Abstract
Force sensing at cadherin-mediated adhesions is critical for their proper function. α-Catenin, which links cadherins to actomyosin, has a crucial role in this mechanosensing process. It has been hypothesized that force promotes vinculin binding, although this has never been demonstrated. X-ray structure further suggests that α-catenin adopts a stable auto-inhibitory conformation that makes the vinculin-binding site inaccessible. Here, by stretching single α-catenin molecules using magnetic tweezers, we show that the subdomains MI vinculin-binding domain (VBD) to MIII unfold in three characteristic steps: a reversible step at ~5 pN and two non-equilibrium steps at 10–15 pN. 5 pN unfolding forces trigger vinculin binding to the MI domain in a 1:1 ratio with nanomolar affinity, preventing MI domain refolding after force is released. Our findings demonstrate that physiologically relevant forces reversibly unfurl α-catenin, activating vinculin binding, which then stabilizes α-catenin in its open conformation, transforming force into a sustainable biochemical signal.
Similar content being viewed by others
Introduction
Cell–matrix and cell–cell adhesions are required in morphogenesis during embryogenesis, tissue development during fetal life, as well as tissue maintenance during adulthood1. In addition to mere cell membrane adhesion, fine-tuning of transmission of mechanical load from cell to extracellular matrix (ECM) and cell to cell is also essential to these processes2,3. The molecular mechanisms underlying cell–ECM mechanosensing processes have been partly unraveled. Although cell–ECM mechanotransduction may rely on more global adaptation of the actomyosin viscoelastic networks4 and activation of mechanosensitive channels5, pioneering works have demonstrated the existence of integrin-associated cytoplasmic proteins with buried sites of phosphorylation such as p130Cas6, and of protein–protein interactions such as talin7,8 that are unmasked upon myosin II-dependent stretching. The tension-dependent conformation switch of these proteins may thus initiate the force-dependent building of adaptor complexes linking cell–matrix adhesions to the tension-generating actomyosin network.
By analogy, cadherin-associated adhesion complexes might have an essential role in transducing forces at cell–cell junctions9,10. These complexes are tension adaptive, actin-cytoskeleton-associated structures, responsive to both external load and tensile force produced by intracellular myosin motors11,12. The mechanism of mechanosensing at cell–cell contacts has only been very recently investigated13,14, and α-catenin appears as a central component of the force transmission pathway.
The αE isoform of α-catenin is expressed ubiquitously in early embryonic cells, and then restricted to epithelia. Its deletion is associated with impaired cadherin-mediated adhesion15,16, tissue growth, and homeostasis17,18,19. It has been recently hypothesized that αE-catenin may act as a mechanotransducer in the pathway that converts mechanical strain on cadherin adhesions into a cue for junction strengthening11. Because vinculin accumulates at mature cell–cell junctions upon actomyosin generated tension11,20,21,22 and binds αE-catenin23,24,25, it has been proposed that α-catenin functions in concert with vinculin. Further analysis of cadherin adhesion strengthening by cell doublet force separation measurement indicates that α-catenin, vinculin and their direct interaction are required for tension-dependent intercellular junction strengthening26. These proteins appear as key candidates for mechanotransduction at cell–cell junctions.
Vinculin is a cytoplasmic actin-binding protein enriched in both focal adhesions and adherens junctions, essential for embryonic development27. At focal adhesions, vinculin has a critical function in linking integrins to F-actin. Vinculin is a compact globular protein composed of successive four α-helix bundles. Five of these α-helix bundles constitute the vinculin head binding to various partners such as talin, whereas the C-terminal constitutes the vinculin tail binding to F-actin. In the cytosol, vinculin is under an inactive head to tail conformation presenting only week affinity for actin. In contrast, vinculin captured at focal adhesions by force-dependent activated talin is stabilized under an open conformation characterized by head to tail dissociation, stabilized by binding of the head to talin and high affinity binding of the tail to F-actin28.
α-Catenin is a complex protein with strong homology with the vinculin head domain, sharing a λ-shape arrangement of α-helix bundles29. At cell–cell junctions, β-catenin directly binds to the N-terminus of α-catenin30,31 and to the intracellular tail of cadherins32,33, forming the cadherin/β/α-catenin complex. α-Catenin possesses a domain of homodimerization and dimerizes in solution (Fig. 1a: DD domain); however, this domain overlaps with a N-terminal β-catenin-binding domain, and homodimerization of α-catenin is inhibited by β-catenin binding29,34. The C-terminus of α-catenin contains an F-actin-binding site25,35, which associates the tertiary cadherin/β-catenin/α-catenin complex to the actin filaments36. Although direct binding has not been observed between the purified components of this complex in solution37, it is still acknowledged that α-catenin dynamically links the complex to F-actin directly, indirectly or both, allowing force transduction and strengthening of adhesions13,26,38. α-Catenin binds to other actin-binding proteins, such as vinculin18,24,25,39, ZO-1 (refs 25, 40), afadin41 and formin-1 (ref. 42)42, through sites distributed in the central part of the molecule. The actin-binding domain of α-catenin located at the C-terminus of the molecule (Fig. 1a: FABD domain) appears to bind to the side of actin filaments, inducing conformational changes of individual filaments and preventing the binding of the branching complex Arp2/3 and the severing protein cofilin43. Thus, α-catenin binding to actin may favour assembly of unbranched filaments that are more protected from severing than dynamic, branched filament arrays44.
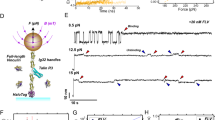
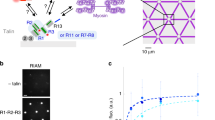
(a) Domain mapping of full-length and sub-domains of mouse αE-catenin. αE-catenin consists of an N-terminal dimerization domain (DD), followed by vinculin-binding domain (VBD; also referred to the MI domain). Interaction between VBD and two other modulation domains (MII-MIII) is suggested to inhibit vinculin binding. αE-catenin contains a C-terminal F-actin-binding domain (FABD) that binds to F-actin and transmits mechanical forces generated by actomyosin contraction. The αCM construct (residues 275–735) is enclosed in the top bracket. (b) Schematics of experimental setup. A single αCM molecule is tethered between a cover glass surface and a paramagnetic bead through NTA-His tag and streptavidin-biotin linkages, respectively. The modulatory MI (VBD) domain is shown in green, MII and MIII domains in yellow. The vinculin-binding helix is marked in red. The VD1 molecule, which is also a helix bundle, is shown in silver. Force was applied to the paramagnetic bead using a pair of permanent magnets. αCM constructs were stretched in the absence or in the presence of VD1 of various concentrations. The model of αCM is adopted from PDB structure 4IGG.
