. 4a). Removing ADH6Sc from Strain M1F(IA)2I34 produced strain M1F(IA)2I34a, which generated an isobutanol titre of 27±3 mg l−1 with nearly undetectable butanol and 4MP titres (Fig. 4b). An alternative to ADH6Sc was identified from Leifsonia sp. strain S749. The new alcohol dehydrogenase LsadhLs was hypothesized to have improved specificity for 4-methyl-valeraldehyde based on substrates that were assayed in vitro57. When lsadhLs was combined with the isobutyraldehyde-specific dehydrogenase Fjoh2967Fj in strain M1Fj(IA)2I34L, selective synthesis of 4MP was achieved over other alcohol by-products (Fig. 4c). Isobutanol titres were reduced to 21±3 mg l−1 similar to those observed with the no alcohol dehydrogenase control. 4MP was produced at 90±7 mg l−1 (0.016±0.001 mol mol−1 glucose) making up 81% of all alcohol products. The dominant by-products were the 4MP precursors acetate (592±34 mg l−1, 0.177±0.010 mol mol−1 glucose) and isobutyrate (1,128±34 mg l−1, 0.229±0.002 mol mol−1 glucose), suggesting that by-product shunts were reduced and further improvement could be made by relieving a downstream rate limitation in modules 2, 3 or 4 (Supplementary Fig. 8). Although 4MP titres were lower with LsadhLs, SDS–PAGE analysis of LsadhLs confirmed strong overexpression in E. coli (Supplementary Fig. 8C). The reduction in titre is likely due to the change from an NADPH-dependent dehydrogenase (Adh6pSc) to an NADH-dependent dehydrogenase (LsadhLs) under the aerobic conditions used. The ratio of NADH/NAD+ has been observed to be lower than that of NADPH/NAD+ under similar culture conditions64.
(a) The desired pathway reactions to 4MP are indicated by bold arrows with the by-product shunt to isobutanol indicated by the dashed arrow. High activity of Adh6pSc on isobutyraldehyde diverts isobutyrate flux to isobutanol. The LsadhLs alcohol dehydrogenase’s selectivity for 4-methyl-valeraldehyde greatly reduces flux to the isobutanol shunt. (b) The alcohol profile of strain M1F(IA)2I34 expressing feaBEc and ADH6Sc contains 168±31 mg l−1 of 4MP but is dominated by 1.046±45 g l−1 of isobutanol. The M1F(IA)2I34a control without ADH6Sc expression produces low to undetectable levels of all three alcohols. (c) Replacing feaBEc with Fjoh2967Fj in strain M1Fj(IA)2I34 reduces isobutanol (797±20 mg l−1) and increases 4MP (192±23 mg l−1) marginally. Replacing ADH6Sc with lsadhLs greatly enhanced alcohol selectivity producing 90±7 mg l−1 4MP with only 20±5 mg l−1 isobutanol. All data are presented as the mean±s.d. (n=3) with in vivo data generated using biological triplicates.
Discussion
Recent efforts to develop microbial pathways for chemical synthesis have moved beyond upregulation of native pathways to include transfer and modification of heterologous pathways to new hosts and modified termination of native host pathways. Only a small number of truly de novo pathway designs have been published and most use isolated heterologous enzymes acting on their cognate substrates18,19,20,50. Engineered pathways to liquid fuels, in particular, have predominantly relied on entirely natural (ethanol, butanol, isoprenoid) or terminally modified natural pathways (FAS, amino-acid αKAE, isoprenoid). The presented work moves beyond modification of natural pathways by successfully demonstrating synthesis of a C6 branched alcohol via an extended de novo pathway, which maintains selectivity, while utilizing multiple naturally occurring enzymes outside their native pathway contexts.
Although one set of modules has been presented in the current work, alternate chemistries could be substituted for or combined with the selected modules to create new pathways to the same or alternate products. For example, an isobutyryl-CoA mutase or branched α-keto-acid decarboxylase route could be used to generate the isobutyryl-CoA precursor in place of modules 1 and 2 (ref. 65). Similarly, a FAS route could be substituted for module 3 to generate the longer saturated acid substrate for module 4 (ref. 66). Using this design, individual alternative modules or module combinations can be directly compared with the existing pathway in vivo. In addition, entirely new classes of branched products (for example, aldehydes, alkanes) could be made by using different module 4 enzymes.
For the presented pathway, an iterative screening approach identified the enzymes catalyzing conversion of the downstream 4-methyl-valeraldehyde and upstream isobutyraldehyde intermediates as key components controlling selectivity of the pathway. Our initial module 4 alcohol dehydrogenase selection, Adh6pSc, proved to be highly active, but non-selective in the full pathway context. Module 4 displayed high activity on our desired substrate, but in vivo results with the full pathway suggested this module had a broad substrate range. Persistent high isobutanol titres from strains expressing modules 1–4 suggested that module 4 enzymes were interacting with isobutyrate and/or isobutyraldehyde. In vitro and in vivo data from module 4 testing implicated the alcohol dehydrogenase, Adh6pSc, as the non-selective enzyme (Figs 2 and 4). By replacing Adh6pSc with the isobutyraldehyde-specific and NADH-dependent alcohol dehydrogenase, LsadhLs, pathway selectivity and overall cofactor utilization were improved.