The α-catenin central domain (αCM, Fig. 1a: VBD+M domains) is an adhesion modulation domain25,45, composed of a vinculin-binding domain MI or VBD23,46, followed by the so-called modulation domains MII and MIII (ref. 47). Biochemical and structural data suggest that the vinculin helix bundle (VD1) forming the vinculin head binds to a single vinculin-binding α-helix (residues 305–355) within the α-catenin VBD domain39,46. Structural data further indicate that α-catenin can adopt an auto-inhibitory conformation48 whereby the VBD domain is interacting with both MII and MIII helix bundles, thus preventing accessibility of the vinculin-binding domain. Vinculin may only bind to α-catenin when the VBD-MII-MIII helix bundles are in an open conformation46,47,48. Altogether, these data suggest that α-catenin may unfurl in a tension-dependent manner, allowing the recruitment of vinculin, and further strengthening of the cadherin complex/F-actin linkage. This pathway may be responsible for the mechanosensitive maturation of intercellular junctions. However, crucial experiments are lacking in support of this hypothesis. No direct evidence demonstrates the force-dependent unfurling exposing the VBD α-helix and the binding of α-catenin to vinculin under force.
To investigate the role of α-catenin in force transduction, we use single-molecule assays that allow us to obtain direct experimental evidence that physiological-range forces regulate α-catenin conformation and α-catenin–vinculin interaction. Thus, we provide here a molecular mechanism by which forces are locally transformed into biochemical signal at cadherin-mediated adhesions, which has significant implications for adherens junction assembly and regulation.
Results
αCM unfolds in three characteristic steps under force
To experimentally test the hypothesis that the modulation domain of αE-catenin (αCM) containing the VBD-MII-MIII helix bundles can unfurl upon application of physiological-range forces39, we first investigated the force response of αCM domain of mouse αE-catenin (275–735), produced with a biotin tag at the N-terminal and a 6-His tag at the C-terminal (Fig. 1a), thanks to a stable high-force magnetic tweezers instrument49,50,51. For this set of experiments, we specifically tethered single αCM molecules between an NTA-Cu2+ functionalized coverslip and a streptavidin-coated paramagnetic bead and subjected the tethered molecules to forces exerted by a pair of permanent magnets (Fig. 1b). We recorded the extension change of the αCM molecule based on the diffraction pattern of the bead at a sampling rate of 200 Hz with nanometer resolution49.
To obtain a mechanical characterization of the molecule, single αCM tethers were subjected to force scanning cycles. In each cycle, force was increased at a constant loading rate of 4 pN s−1 from ~1 pN to ~40 pN, which was followed with a force-decrease step, reducing force exponentially with time back to <1 pN that minimizes duration of the tether being held at large forces (Fig. 2a). The chosen loading rate of 4 pN s−1 is within a physiologically relevant range, as recently estimated for integrin stretching during cell retraction52. During the force-increase step, we identified unfolding events by sudden increases in the extension of the αCM tether, whereas we also identified refolding events during the force-decrease step by sudden decreases in the extension. To allow refolding of the αCM molecule following the force-decrease step, we held the αCM tether at<1 pN for 1 min. During this period of time, the refolding of αCM appeared taking place with a >90% probability (Supplementary Fig. 1a). The unfolding and refolding force-responses that we observed indicate that αCM behaves like a nonlinear spring, which may be important for its potential in vivo function as a mechanosensor.
(a) Extension change of αCM recorded at 200 Hz in a typical force cycle experiment. During the stretch phase (force-increase) at a loading rate of 4 pN s−1, an unfolding step was observed at ~5 pN (blue box), and two additional ones occurred at higher forces (~15 pN; red box). Two refolding events, one at ~5 pN with>20 nm and another at ~2 pN with >10 nm, were observed during relaxation (force-decrease). The inset shows that the extension at ~5 pN is reversible. (b) Four force-extension curves recorded in four sequential stretch-relax cycles. The stretch curves are indicated in colour, whereas the relax curves are indicated in grey scale. Unfolding steps are marked by up arrows and refolding steps are marked by down arrows. Data in b are smoothed using 0.05-s time intervals. (c) Stepwise extension fluctuation at constant forces. Raw data are shown in red, and the black lines are fitted states using Hidden Markov method (Methods: ‘Determination of transition rates at constant forces’). (d) Typical force-extension curves obtained from four force cycles for L344P αCM mutant on a single tether. The red up arrow marks the missing ~5 pN unfolding step.
In a typical force-increase scan, we found that the αCM unfolded in three characteristic steps (Fig. 2a). Because both extension and force at any time were known, the time traces of extension and force could be converted into force-extension curves (Fig. 2b). Representative unfolding and refolding force-extension curves obtained during successive force cycles on a single αCM tether shows remarkable reproducibility (Fig. 2b). We confirmed this reproducibility on multiple (>10) independent αCM molecules (Supplementary Fig. 1b for another example).
At a constant loading rate of 4 pN s−1, the first step occurred at 5.2±0.7 pN (mean±s.d.) and involved a reversible extension change of 16.3±4 nm (mean±s.d.) between two conformational states, which can be understood by unfolding the bundled α-helices in the VBD domain into a linear chain of extended α-helices as discussed in the Discussion section. The inset in Fig. 2a clearly shows that the extension of the tether fluctuates at forces ~5 pN indicating that the transition of the corresponding domain is near equilibrium when the αCM tether is subjected to this force magnitude.
The two higher force steps were overlapping at about 12.5±2.0 pN and had similar step sizes of 23.3±6.6 nm corresponding to 105±30 residues. These unfolding sizes fit the domain sizes of αCM (VBD: 277–393, MII-MIII: 396–631, Fig. 1a)25,45. As the unfolding forces of the two higher force steps were close to each other, they could combine into a single large step of ~50 nm. During the following force-decrease scan, we observed two apparent refolding steps at low forces, one at ~5 pN, and another at ~2 pN (Fig. 2a).
The co-existence of unfolding and refolding steps of same sizes around 5 pN during the stretching process suggests that it corresponds to a near-equilibrium transition operating in this range of forces (Supplementary Fig. 1c). We were particularly interested in this transition because its force dependence is close to the range of forces generated by a single myosin motor53. If this transition is near-equilibrium in this range of forces it should be largely insensitive to loading rate. To further demonstrate the reversibility of this transition, constant force experiments were carried out in the 4–5 pN range (Fig. 2c). Slight increase in force from ~4.1 to ~4.8 pN switched the predominant folded state to the predominant unfolded state. Analysis of the dwell times of each state determined the force-dependent unfolding and refolding transition rates and a critical force of ~4.7 pN (Supplementary Fig. 1d). These results confirmed that the ~5 pN transition was insensitive to loading rate in the range tested and reversible around a force of 5 pN, which may be important for physiological myosin-driven conformational switch of α-catenin.
Altogether, the highly reproducible force-extension curve we observed characterizes the mechanical response of αCM, and can serve as the basis for further analysis of the force-dependent interaction between αCM and the VD1 domain of the vinculin head.