As with alcohol dehydrogenase candidates, we initially selected aldehyde dehydrogenases previously validated for an isobutyraldehyde substrate in an engineered pathway. Two endogenous enzymes, PuuCEc and FeaBEc, were previously identified as the most effective E. coli aldehyde dehydrogenases for isobutyraldehyde oxidation to isobutyrate44. Of the two E. coli aldehyde dehydrogenases, FeaBEc proved to successfully synthesize 4MP from glucose in strain M1F2P34 expressing modules 1, 2, 3 and 4 (Fig. 3b). Based on in vitro data, one may predict PuuCEc to function more effectively because its kcat/Km is more consistent across substrate lengths, whereas the kcat/Km of FeaBEc actually increases by an order of magnitude between propionaldehyde and hexanaldehyde substrates67,68. In vivo results disproved this prediction with only FeaBEc-producing 4MP (Fig. 3a). The better performance of FeaBEc in the context of the full pathway may be explained by reported Km values for the two dehydrogenases. FeaBEc has Km values below 100 μM for relevant substrates, whereas the Km values for PuuCEc are 1 mM. PuuCEc and FeaBEc were tested in strains expressing ADH6Sc. Like FeaBEc, Adh6pSc has reported Km values for relevant substrates in the 100–200 μM range53. Adh6pSc was observed to have kcat values (~100 s−1) an order of magnitude higher than values observed for FeaBEc and PuuCEc (~10 s−1) for related aliphatic aldehydes. Together, these observed kinetics support the hypothesis that Adh6pSc out-competes PuuCEc and FeaBEc for the isobutyraldehyde substrate. Isobutyraldehyde and reducing equivalents are diverted to isobutanol, lowering 4MP titres (Supplementary Fig. 5). Strain M1P2P34 with PuuCEc produces significantly more isobutanol than strain M1F2P34 with FeaBEc, as expected based on observed Km values.
In addition, in vitro data suggested that even though FeaBEc functioned as an isobutyraldehyde dehydrogenase, it may also favour a 4-methyl-valeraldehyde substrate. The potential futile cycle created by activity on 4-methyl-valeraldehyde was avoided by using the isobutyraldehyde-specific dehydrogenase Fjoh2967Fj from F. johnsonaie45. Replacing feaBEc with Fjoh2967Fj led to increased isobutyrate and eliminated detectable 4MV production (Fig. 3). Combining more selective alcohol and aldehyde dehydrogenases led to a highly selective overall pathway with the major by-product being overflow of the upstream intermediate isobutyrate (Supplementary Fig. 6). Together, the results from alcohol and aldehyde dehydrogenase selection highlight the importance of considering both high activity and required selectivity when utilizing retro-biosynthetic screening. Proposing potential upstream pathways is required to identify intermediates, which could have cross-reactivity with downstream enzymes.
Further engineering of the CoA-dependent 4MP pathway is warranted given the potential high-energy yield. Dugar and Stephanopoulos have outlined the importance of balancing reducing equivalents generated and consumed in a recombinant pathway if high yields are desired38. Using the current 4MP pathway enzymes, the overall reaction can be written as:

The reducing equivalents of the pathway are balanced, but some are contained in different cofactors. The maximum pathway energy efficiency (γP) can be calculated using the degrees of reductance and pathway stoichiometry for a glucose substrate and 4MP product. Maximum pathway energy efficiency for the αKAE pathway and the presented CoA-dependent pathway are 75% and 100%, respectively. Accounting for cofactor requirements, the adjusted pathway energy efficiencies  are 24% and 45% for the αKAE and CoA pathways, respectively (Supplementary Methods). If alternative enzymes are identified or engineered to accept NADH in place of NADPH, maximum pathway yields could be achieved under anaerobic fermentation. The maximum adjusted efficiency values for these pathway architectures then become 28% (αKAE) and 100% (CoA). The yield calculations highlight how our rational design approach leads to a pathway architecture with high-yield potential unlike inherently limited pathways utilizing modification of amino-acid synthesis.
are 24% and 45% for the αKAE and CoA pathways, respectively (Supplementary Methods). If alternative enzymes are identified or engineered to accept NADH in place of NADPH, maximum pathway yields could be achieved under anaerobic fermentation. The maximum adjusted efficiency values for these pathway architectures then become 28% (αKAE) and 100% (CoA). The yield calculations highlight how our rational design approach leads to a pathway architecture with high-yield potential unlike inherently limited pathways utilizing modification of amino-acid synthesis.