~5 pN force unfurls αCM exposing the vinculin-binding site
To determine whether the αCM near-equilibrium unfolding at ~5 pN corresponds to the unfurling of the VBD domain allowing the stabilization of the VD1-binding α-helix predicted by structural data39,48, we performed glutathione S-transferase (GST) pull-down experiments. WT αCM proteins were specifically pulled down with GST-VD1 (Supplementary Fig. 2a), as expected from the presence of the VBD domain. To disrupt the VD1-binding α-helix, we introduced a leucine to proline point mutation in the middle of the α-helix (construct L344P αCM)39. As previously reported39, this mutation inhibited the binding of αCM to GST-VD1 (Supplementary Fig. 2a). We repeated the force-cycling experiments on L344P αCM. The 5 pN reversible unfolding/refolding step was not observed with the mutant protein (Fig. 2d and Supplementary Fig. 2b,c) indicating that the VD1-binding α-helix was required for proper unfolding/refolding of αCM at ~5 pN. It further suggested that the transition at ~5 pN involves the unfurling of the VBD domain allowing the formation of the vinculin-binding α-helix.
We then determined whether αCM mechanical stretching influences VD1 binding. To do so, we counted the number of photobleaching events of Alexa 488-labelled VD1 bound to αCM maintained or not under force. Alexa 488 was conjugated to VD1 following the VD1 labelling protocol used in a previous study of talin-VD1 binding7 (Methods: ‘Photobleaching counting’). Three typical time trajectories of the fluorescence intensity with zero, one and three photobleaching events, respectively, are presented in Fig. 3a. The number of photobleaching events was very low when no force was applied and increased fivefold when force was increased from 0 to 20 pN (Fig. 3b, upper panel). Significantly, more tethers showed one photobleaching event at 20 pN force compared with that at 0 pN force. This result is consistent with previous reports suggesting the binding of only one vinculin-binding α-helix per α-catenin central domain24. We performed similar experiments with a construct containing three αCM domains. Consistently, using this artificial 3x WT αCM molecule-bearing three potential vinculin-binding sites, we also observed an increase of photobleaching events when forces were increased from 0 to 20 pN, with a number of photobleaching events reaching three at 20 pN (Fig. 3b). The rare cases where we observed more than one photobleaching event on WT αCM, or more than three photobleaching events on 3x WT, were likely due to low probability of double or multiple coupling of the Alexa 488 dyes on some VD1 molecules. Altogether these results demonstrate that the mechanical stretching of αCM strongly activates the binding of a single VD1 per VBD domain.
(a) Three representative time-lapse traces of the fluorescent intensity measured below individual tethered beads: 0 (WT αCM, no VD1 in solution), 1 (WT αCM, VD1 in solution) and 3 photobleaching events (3x WT αCM, VD1 in solution) were shown for each case. Dashed lines are drawn for visual guidance. (b) Histograms of the percentage of beads with different numbers of photobleaching events observed for WT αCM and 3x WT αCM constructs. The number of individual tethered beads tested for each condition is denoted by ‘n’ in figure panel. The P-value was obtained from the χ2 test with the null hypothesis that the photobleaching statistics is independent on whether force is applied.
VD1 binds stretched αCM and inhibits MI domain refolding
To correlate the force-dependent binding of VD1 to one of the specific unfolding steps of the αCM molecule described above, we investigated the effect of VD1 on the force responses of αCM. We carried out force cycle experiments in the presence of VD1 in the solution. In a typical force cycle in the presence of 10 nM VD1, the ~5 pN unfolding step disappeared in the force-increase phase, whereas the two unfolding steps at 10–18 pN remained (Fig. 4a). In the force-decrease phase, we observed a small (~2.5 nm) step at ~9 pN, which was not observed in the absence of VD1, followed by two larger (>15 nm) steps at lower forces also observed in the absence of VD1.
(a) Force-extension curve of αCM in a typical stretch-relax force cycle in the presence of 10 nM VD1. The inset shows a ~3 nm VD1 dissociation unfolding step at ~37 pN. (b) Unfolding force histogram obtained from repeating stretch-relax force cycles on a single αCM at different VD1 concentrations. (c) Percentages of observed unfolding events at <7.5 pN and >7.5 pN unfolding cycles at different VD1 concentrations. The significance indicated by the asterisks is based on P-value calculation from the χ2 test with the null hypothesis that the percentages of the unfolding events are independent on the VD1 concentration. Four asterisks denote a P-value smaller than 0.0001. The data were collected from eight independent tethers. The error bar denotes 95% confidence interval of probability estimation. (d) Three force-extension curves recorded during three sequential stretch processes between 1 and 9 pN in the presence of 0 nM VD1 (coloured traces), where only the ~5 pN unfolding/refolding events occurred. After each stretch, force was jumped to 0.1 pN for 1 min for refolding before the next stretch was performed in the presence of 10 nM VD1 (black and grey traces). The ~5 pN unfolding/refolding event was observed during the first stretch cycle (black), but was lost in the subsequent three stretches. Data in d are smoothed using 0.05-s time interval.
Figure 4b shows unfolding force distributions normalized by the number of force cycles at different VD1 concentrations recorded during force-increase phases on a single tether with more than 30 unfolding events for each condition, which clearly reveals that the frequency of the ~5 pN transition decreases as the VD1 concentration increases. Figure 4c summarizes results from eight different tethers recorded in the absence or in the presence of VD1 at 1 and 10 nM concentrations. The ~5 pN unfolding events were significantly reduced in the presence of 1 nM VD1, and nearly disappeared in the presence of 10 nM VD1. Similar results were obtained at higher VD1 concentrations (Supplementary Fig. 3).
Interestingly, we found an additional unfolding step at >35 pN associated with a very small step size of ~3.0 nm appearing in the presence of VD1 in solution, which was not observed in the absence of VD1 (Fig. 4a). Considering that high forces could favour a transition of the vinculin-binding α-helix from the VD1-bound state to a VD1-unbound, extended random coiled peptide chain state, this step could be interpreted as the dissociation of VD1 bound to the vinculin-binding α-helix. Therefore, at a sufficiently high force, VD1 dissociation should occur, accompanied by a small unfolding step during the subsequent helix-to-coil transition of the vinculin-binding α-helix. The ~3 nm unfolding step size observed at ~39 pN was consistent with the estimated extension increase during helix-to-coil transition of the vinculin-binding α-helix at 30–40 pN (Methods: ‘Helix-to-coil transition’).
The fact that we observed only one ~3 nm unfolding step at high forces again strongly indicated that a single VD1-binding helix exists in the αCM construct, which was consistent with our photobleaching experiments (Fig. 3). Similarly, the ~2.5 nm refolding step at ~9 pN that we only observed in the presence of VD1 may be due to the re-binding of VD1 to the unfolded vinculin-binding α-helix of αCM, which induces formation of the α-helix structure of the vinculin-binding site. Consistently, when experiments were performed, at a lower concentration of VD1 (1 nM) to reduce the chance of re-binding during force decrease, we observed that displacement of the bound VD1 at high force could restore the ~5 pN transitions (Supplementary Fig. 3c),
To investigate whether activation of VD1 binding to αCM requires or not complete unfolding of αCM, we performed force-cycling experiments up to 9 pN at a loading rate of 4 pN s−1. We selected this maximal force because it only allowed unfolding of the weakest domain at ~5 pN. We found that unfolding the weakest domain was sufficient to activate VD1 binding that abolished the ~5 pN unfolding/refolding events (Fig. 4d). The results indicated that unfolding the VBD domain alone was sufficient to activate VD1 binding. Taken together, these results suggested that the vinculin-binding α-helix in αCM could be reversibly exposed by mechanical force of ~5 pN. In addition, they indicated that VD1 binding to the exposed vinculin-binding α-helix inhibited αCM refolding at low force, and that the bound VD1 could be displaced at large forces.
VD1 binds to and locks αCM in its open conformation
To evaluate to which extent mechanical force increased the affinity of VD1 binding to αCM, we incubated αCM tethers in the presence of 10–100 nM VD1 at low force (1–2 pN) for 10 min before the first stretching. Should VD1 spontaneously bind to the αCM tether at low force during the time, we would expect to observe a loss of the ~5 pN unfolding/refolding step during the first force-increase scan. In three out of four independent experiments, we observed the ~5 pN transition and the two higher force unfolding steps in the first force-increase scan (Fig. 5a, and Supplementary Fig. 4a,b), indicating that no VD1 binding had occurred within 10 min incubation at ~2 pN. In addition, we found that the ~5 pN transition was lost in the subsequent force cycles, whereas the two higher force species remained. These results were consistent with VD1 binding occurring after the first stretching procedure, which locked the vinculin-binding α-helix of the MI domain in an open state. In the fourth experiment at 100 nM VD1, the ~5 pN transition step disappeared in the first force-increase scan, indicating VD1 binding before the force-increase scan. A close examination of the extension time trace revealed that a spontaneous binding of VD1 occurred at ~2 pN at minutes after we introduced VD1 solution, as indicated by an abrupt extension increase (Fig. 5b). As VD1 binds to unfolded αCM in seconds under >5 pN forces, these results indicated that mechanical forces above 5 pN strongly promoted VD1 binding to αCM.
(a) Extension change of αCM in multiple stretch-relax force cycles in the presence of 10 nM VD1. The VD1 solution was slowly introduced at a low force (~1 pN) to avoid αCM unfolding by fluid-drag force. After the VD1 solution was introduced, the αCM molecule was held at ~2 pN for 10 min in the VD1 solutions before being stretched for the first time. Data were smoothed with 0.05-s time window. (b) In one out of four experiments after introduction of 100 nM VD1 solution at 2 pN, spontaneous binding of VD1 occurred indicated by a stepwise unfolding event. For clarity, data are smoothed using 0.5-s time interval.
VD1 bound to exposed VBD has a slow off-rate at low force
Then, to quantify the stability of VD1 association to mechanically exposed VD1-binding α-helix under force, we carried out force-jump experiments in which force was alternated between 8 pN to promote rapid VD1 binding, and a high force to dissociate the VD1 from the VD1-binding α-helix. We found that a single VD1 dissociation event, indicated by a ~3-nm unfolding step, occurred within 10 s after force was switched to ~40 pN (Fig. 6a), which was not observed for the L344P mutant (Fig. 6b).
(a) Extension change of WT αCM during force jump between ~8 and ~40 pN in the presence of 100 nM VD1. (b) Extension change of L344P αCM during the same force jump procedure. (c) The histogram of lifetime of VD1 bound to αCM at different forces. Red curves are fittings with exponential decay function from which the dissociation rates were obtained. The number of events observed in each condition is denoted by ‘n’ in figure panel. The data were obtained from three individual tethers. (d) The logarithm of the dissociation rate is roughly linearly dependent on force. Red line denotes linear fitting of the log-scaled force-dependent off-rate. From the slope, the transition distance is estimated to be ~1 nm, and the dissociation rate at no force is estimated to be 9.3 × 10−6 s−1 by extrapolation based on the Bell’s model69. Y error bars denote 95% confidence interval of off-rate estimation, X error bars denote the ~5% relative error of our force calibration method.
To quantify the force-dependent off-rate of VD1 bound to αCM, we repeated such force-jump for many cycles on native αCM at a variety of forces. The cumulative dissociation time distribution (the percentage of vinculin dissociated at time t) at various high forces from three independent tethers was obtained (Fig. 6c). At each force, the time window of observation was chosen so that in more than 90% of the force-jump cycles vinculin dissociation events were observed. The cumulative dissociation distributions were fitted by an exponential decay function 1−exp(−tKoff), where the parameter Koff characterizes the rate of dissociation at force f. We found that the logarithm of Koff(f) roughly linearly depended on force (Fig. 6d), from which the off-rate at low forces could be estimated by linear extrapolation. At <10 pN, the extrapolated off-rate is <10−5 s−1. Such an ultraslow dissociation rate of VD1 from exposed VD1-binding α-helix may explain why the bound VD1 could inhibit refolding of αCM. Altogether, these data indicated a strong stability of the vinculin/α-catenin complex with a rapid formation and slow dissociation in the tension range 5–30 pN.
Discussion
We report here on the mechanical response of the central domain of αE-catenin (αCM) and its force-dependent binding to the vinculin head (VD1), characterized at the single-molecule level, in a physiologically relevant range of forces. Our results show that αCM displays three major unfolding steps, occurring at different forces in the range of 5–20 pN under a physiologically loading rate of 4 pN s−1. We identified the MI/VBD domain of α-catenin, which contains the VD1-binding site made of a single α-helix, as the weakest domain that unfolded at ~5 pN. This unfolding step was lost when a single point mutation (L344P) is introduced in the α-helix39, as well as when VD1 is bound to αCM.
The two larger force-unfolding events are unrelated to binding of VD1, and are attributed to the unfolding of the sub-domains MII and MIII also constituted of helix bundles but not predicted to contain vinculin-binding sites. Structural data suggest that in the folded state, MI/VBD interacts with both MII and MIII (ref. 47); therefore this interaction may stabilize VBD in its closed conformation, as well as contributing to the stabilization of the MI-MIII subdomains. Our results are consistent with a hypothesis that the interaction of the three sub-domains contributes to the stabilization of α-catenin under its close conformation, whereas a moderate 5 pN stretching force disrupt this inter-domain interaction resulting in exposing the vinculin-binding site in the MI/VBD domain.