This work has identified a novel pathway for the selective synthesis of the branched medium-chain length alcohol 4MP. The highest titres (193±23 mg l−1) were achieved with strain M1Fj(IA)2I34, which expresses both Fjoh2967Fj and ADH6Sc. Selectivity was achieved by replacing ADH6Sc with lsadhLs in strain M1Fj(IA)2I34L. The 90±7 mg l−1 of 4MP produced by M1Fj(IA)2I34L represented 81% of observed alcohol products. In comparison, of the nine alcohols generated in the previous demonstration of microbial 4MP synthesis using α-KAE, 4MP (202.4±1.1 mg l−1) makes up 14% of the total alcohol product28. High potential efficiency and selectivity make our CoA pathway a preferred candidate for future engineering. Currently, the major by-products of the CoA-dependent route are the acids, acetate, isobutyrate and butyrate (Supplementary Fig. 8). We expect that a combination of tuning thioesterase/transferase activities of the host to selectively cleave the longer 4-methyl-valeryl-CoA intermediate and relieving module 3 rate limitations will further enhance titres. Ultimately, screening or engineering for NADH-dependent enzymes should produce a high yielding fermentative pathway. Our existing pathway can also be adapted to produce other branched medium-chain products by testing new downstream modules. Finally, we believe the pathway design approach described here can be useful for creation of new metabolic pathways, which rely on long de novo routes. Retro-biosynthetic screening within a designed pathway framework enables exploration of enzymatic diversity using a small number of assays while preserving a maximally efficient biochemical conversion.
Methods
Bacterial strains and plasmids
E. coli MG1655(DE3) ΔendA ΔrecA described previously was the host strain for production experiments, alcohol toxicity experiments and for protein expression analysis using cell lysates69. E. coli DH10B (Invitrogen) and ElectroTen-Blue (Stratagene) were used in plasmid cloning transformations and for plasmid propagation. E. coli BL21Star(DE3) (Life Technologies) was used for expression of carNi for purification (see Supplementary Tables 1 and 2 for strain details).
A codon-optimized version of S. cerevisiae ADH6 was purchased from DNA 2.0 and codon-optimized versions of N. iowensis car and B. subtilis sfp were purchased from GenScript (see Supplementary Methods for codon-optimized sequences). Codon-optimized T. denticola ter and E. gracilis ter were purchased from GenScript15. Leifsonia sp. strain S749 lsadh was purchased as a codon-optimized GeneArt String from Life Technologies. All other genes were amplified from genomic DNA (gDNA). B. subtilis PY79, E. coli MG1655, P. putida KT2440, C. necator (formerly R. eutropha) H16, M. elsdenii, R. palustris CGA009, P. syringae DC3000, C. acetobutylicum ATCC 824 and S. oneidensis MR-1 gDNA were prepared using the Wizard Genomic DNA purification Kit (Promega). P. aeruginosa PAO1-LAC (ATCC #47085), F. johnsonaie (ATCC #17061) and V. parahaemolyticus EB 101 (ATCC #17802) gDNA were purchased from American Type Culture Collection. Custom oligonucleotide primers were purchased (Sigma-Genosys) for PCR amplification of genes from gDNA using either Phusion High-Fidelity DNA polymerase (Finnzymes, Thermo Scientific Molecular Biology) or Q5 High-Fidelity DNA polymerase (New England Biolabs). Synthetic operons were constructed using a modified Splice by Overlap Extension PCR method.
The compatible vector set pETDuet-1, pCDFDuet-1, pACYCDuet-1 and pCOLADuet-1 was used to express single genes or synthetic operons under control of a T7lac promoter and individual ribosome-binding sites. Plasmids were constructed using standard molecular biology techniques with restriction enzymes and T4 DNA ligase purchased from New England Biolabs. Ligation products in pETDuet-1, pACYCDuet-1 and pCOLADuet-1 were used to transform E. coli DH10B and pCDFDuet-1 products were used to transform E. coli ElectroTen-Blue. Propagated constructs were purified using a QIAprep Miniprep Kit (Qiagen) and agarose gel fragments were purified using a Zymoclean Gel DNA Recovery Kit (Zymo Research). Completed constructs were used to co-transform E. coli MG1655(DE3) ΔendA ΔrecA (see Supplementary Table 3 for plasmid details).
Splice by overlap extension
Initial PCR products with homologous ends were added to a PCR mixture without additional primers and cycled through a standard PCR cycle four times with annealing temperatures set at 6 °C above, 3 °C above and at the designed melting temperature for the homology. The upstream primer for the upstream gene and the downstream primer for the downstream gene in the designed operon were then added to amplify the full-length product. A standard PCR method using the annealing temperature for the primer pair was used for final amplification.