The ~5 pN unfolding and folding transitions are nearly at equilibrium within the loading rates tested in the experiments. These transitions involved a step size of ~16 nm with a critical force of ~4.7 pN at which folding and unfolding are balanced. This transition cannot be explained by the disruption of the inter-domain interactions alone; otherwise much smaller transition steps would be expected as each subdomain has a dimension of only ~3 nm in the folded state. Together with data obtained from L344P mutation and VD1-binding studies, the ~16 nm step involved in this transition is consistent with a picture that the disruption of the inter-domain interactions at ~5 pN leads to a subsequent unfolding of the MI/VBD α-helix bundle. The extension of an unfolded peptide in random coil and in α-helix conformations is similar, ~17 nm estimated for a chain of extended helices with total ~32 helical turns of 116 residues in the MI domain, and ~15 nm of the same number of residues in the random coil conformation at ~5 pN estimated based on the worm-like chain polymer model with a bending persistence length of ~0.5 nm (reported from 0.4–0.6 nm)54,55,56,57. Considering that VD1-binding occurred at this force range, we reason that the unbundled α-helices in the MI domain at ~5 pN likely assumed a conformation as a chain of extended α-helices rather than a randomly coiled peptide chain.
We then demonstrated and characterized in details the kinetics of the force-dependent binding of the VD1 to αCM. Indeed, our results revealed that a characteristic force response of αCM controlled the level of VD1 binding. αCM unfolding at ~5 pN was sufficient to trigger VD1 binding at a nanomolar concentration, which prevented refolding of αCM VBD even after force was released. This binding was observed in the presence of VD1 at concentrations as low as 1 nM and was maximum in the presence of 10 nM VD1, implying a strong affinity of VD1 to αCM VBD triggered by >5 pN force. Overall, the effect of VD1 binding to the vinculin-binding α-helix in αCM on the characteristic force response of αCM is similar to the leucine to proline mutation at the residue 344 of αCM, both abolishing the ~5 pN unfolding/refolding transitions. These results are also consistent with our observation that VD1 did not bind to L344P αCM in in-vitro pull-down assays.
To estimate the increase in the binding affinity of VD1 to αCM by force, we used results from published isothermal titration calorimetry studies, which indicate a KD of 82±19 nM for αCM binding to VD1 at zero force58. Based on these calculations, and considering that the apparent VD1 binding to mechanically stretched αCM at ~5 pN was observed at ~1 nM VD1 and maximum at ~10 nM VD1, we estimated that a force of several piconewton increased the binding affinity of VD1 to αCM by about 100-folds, which may be sufficient to activate auto-inhibited vinculin by competing off its head-to-tail association59. Overall, these data provide molecular evidences in strong support of α-catenin as being the force-dependent molecular switch acting at cell–cell junctions to recruit vinculin (Fig. 7), at the basis of the reported tension-dependent growth and adaptation of these junctions12,22,26,60.
At low force (<5 pN), αCM exists under an auto-inhibited helix bundle conformation that prevents VD1 binding. At high force (>30 pN), VD1 binding is also inhibited because the α-helix conformation of the vinculin-binding site bound with VD1 is unstable. In the intermediate force range (5–30 pN), the auto-inhibited conformation of αCM is released, and the vinculin-binding site is exposed for VD1 binding.
Our results revealed that vinculin head binding is biphasically dependent on force. In the small force range (<5 pN) where αCM existed as natively folded, auto-inhibitory helical bundles, and in the high force range (>30 pN) where the VD1-binding site was converted from its α-helix conformation to an extended random coiled peptide chain, binding of VD1 was strongly inhibited. Between 5 and 30 pN, binding of VD1 was significantly promoted. As 5 pN is close to the force that can be exerted by a single myosin motor53, the low-force regime is likely involved during early adhesion formation with low myosin activity. The high-force VD1 dissociation is also physiologically relevant, as a 30-pN force can be produced by several cooperative myosins or external forces exerted on cells. Ultimately, locking α-catenin in a partially unfolded conformation may regulate cellular adhesion strengthening in a force-dependent manner. Similarly, high forces achieved upon mobilization of multiple myosin molecules pulling on a single αE-catenin molecule via F-actin may eventually release vinculin at >30 pN and then lead to the disruption of the mechanosensitive link61. This may explain the disruption of cadherin adhesions observed upon increased cell contraction obtained by increasing RhoA activity62. To elucidate these mechanisms, further experiments in cellulo will be required in the future.
The force-dependent process centered on α-catenin we unraveled here may further cooperate with the binding of other α-catenin partners to regulate the strength of the cytoskeleton linkage to cell–cell junctions. A ~5 pN force deforms the modulation domain of α-catenin and initializes binding of VD1. The resulting lockage of α-catenin in a partially unfolded conformation may have a downstream impact on interactions between α-catenin and other junctional and cytoskeleton proteins, such as α-catenin itself, afadin, ZO-1, α-actinin and F-actin. However, further understanding of these processes will require precise structural and biochemical data on these complexes that are lacking so far. Up to date, a complete structural description of αE-catenin under both its closed and open conformations is still lacking.
The exact sequential mechanistic pathway by which α-catenin and vinculin bind together at cell–cell contacts has been highly debated within the field. Bakolitsa et al. proposed that αE-catenin VBD binding to vinculin can activate vinculin58; however, previous data suggested that αE-catenin lacking the actin-binding domain, or the VBD domain alone, bound poorly to full-length vinculin in solution unless F-actin is present in the solution39,46. These results suggested that αE-catenin or the VBD domain alone were not sufficient to activate vinculin by direct interaction with the vinculin head. Nonetheless, as αE-catenin and the VBD domain were not placed under force in those experiments, those results do not rule out the possibility that αE-catenin alone can directly activate vinculin when it is placed under force. Our results demonstrated that VD1 bound to mechanically exposed VBD in αCM under 5–30 pN forces with a nanomolar dissociation constant. This is significantly lower than the KD=~50–90 nM determined for the head-to-tail association of vinculin59,63. Such a strong affinity between VD1 and mechanically stretched αE-catenin suggests that it may be able to compete off the vinculin head-to-tail association of auto-inhibited vinculin, which warrants future studies. In in vivo conditions, such stably associated activated vinculin may then bind itself to other molecular partners of the junctional complex, such as of F-actin, β-catenin64 and PtdIns (4,5)P2 (ref. 58), further contributing to the tension-dependent maintenance, strengthening and force-adaptation of cell–cell junctions.
To our knowledge, this work provides the first molecular demonstration of a tension-dependent conformational switch for a cadherin-associated protein. It provides a molecular mechanism explaining the local mechanosensitivity of cell–cell junctions, based on tension-dependent unfurling of α-catenin and recruitment of vinculin, a pathway that has been overwhelmingly hypothesized but surprisingly never proven so far. Further studies will be needed to determine whether the pathway is unique at cell–cell junctions or cooperate with similarly operating molecular mechanosensors and/or with more globally distributed mechano-adaptation of the actomyosin cytoskeleton4. Together with previous studies on force-dependent binding of vinculin to talin at integrin-mediated cell–matrix adhesions7,8, this work indicates that tension-dependent unfurling of mechanosensitive proteins and recruitment of actin-binding adaptor proteins to adhesion plaques is a central mode of mechanotransduction at adhesion sites. Further dissection of cell-context modulation of this pathway has broad implications for the understanding of cell adaptation to cell–ECM and cell–cell transduced mechanical load, an important factor of tissue reshaping, tumour progression and collective cell migration.