Culture conditions
For all production experiments, triplicate seed cultures were grown from isolated colonies at 30 °C overnight in 3 ml LB medium in a 14-ml culture tube on a rotary shaker at 250 r.p.m. All production cultures were inoculated with 1% inoculum from overnight seed culture and grown at 30 °C on a rotary shaker at 250 r.p.m. Cultures were induced with 0.5 mM isopropyl-β-D-thiogalactoside (IPTG) when OD600 values reached 0.6–1.0 corresponding to mid-exponential phase. For constructs designed for 4MV production, 50 ml cultures in 250 ml shake flasks were used, and for constructs designed to produce 4MP, 3 ml cultures in 1 inch diameter 50 ml screw-cap culture tubes (Pyrex VISTA) were used. Unless otherwise stated, 1 ml culture samples were taken 48 h post induction, centrifuged to pellet cells, and the supernatant was removed for analysis.
For production of 4MV and 4MP from glucose and isobutyrate, LB medium supplemented with 1% glucose and 10 mM isobutyrate was used. For production of 4MV and 4MP from glucose, LB medium supplemented with 1.2% glucose was used. Samples were taken 48 h post induction except for initial experiments with strains M1F2P34 and M1P2P34 when samples were taken 72 h post induction.
For assessing toxicity of isobutanol and 4MP, an MG1655(DE3) ΔendA ΔrecA seed culture was grown overnight in LB medium. Duplicate 3 ml cultures in LB+1.2% glucose+alcohol were inoculated to an initial OD600 of 0.1. Cultures were contained in the same 50 ml screw-cap tubes used in production experiments. Cultures contained 1 mM isobutanol, 5 mM isobutanol, 10 mM isobutanol, 1 mM 4MP, 5 mM 4MP, 10 mM 4MP or no alcohol. Growth was monitored by optical density and the growth rate was calculated from a linear regression of the natural log of the OD600 values for the 1.5, 2 and 2.5 h post inoculation time points.
Relative activity assay for purified His-Car
An overnight culture of BL21 Star (DE3) (Invitrogen) harbouring pET/His-Car-RBS2-Sfp was used as 10% (v/v) inoculum in 2 l of LB broth. The culture was incubated at 30 °C and 250 r.p.m., and expression was induced using a final concentration of 1 mM IPTG at OD 0.6. Cells were harvested after 20 h using centrifugation and resuspended in a buffer containing 50 mM Tris-HCl pH 8.0, 300 mM NaCl and 10% glycerol. Cells were subsequently lysed using sonication. The supernatant was collected and applied to a column containing Ni-NTA resin (Qiagen). Affinity chromatography was performed using step-wise increasing concentrations of imidazole. Fractions containing purified His6-Car were dialyzed overnight at 4 °C into 50 mM Tris-HCl, pH 7.5, 50 mM NaCl, 1 mM DTT and 10% glycerol. Dialyzed enzyme was then flash frozen using liquid nitrogen and stored at −80 °C.
The activity of His-Car on various substrates was determined by measuring changes in absorbance at 340 nm for up to 5 min in 96-well microplates (Tecan Infinite F200 Pro). Reactions were prepared as follows: 100 mM Tris-HCl, pH 7.5, 10 mM MgCl2, 0.6 mM NADPH, 1 mM ATP, 224 nM His-Car and 50 mM pH neutralized acid substrate. All substrates were assayed in triplicate. For KM and Vmax determinations, substrates were assayed at five different concentrations.
SDS–PAGE analysis
E. coli MG1655(DE3) ΔendA ΔrecA was transformed with empty pETDuet-1, pET-(bktBCn-terTd)-(phaBCn-phaJ4bCn), pET-(bktBCn-terTd)-(phaJ4bCn-phaBCn) pACYC-(carNi-sfpBs)-ADH6Sc or pACYC-(carNi-sfpBs)-lsadhLs. Single colonies from plates of each transformation were grown overnight in 3 ml of LB with appropriate antibiotic. Shake flask cultures (250 ml flasks) containing 50 ml LB+0.6% glucose were inoculated at 1% inoculum from overnight LB cultures and incubated with agitation at 30 °C and 250 r.p.m. Shake flasks were induced with 0.5 mM IPTG OD600 values between 0.5 and 0.6.
Five and a half hours after induction, 7 ml of each culture were sampled and pelleted by centrifugation. Cell pellets were resuspended in 1 ml of 10 mM Tris-HCl at pH 8.0 and added to 1.7 ml microcentrifuge tubes containing 500 μl of 0.1 mm diameter glass beads (Scientific Industries, Inc., Disruptor Beads, SI-BG01). Samples were then vortexed for 10 min.
After lysis, samples were pelleted by centrifugation (6,000 g, 4 °C, 10 min) and the supernatant was removed as soluble lysate. Total protein was quantified by the Bradford assay method using Bio-Rad Protein Assay Dye Reagent (Bio-Rad, Cat #500-0006) and a bovine serum album standard70. A Bio-Rad 10% Mini-PROTEAN TGX gel (Bio-Rad, Cat #456-1034) was run using the Mini-PROTEAN Tetra Cell electrophoresis set up. Bio-Rad Precision Plus Protein All Blue Standard (Bio-Rad, Cat #161-0373) and 10 μg of total protein for each sample was loaded on the gel. After running at 200 V for 33 min, the gel was washed with deionized water before staining with Bio-Rad Bio-Safe Coomassie Stain (Bio-Rad, Cat #161-0786).
Metabolite analysis
Culture samples were pelleted by centrifugation and supernatant was removed for HPLC analysis with an Agilent 1,200 series instrument (Agilent) with a refractive index detector. Analytes were separated using the Aminex HPX-87H anion exchange column (Bio-Rad Laboratories) with a 5 mM sulfuric acid mobile phase at 35 °C and a flow rate of 0.6 ml min−1. Commercial standards of glucose, α-KIV, acetate, acetoin, isobutyrate, butyrate, isobutanol, butanol, 4MV and 4MP were used for quantification of experimental samples by linear interpolation of external standard curves.
Additional information
How to cite this article: Sheppard, M. J. et al. Retro-biosynthetic screening of a modular pathway design achieves selective route for microbial synthesis of 4-methyl-pentanol. Nat. Commun. 5:5031 doi: 10.1038/ncomms6031 (2014).
References
Bachmann, B. O. Biosynthesis: is it time to go retro? Nat. Chem. Biol. 6, 390–393 (2010).
Weeks, A. M. & Chang, M. C. Y. Constructing de novo biosynthetic pathways for chemical synthesis inside living cells. Biochemistry 50, 5404–5418 (2011).
Prather, K. L. J. & Martin, C. H. De novo biosynthetic pathways: rational design of microbial chemical factories. Curr. Opin. Biotechnol. 19, 468–474 (2008).
Pfeifer, B. A., Admiraal, S. J., Gramajo, H., Cane, D. E. & Khosla, C. Biosynthesis of complex polyketides in a metabolically engineered strain of E. coli. Science 291, 1790–1792 (2001).
Atsumi, S. et al. Metabolic engineering of Escherichia coli for 1-butanol production. Metab. Eng. 10, 305–311 (2008).
Nielsen, D. R. et al. Engineering alternative butanol production platforms in heterologous bacteria. Metab. Eng. 11, 262–273 (2009).
Lee, S. Y., Yim, K. S., Chang, H. N. & Chang, Y. K. Construction of plasmids, estimation of plasmid stability, and use of stable plasmids for the production of poly(3-hydroxybutyric acid) by recombinant Escherichia coli. J. Biotechnol. 32, 203–211 (1994).
Choi J-i, Lee SY. & Han, K. Cloning of the alcaligenes latus polyhydroxyalkanoate biosynthesis genes and use of these genes for enhanced production of poly(3-hydroxybutyrate) in Escherichia coli. Appl. Environ. Microbiol. 64, 4897–4903 (1998).
Wenzel, S. C. & Müller, R. Recent developments towards the heterologous expression of complex bacterial natural product biosynthetic pathways. Curr. Opin. Biotechnol. 16, 594–606 (2005).
Voelker, T. A. & Davies, H. M. Alteration of the specificity and regulation of fatty acid synthesis of Escherichia coli by expression of a plant medium-chain acyl-acyl carrier protein thioesterase. J. Bacteriol. 176, 7320–7327 (1994).
Youngquist, J. T. et al. Production of medium chain length fatty alcohols from glucose in Escherichia coli. Metab. Eng. 20, 177–186 (2013).
Atsumi, S., Hanai, T. & Liao, J. C. Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451, 86–89 (2008).
Atsumi, S. et al. Engineering the isobutanol biosynthetic pathway in Escherichia coli by comparison of three aldehyde reductase/alcohol dehydrogenase genes. Appl. Microbiol. Biotechnol. 85, 651–657 (2010).
Bond-Watts, B. B., Bellerose, R. J. & Chang, M. C. Y. Enzyme mechanism as a kinetic control element for designing synthetic biofuel pathways. Nat. Chem. Biol. 7, 222–227 (2011).
Tseng, H.-C. & Prather, K. L. J. Controlled biosynthesis of odd-chain fuels and chemicals via engineered modular metabolic pathways. Proc. Natl Acad. Sci. 109, 17925–17930 (2012).
Dekishima, Y., Lan, E. I., Shen, C. R., Cho, K. M. & Liao, J. C. Extending carbon chain length of 1-butanol pathway for 1-hexanol synthesis from glucose by engineered Escherichia coli. J. Am. Chem. Soc. 133, 11399–11401 (2013).
Marcheschi, R. J. et al. A synthetic recursive ‘+1’ pathway for carbon chain elongation. ACS Chem. Biol. 7, 689–697 (2012).