Methods
Protein expression and purification
A PCR product of the mouse αE-catenin central domain (αCM: residues: 275–735) was cloned into the di-cistronic pDW363 vector, co-expressing the Escherichia coli biotin holoenzyme synthetase (Bir A)65. Briefly, the 822–2266 nucleotide fragment of the coding sequence of the mouse αE-catenin cDNA (NM_009818) was amplified by PCR and cloned in phase between the XhoI and BamHI sites in N-terminal of the lysine-containing sequence recognized by Bir A, replacing the MalE-coding sequence. A 6His-coding sequence followed by a stop codon was added in C-terminal of the αE-catenin sequences. Unique KpnI and ApaI sites were introduced on both sides of the recombinant αE-catenin, thanks to the following forward: 5′-CTGGTGGCTCGAGCGGTACCGGCGGAGAGCTGGCATACGCT-3′, and reverse: 5′-ATGACCGACTTCACCCGAGGCAAAGGGCCCGGGGCCGGGCATCATCACCATCACCATTGAGGATCCATCATC-3′ primers. We then derived by PCR from this WT Biot-αEcat-6His construct, designated WT αCM throughout this study, a L344P Biot-αEcat-6His named thereafter L334P αCM bearing the L344P point mutation in the VBD domain reported previously to impair vinculin binding39. A 3 × WT αCM biotin and 6His-tagged construct, bearing three repeats of the KpnI-ApaI fragment, was also constructed in the pDW363 vector by combination of PCR amplification and In-Fusion recombination (Clontech). All constructs were validated by full sequencing of the open reading frame. Recombinant Biot-6His fusion proteins were expressed in E. coli BL21 strain and purified using Protino Ni-TED columns (Macherey-Nagel). Protein purity was evaluated by SDS–polyacrylamide gel electrophoresis electrophoresis and protein concentration measured using absorbance at 280 nm. The VD1 (residues: 1–258) construct was expressed as described in previous studies7.
GST pull-down and western blotting
A measure of 10 μg of Biot-αE-cat-6His (WT αCM or L334P αCM) and 4 μg of GST-VD1 (or GST) were mixed gently with 10 μl of glutathione-sepharose beads in PBS (final volume, 200 μl) for 30 min at room temperature. The beads were washed three times with PBS, then centrifuged for 2 min at 4 °C and incubated in 50 μl of 20 mM reduced glutathione, 50 mM Tris-HCl buffer pH 8 for 10 min at room temperature. GST-VD1 (or GST) with associated αE-catenin were recovered in the supernatant after centrifugation. Proteins were resolved on 10% polyacrylamide gels (Invitrogen) by conventional SDS–polyacrylamide gel electrophoresis and Coomassie blue staining.
Single-molecule manipulation
The single-molecule α-catenin pulling experiments were done on a home-made high-force magnetic tweezers setup with laminar flow channels with a NTA-Cu2+ functionalized coverslip for specific immobilization of αCM. The force calibration was done for each individual tethered bead measured in experiments, which had an ~5% relative error. The details of the design and control of the magnetic tweezers setup were published previously49. Protocols of coverslip functionalization, sample preparation, single-protein stretching experiments and force calibration have been published in our previous studies of Filamin A protein50,51.
Briefly, the channels were prepared by the following procedure: 20 × 32 mm2 1.5# coverslips were cleaned by ultrasonication in detergent, acetone and isopropanol for 30 min each, followed by 15 min of oxygen plasma treatment. The cleaned coverslips were silanized by incubation in 1% APTES (Sigma) in methanol for 20 min, rinsed thoroughly with methanol and cured for 20 min in oven at 110 °C. A laminar flow channel was made by sandwiching the silanized coverslip and a 20 × 20 mm2 clean non-functionalized coverslip with two parallel stripes of double-sided tape as spacer. The laminar flow channel was further treated by 0.5% glutaraldehyde for 1 h followed by incubation with 3 μm diameter amine-coated polysterene beads (polysciences) for 20 min and then with 10 μg ml−1 NH2-NTA for 6 h. The channel was then treated with 500 mM Tris pH 7.4 for 20 min and 0.04% (w/v) CuSO4 for 10 min. The Cu2+ charged channels were then blocked in PBS, 1% BSA, 0.0002% Tween-20 overnight before stretching experiment.
6His and biotin double-tagged αCM molecule (0.002 mg ml−1) were introduced in the channel and incubated for 20 min before introduction of 2.8 μm diameter streptavidin-coated paramagnetic beads (M270 streptavidin, Dynabeads) to form tethers. A syringe pump was used for slow buffer exchange (~5 μl min−1). The drag force applied to αCM during buffer exchange was estimated to be less than 1 pN. The pulling experiments were done in 40 mM phosphate buffer pH 7.4, 300 mM NaCl, 5 mM β-mercaptoethanol and 10 mg ml−1 BSA at 22 °C with various VD1 concentrations.
Raw extension data were recorded at a sampling rate of 200 Hz. In Figs 2b,d, 4a,d and 5a,b, those are smoothed using the fast Fourier transform (FFT)-smooth function of OriginPro 9.0 in order to improve the clarity of data representation. The unfolding steps of the molecule were detected using an in-house written Matlab programme. Briefly, a delta-function was calculated from the raw time lapse data. Candidate unfolding step positions were picked by peaks in the delta-function with height above a threshold of 10 nm. To determine the unfolding step size, the data points across the individual steps were fitted to a Heaviside step function with an additional linear term that take into account the continuous extension increase because of force change in the force scans. Finally, a t-test was carried out to test the height difference before and after the individual identified steps. Only statistically significant (P-value<0.01) steps were considered.
Force calibrations methods
The force applied to a bead depends on both the magnets–bead distance and the maximal magnetization of the bead. The latter is determined by the number of nano-magnetites in the bead, which has a large variation from one bead to another. At a fixed magnets–bead distance, this results in a large variation in force. According to our previous studies, forces applied to the weakest and the strongest M280 dynabeads at the same magnets–bead distance can differ by 100% (ref. 49).
In the single-protein manipulation studies of unfolding and refolding of αCM molecules, force was calibrated for individual beads based on their thermal fluctuations under force. A small relative error in force calibration (~5%) was ensured, which was mainly caused by the variation in the size of beads49.
In the photobleaching assay, a fixed magnets–bead distance was maintained and the sample chamber was scanned over 1 h time scale such that tethers in the chamber were subjected to the force as homogeneous as possible. Then, photobleaching assay was performed. As in this assay, the thermal motions of the beads were not tracked, the force estimated represented the average force and therefore had a significantly larger error compared with the single-molecule manipulation assay. To ensure most beads were subject to large enough force to unfold the αCM molecules, a fixed magnets–bead distance was chosen to have a relatively large average value of 20 pN.