Dueber, J. E. et al. Synthetic protein scaffolds provide modular control over metabolic flux. Nature Biotech. 27, 753–759 (2009).
Niu, W., Molefe, M. N. & Frost, J. W. Microbial synthesis of the energetic material precursor 1,2,4-butanetriol. J. Am. Chem. Soc. 125, 12998–12999 (2003).
Hansen, E. H. et al. De Novo biosynthesis of vanillin in fission yeast (Schizosaccharomyces pombe) and Baker's Yeast (Saccharomyces cerevisiae). Appl. Environ. Microbiol. 75, 2765–2774 (2009).
Shen, C. R. et al. Driving forces enable high-titer anaerobic 1-butanol synthesis in Escherichia coli. Appl. Environ. Microbiol. 77, 2905–2915 (2011).
Withers, S. T., Gottlieb, S. S., Lieu, B., Newman, J. D. & Keasling, J. D. Identification of isopentenol biosynthetic genes from Bacillus subtilis by a screening method based on isoprenoid precursor toxicity. Appl. Environ. Microbiol. 73, 6277–6283 (2007).
Torella, J. P. et al. Tailored fatty acid synthesis via dynamic control of fatty acid elongation. Proc. Natl Acad. Sci. 110, 11290–11295 (2013).
Youngquist, J. T., Rose, J. & Pfleger, B. Free fatty acid production in Escherichia coli under phosphate-limited conditions. Appl. Microbiol. Biotechnol. 97, 5149–5159 (2013).
Atsumi, S. & Liao, J. C. Metabolic engineering for advanced biofuels production from Escherichia coli. Curr. Opin. Biotechnol. 19, 414–419 (2008).
Lee, S. K., Chou, H., Ham, T. S., Lee, T. S. & Keasling, J. D. Metabolic engineering of microorganisms for biofuels production: from bugs to synthetic biology to fuels. Curr. Opin. Biotechnol. 19, 556–563 (2008).
Lennen, R. M. & Pfleger, B. F. Microbial production of fatty acid-derived fuels and chemicals. Curr. Opin. Biotechnol. 24, 1044–1053 (2013).
Zhang, K., Sawaya, M. R., Eisenberg, D. S. & Liao, J. C. Expanding metabolism for biosynthesis of nonnatural alcohols. Proc. Natl Acad. Sci. 105, 20653–20658 (2008).
Ragauskas, A. J. et al. The path forward for biofuels and biomaterials. Science 311, 484–489 (2006).
Fichman, B. T. et al. Annual Energy Review 2011 ed. Energy USDo United States Energy Information Administration (2012).
Atsumi, S., Higashide, W. & Liao, J. C. Direct photosynthetic recycling of carbon dioxide to isobutyraldehyde. Nature Biotech. 27, 1177–1180 (2009).
Machado, I. M. P. & Atsumi, S. Cyanobacterial biofuel production. J. Biotechnol. 162, 50–56 (2012).
Wallner, T., Ickes, A. & Lawyer, K. inProceedings of the FISITA 2012 World Automotive Congress Vol. 3, 15–26Springer (2012).
Chao, J. & Rossini, F. D. Heats of combustion formation and isomerization of 19 alkanols. J. Chem. Eng. Data 10, 374–379 (1965).
Goenaga, J. M., Gayol, A., Concha, R. G., Iglesias, M. & Resa, J. M. Effect of temperature on thermophysical properties of ethanol+aliphatic alcohols (C4-C5) mixtures. Monatsh. Chem. 138, 403–436 (2007).
Hales, J. L. & Ellender, J. H. Liquid densities from 293 to 490 K of nine aliphatic alcohols. J. Chem. Thermodyn. 8, 1177–1184 (1976).
Hussein, N. M. & Asfour, A-FA. Densities and kinematic viscosities of ten binary 1-alkanol liquid systems at temperatures of (293.15 and 298.15) K. J. Chem. Engin. Data 54, 2948–2952 (2009).
Dugar, D. & Stephanopoulos, G. Relative potential of biosynthetic pathways for biofuels and bio-based products. Nature Biotech. 29, 1074–1078 (2011).
Hopwood, D. A. & Sherman, D. H. Molecular genetics of polyketides and its comparison to fatty acid biosynthesis. Annu. Rev. Genet. 24, 37–62 (1990).
Lichtenthaler, H. K., Rohmer, M. & Schwender, J. Two independent biochemical pathways for isopentenyl diphosphate and isoprenoid biosynthesis in higher plants. Physiol. Plant. 101, 643–652 (1997).
Dickson, R. C. Thematic review series: sphingolipids. New insights into sphingolipid metabolism and function in budding yeast. J. Lipid. Res. 49, 909–921 (2008).
Kim, Y., Ingram, L. O. & Shanmugam, K. T. Dihydrolipoamide dehydrogenase mutation alters the NADH sensitivity of pyruvate dehydrogenase complex of Escherichia coli K-12. J. Bacteriol. 190, 3851–3858 (2008).
Dellomonaco, C., Clomburg, J. M., Miller, E. N. & Gonzalez, R. Engineered reversal of the β-oxidation cycle for the synthesis of fuels and chemicals. Nature 476, 355–359 (2011).
Zhang, K., Woodruff, A. P., Xiong, M., Zhou, J. & Dhande, Y. K. A synthetic metabolic pathway for production of the platform chemical isobutyric acid. ChemSusChem 4, 1068–1070 (2011).
Yamanaka, Y. et al. Thermostable aldehyde dehydrogenase from psychrophile, Cytophaga sp. KUC-1: enzymological characteristics and functional properties. Biochem. Biophys. Res. Commun. 298, 632–637 (2002).
Crosby, H. A., Pelletier, D. A., Hurst, G. B. & Escalante-Semerena, J. C. System-wide studies of n-lysine acetylation in Rhodopseudomonas palustris reveal substrate specificity of protein acetyltransferases. J. Biol. Chem. 287, 15590–15601 (2012).
Dennis, D., McCoy, M., Stangl, A., Valentin, H. E. & Wu, Z. Formation of poly(3-hydroxybutyrate-co-3-hydroxyhexanoate) by PHA synthase from Ralstonia eutropha. J. Biotechnol. 64, 177–186 (1998).
Kawashima, Y. et al. Characterization and functional analyses of R-specific enoyl coenzyme A hydratases in polyhydroxyalkanoate-producing Ralstonia eutropha. Appl. Environ. Microbiol. 78, 493–502 (2012).
Hoffmeister, M., Piotrowski, M., Nowitzki, U. & Martin, W. Mitochondrial trans-2-Enoyl-CoA reductase of wax ester fermentation from Euglena gracilis defines a new family of enzymes involved in lipid synthesis. J. Biol. Chem. 280, 4329–4338 (2005).
Martin, C. H., Dhamankar, H., Tseng, H.-C., Sheppard, M. J. & Reisch, C. R. Prather KLJ. A platform pathway for production of 3-hydroxyacids provides a biosynthetic route to 3-hydroxy-γ-butyrolactone. Nat. Commun. 4, 1414 (2013).
Tucci, S. & Martin, W. A novel prokaryotic trans-2-enoyl-CoA reductase from the spirochete Treponema denticola. FEBS Lett. 581, 1561–1566 (2007).
McMahon, M. D. & Prather, K. L. J. Functional screening and in vitro analysis reveal thioesterases with enhanced substrate specificity profiles that improve short-chain fatty acid production in Escherichia coli. Appl. Environ. Microbiol. 80, 1042–1050 (2014).
Larroy, C., Fernández, M. R., González, E., Parés, X. & Biosca, J. A. Characterization of the Saccharomyces cerevisiae YMR318C (ADH6) gene product as a broad specificity NADPH-dependent alcohol dehydrogenase: relevance in aldehyde reduction. Biochem. J. 361, 163–172 (2002).
Li, T. & Rosazza, J. P. Purification, characterization, and properties of an aryl aldehyde oxidoreductase from Nocardia sp. strain NRRL 5646. J. Bacteriol. 179, 3482–3487 (1997).
Venkitasubramanian, P. & Daniels, L. Rosazza JPN. Reduction of carboxylic acids by nocardia aldehyde oxidoreductase requires a phosphopantetheinylated enzyme. J. Biol. Chem. 282, 478–485 (2007).
Inoue, K., Makino, Y., Dairi, T. & Itoh, N. Gene cloning and expression of Leifsonia alcohol dehydrogenase (LSADH) involved in asymmetric hydrogen-transfer bioreduction to produce (R)-form chiral alcohols. Biosci. Biotechnol. Biochem. 70, 418–426 (2006).
Inoue, K., Makino, Y. & Itoh, N. Purification and characterization of a novel alcohol dehydrogenase from Leifsonia sp. Strain S749: a promising biocatalyst for an asymmetric hydrogen transfer bioreduction. Appl. Environ. Microbiol. 71, 3633–3641 (2005).
Jörnvall, H., Persson, B. & Jeffery, J. Characteristics of alcohol/polyol dehydrogenases. Eur. J. Biochem. 167, 195–201 (1987).
De Smidt, O., Du Preez, J. C. & Albertyn, J. The alcohol dehydrogenases of Saccharomyces cerevisiae: a comprehensive review. FEMS Yeast. Res. 8, 967–978 (2008).
Akhtar, M. K., Turner, N. J. & Jones, P. R. Carboxylic acid reductase is a versatile enzyme for the conversion of fatty acids into fuels and chemical commodities. Proc. Natl Acad. Sci. USA 110, 87–92 (2013).