Photobleaching counting
The photobleaching counting of the number of Alexa 488-labelled VD1 bound to a single WT αCM or 3X αCM under force were carried out using a total internal reflection microscope similar to the one described in the previous study of talin–vinculin interactions7. Force estimated in the photobleaching assay represented the average force and therefore has a significantly larger error compared with the single-molecule manipulation assay. Labelling and purification of the VD1 with Alexa 488 dye were based on the commercial microscale protein labelling kit (A30006) following the manufacturer’s protocol. The Alexa Fluro-488 reactive dye has a tetrafluorophenyl ester, which reacts efficiently with primary amines of proteins to form stable dye–protein conjugates.
The solution containing 50 μg ml−1 labelled VD1 was added to the flow-chamber coated with WT αCM or WT 3X αCM constructs. These constructs were subjected to force and incubated for 1 h, then unbound VD1 was flushed away before counting the number of bound VD1 by photobleaching events similar to previous studies of talin–vinculin interaction.
The images of TIRF fluorescence at 488 nm were taken for 1 min using Olympus IX71 (TIRF) microscope with a 1.45 numerical aperture, × 100 objective. Images were further analysed using in-house written Igor programme. Briefly, a region of interest containing a tethered magnetic bead was chosen and the total fluorescence intensity of this region of interest was recorded. The auto-fluorescence of the magnetic beads contributed to a background intensity, which was stable at our experimental time (~60 s) without any abrupt changes. For a tether with bound labelled VD1, photobleaching of single-dye molecules showed an abruptly decrease in fluorescence intensity, which allowed us to count the number of labelled VD1 associated to the tether.
Single-dye photobleaching events were identified following the method developed by del Rio et al. in their paper on force-dependent vinculin binding to talin7. Spontaneous stepwise decreases of fluorescent intensity in the time trajectories of the total fluorescence underneath the magnetic beads were identified as photobleaching steps of Alexa 488-labelled VD1. These photobleaching steps were not present in control experiments when Alexa 488-labelled VD1 was not added. In that case, only smooth exponential decay of auto-fluorescence of the bead was observed.
The number of tethered beads tested for the WT αCM at no force and 20 pN of applied force was 89 and 61, respectively. For the WT 3X αCM, the total number of tethered beads tested was 51 and 55 for no force and 20 pN force, respectively.
Determination of transition rates at constant forces
The force-dependent kinetic rates of the near-equilibrium unfolding/refolding transitions between 4 and 6 pN were obtained by fitting constant force extension fluctuation time traces to a Modified Hidden Markov model. An in-house written Matlab programme based on the Viterbi algorithm was used to find emission distribution and corresponding folded/unfolded state sequence with maximum likelihood66. The lifetime distributions of each state were then fitted to an exponential distribution to obtain their characteristic lifetime t. The unfolding and refolding transition rates at a given force were calculated as the inverse of the fitted characteristic lifetime of the unfolded and folded states. The distance to the transition state was calculated by the slope of the linear fit of the natural logarithm of the unfolding and refolding rates at different forces as described previously67.
Helix-to-coil transition estimation
Crystal structures were available for vinculin head binding to the VD1-binding α-helix of αE-catenin such as 4EHP. VD1 can only bind to the VBD of αE-catenin if the VD1-binding domain is under α-helix conformation. When interacting with VD1, the vinculin-binding site of αE−catenin adopts an α-helical structure with six turns of alpha helix that consists of around 22 residues and span a length of around 3.2 nm (0.54 nm per turn). Although at 25–40 pN, the length of a random coil of 22 amino acids is predicted to be 6.1–6.7 nm based on the worm-like-chain polymer model68 with a persistent length of 0.5 nm estimated from previous protein unfolding experiments56 (0.4–0.6 nm). Therefore, the size of steps expected for the helix-to-coil transition of the vinculin-binding α-helices when the vinculin head domain dissociates is ~2.9–3.5 nm.
Additional information
How to cite this article: Yao, M. et al. Force-dependent conformational switch of α-catenin controls vinculin binding. Nat. Commun. 5:4525 doi: 10.1038/ncomms5525 (2014).
Change history
04 March 2015
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Edelman, G. M. Cell adhesion molecules in the regulation of animal form and tissue pattern. Annu. Rev. Cell Biol. 2, 81–116 (1986).
Mammoto, T. & Ingber, D. E. Mechanical control of tissue and organ development. Development 137, 1407–1420 (2010).
Zhang, H. & Labouesse, M. Signalling through mechanical inputs: a coordinated process. J. Cell Sci. 125, 3039–3049 (2012).
Trichet, L. et al. Evidence of a large-scale mechanosensing mechanism for cellular adaptation to substrate stiffness. Proc. Natl Acad. Sci. USA 109, 6933–6938 (2012).
Morris, C. E. Mechanosensitive ion channels. J. Membr. Biol. 113, 93–107 (1990).
Sawada, Y. et al. Force sensing by mechanical extension of the Src family kinase substrate p130Cas. Cell 127, 1015–1026 (2006).
del Rio, A. et al. Stretching single talin rod molecules activates vinculin binding. Science 323, 638–641 (2009).
Yao, M. et al. Mechanical activation of vinculin binding to talin locks talin in an unfolded conformation. Sci. Rep. 4, 4610 (2014).
Mege, R. M., Gavard, J. & Lambert, M. Regulation of cell-cell junctions by the cytoskeleton. Curr. Opin. Cell Biol. 18, 541–548 (2006).
Ganz, A. et al. Traction forces exerted through N-cadherin contacts. Biol. Cell 98, 721–730 (2006).
Yonemura, S., Wada, Y., Watanabe, T., Nagafuchi, A. & Shibata, M. Alpha-catenin as a tension transducer that induces adherens junction development. Nat. Cell Biol. 12, 533–542 (2010).
Ladoux, B. et al. Strength dependence of cadherin-mediated adhesions. Biophys. J. 98, 534–542 (2010).
Huveneers, S. & de Rooij, J. Mechanosensitive systems at the cadherin-F-actin interface. J. Cell Sci. 126, 403–413 (2013).
Dufour, S., Mege, R. M. & Thiery, J. P. Alpha-catenin, vinculin, and F-actin in strengthening E-cadherin cell-cell adhesions and mechanosensing. Cell. Adh. Migr. 7, 345–350 (2013).
Watabe, M., Nagafuchi, A., Tsukita, S. & Takeichi, M. Induction of polarized cell-cell association and retardation of growth by activation of the E-cadherin-catenin adhesion system in a dispersed carcinoma line. J. Cell Biol. 127, 247–256 (1994).
Hirano, S., Kimoto, N., Shimoyama, Y., Hirohashi, S. & Takeichi, M. Identification of a neural alpha-catenin as a key regulator of cadherin function and multicellular organization. Cell 70, 293–301 (1992).