Venkitasubramanian, P., Daniels, L., Das, S., Lamm, A. S. & Rosazza, J. P. N. Aldehyde oxidoreductase as a biocatalyst: reductions of vanillic acid. Enzyme. Microb. Technol. 42, 130–137 (2008).
Taguchi, S. et al. A microbial factory for lactate-based polyesters using a lactate-polymerizing enzyme. Proc. Natl Acad. Sci. 105, 17323–17327 (2008).
Lim, H. N., Lee, Y. & Hussein, R. Fundamental relationship between operon organization and gene expression. Proc. Natl Acad. Sci. 108, 10626–10631 (2011).
Tseng, H.-C., Martin, C. H., Nielsen, D. R. & Prather, K. L. J. Metabolic engineering of Escherichia coli for enhanced production of (R)- and (S)-3-hydroxybutyrate. Appl. Environ. Microbiol. 75, 3137–3145 (2009).
Cracan, V., Padovani, D. & Banerjee, R. IcmF is a fusion between the radical B12 enzyme isobutyryl-CoA mutase and its G-protein chaperone. J. Biol. Chem. 285, 655–666 (2010).
Howard, T. P. et al. Synthesis of customized petroleum-replica fuel molecules by targeted modification of free fatty acid pools in Escherichia coli. Proc. Natl Acad. Sci. USA 110, 7636–7641 (2013).
Rodríguez-Zavala, J. S., Allali-Hassani, A. & Weiner, H. Characterization of E. coli tetrameric aldehyde dehydrogenases with atypical properties compared to other aldehyde dehydrogenases. Protein Sci. 15, 1387–1396 (2006).
Jo, J.-E. et al. Cloning, expression, and characterization of an aldehyde dehydrogenase from Escherichia coli K-12 that utilizes 3-Hydroxypropionaldehyde as a substrate. Appl. Microbiol. Biotechnol. 81, 51–60 (2008).
Tseng, H.-C., Harwell, C., Martin, C. & Prather, K. Biosynthesis of chiral 3-hydroxyvalerate from single propionate-unrelated carbon sources in metabolically engineered E. coli. Microb. Cell. Fact. 9, 96 (2010).
Zor, T. & Selinger, Z. Linearization of the Bradford protein assay increases its sensitivity: theoretical and experimental studies. Anal. Biochem. 236, 302–308 (1996).
Acknowledgements
This research was supported in part by an award from the Department of Energy (DOE), Office of Science Graduate Fellowship Program (DOE SCGF). The DOE SCGF Program was made possible in part by the American Recovery and Reinvestment Act of 2009. The DOE SCGF program is administered by the Oak Ridge Institute for Science and Education for the DOE. ORISE is managed by Oak Ridge Associated Universities (ORAU) under DOE contract number DE-AC05-06OR23100. All opinions expressed in this paper are the author’s and do not necessarily reflect the policies and views of DOE, ORAU or ORISE. This research was also supported by the Institute for Collaborative Biotechnologies through grant W911NF-09-0001 from the US Army Research Office. The content of the information does not necessarily reflect the position or the policy of the Government, and no official endorsement should be inferred.
Author information
Authors and Affiliations
Contributions
M.J.S., A.M.K. and K.L.J.P designed the research. M.J.S., A.M.K. and S.J.W. performed all experiments. K.L.J.P. supervised the research. All authors wrote, reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.J.S., A.M.K. and K.L.J.P. are authors on a patent application entitled 'icrobial production of branched medium chain alcohols, such as 4-ethylpentanol' application number 61/899,129 (1 November 2013). The remaining authors declare no competing financial interests
Supplementary information
Supplementary Information
Supplementary Figures 1-8, Supplementary Tables 1-3, Supplementary Note 1, Supplementary Methods and Supplementary References (PDF 4835 kb)
Rights and permissions
About this article
Cite this article
Sheppard, M., Kunjapur, A., Wenck, S. et al. Retro-biosynthetic screening of a modular pathway design achieves selective route for microbial synthesis of 4-methyl-pentanol. Nat Commun 5, 5031 (2014). https://doi.org/10.1038/ncomms6031
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms6031
This article is cited by
-
Rebooting life: engineering non-natural nucleic acids, proteins and metabolites in microorganisms
Microbial Cell Factories (2022)
-
Efficient biosynthesis of cinnamyl alcohol by engineered Escherichia coli overexpressing carboxylic acid reductase in a biphasic system
Microbial Cell Factories (2020)
-
Engineering nature for gaseous hydrocarbon production
Microbial Cell Factories (2020)
-
Engineered microbial biofuel production and recovery under supercritical carbon dioxide
Nature Communications (2019)
-
Technical Advances to Accelerate Modular Type I Polyketide Synthase Engineering towards a Retro-biosynthetic Platform
Biotechnology and Bioprocess Engineering (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.