Silvis, M. R. et al. Alpha-catenin is a tumor suppressor that controls cell accumulation by regulating the localization and activity of the transcriptional coactivator Yap1. Sci. Signal 4, ra33 (2011).
Kobielak, A. & Fuchs, E. Alpha-catenin: at the junction of intercellular adhesion and actin dynamics. Nat. Rev. Mol. Cell Biol. 5, 614–625 (2004).
Lien, W. H., Klezovitch, O., Fernandez, T. E., Delrow, J. & Vasioukhin, V. AlphaE-catenin controls cerebral cortical size by regulating the hedgehog signaling pathway. Science 311, 1609–1612 (2006).
Sumida, G. M., Tomita, T. M., Shih, W. & Yamada, S. Myosin II activity dependent and independent vinculin recruitment to the sites of E-cadherin-mediated cell-cell adhesion. BMC Cell Biol. 12, 48 (2011).
Miyake, Y. et al. Actomyosin tension is required for correct recruitment of adherens junction components and zonula occludens formation. Exp. Cell Res. 312, 1637–1650 (2006).
le Duc, Q. et al. Vinculin potentiates E-cadherin mechanosensing and is recruited to actin-anchored sites within adherens junctions in a myosin II-dependent manner. J. Cell Biol. 189, 1107–1115 (2010).
Watabe-Uchida, M. Alpha-catenin-vinculin interaction functions to organize the apical junctional complex in epithelial cells. J. Cell Biol. 142, 847–857 (1998).
Weiss, E. E., Kroemker, M., Rudiger, A. H., Jockusch, B. M. & Rudiger, M. Vinculin is part of the cadherin-catenin junctional complex: complex formation between alpha-catenin and vinculin. J. Cell Biol. 141, 755–764 (1998).
Imamura, Y., Itoh, M., Maeno, Y., Tsukita, S. & Nagafuchi, A. Functional domains of alpha-catenin required for the strong state of cadherin-based cell adhesion. J. Cell Biol. 144, 1311–1322 (1999).
Thomas, W. A. et al. α-Catenin and vinculin cooperate to promote high E-cadherin-based adhesion strength. J. Biol. Chem. 288, 4957–4969 (2013).
Peng, X., Nelson, E. S., Maiers, J. L. & DeMali, K. A. New insights into vinculin function and regulation. Int. Rev. Cell. Mol. Biol. 287, 191–231 (2011).
Izard, T. et al. Vinculin activation by talin through helical bundle conversion. Nature 427, 171–175 (2004).
Rangarajan, E. S. & Izard, T. Dimer asymmetry defines alpha-catenin interactions. Nat. Struct. Mol. Biol. 20, 188–193 (2013).
Huber, O., Krohn, M. & Kemler, R. A specific domain in alpha-catenin mediates binding to beta-catenin or plakoglobin. J. Cell Sci. 110, (Pt 15): 1759–1765 (1997).
Aberle, H. et al. Assembly of the cadherin-catenin complex in vitro with recombinant proteins. J. Cell Sci. 107, 3655–3663 (1994).
Stappert, J. & Kemler, R. A short core region of E-cadherin is essential for catenin binding and is highly phosphorylated. Cell Commun. Adhes. 2, 319–327 (1994).
Huber, A. H., Nelson, W. J. & Weis, W. I. Three-dimensional structure of the armadillo repeat region of beta-catenin. Cell 90, 871–882 (1997).
Pokutta, S. & Weis, W. I. Structure of the dimerization and beta-catenin-binding region of alpha-catenin. Mol. Cell 5, 533–543 (2000).
Pokutta, S., Drees, F., Yamada, S., Nelson, W. J. & Weis, W. I. Biochemical and structural analysis of alpha-catenin in cell-cell contacts. Biochem. Soc. Trans. 36, 141–147 (2008).
Nagafuchi, A., Ishihara, S. & Tsukita, S. The roles of catenins in the cadherin-mediated cell adhesion: functional analysis of E-cadherin-alpha catenin fusion molecules. J. Cell Biol. 127, 235–245 (1994).
Yamada, S., Pokutta, S., Drees, F., Weis, W. I. & Nelson, W. J. Deconstructing the cadherin-catenin-actin complex. Cell 123, 889–901 (2005).
Desai, R. et al. Monomeric alpha-catenin links cadherin to the actin cytoskeleton. Nat. Cell Biol. 15, 261–273 (2013).
Peng, X., Maiers, J. L., Choudhury, D., Craig, S. W. & DeMali, K. A. Alpha-Catenin uses a novel mechanism to activate vinculin. J. Biol. Chem. 287, 7728–7737 (2012).
Itoh, M. Involvement of ZO-1 in cadherin-based cell adhesion through its direct binding to alpha catenin and actin filaments. J. Cell Biol. 138, 181–192 (1997).
Pokutta, S., Drees, F., Takai, Y., Nelson, W. J. & Weis, W. I. Biochemical and structural definition of the l-afadin- and actin-binding sites of alpha-catenin. J. Biol. Chem. 277, 18868–18874 (2002).
Kobielak, A., Pasolli, H. A. & Fuchs, E. Mammalian formin-1 participates in adherens junctions and polymerization of linear actin cables. Nat. Cell Biol. 6, 21–30 (2004).
Hansen, S. D. et al. Alpha-E-catenin actin binding domain alters actin filament conformation and regulates binding of nucleation and disassembly factors. Mol. Biol. Cell 24, 3710–3720 (2013).
Benjamin, J. M. et al. AlphaE-catenin regulates actin dynamics independently of cadherin-mediated cell-cell adhesion. J. Cell Biol. 189, 339–352 (2010).
Yang, J., Dokurno, P., Tonks, N. K. & Barford, D. Crystal structure of the M-fragment of alpha-catenin: implications for modulation of cell adhesion. EMBO J. 20, 3645–3656 (2001).
Choi, H. J. et al. alphaE-catenin is an autoinhibited molecule that coactivates vinculin. Proc. Natl Acad. Sci. USA 109, 8576–8581 (2012).
Ishiyama, N. et al. An autoinhibited structure of alpha-catenin and its implications for vinculin recruitment to adherens junctions. J. Biol. Chem 288, 15913–15925 (2013).
Rangarajan, E. S. & Izard, T. The cytoskeletal protein alpha-catenin unfurls upon binding to vinculin. J. Biol. Chem. 287, 18492–18499 (2012).
Chen, H. et al. Improved high-force magnetic tweezers for stretching and refolding of proteins and short DNA. Biophys. J. 100, 517–523 (2011).
Chen, H. et al. Differential mechanical stability of filamin A rod segments. Biophys. J. 101, 1231–1237 (2011).
Chen, H. et al. Mechanical perturbation of filamin A immunoglobulin repeats 20-21 reveals potential non-equilibrium mechanochemical partner binding function. Sci. Rep. 3, 1642 (2013).
Moore, S. W., Roca-Cusachs, P. & Sheetz, M. P. Stretchy proteins on stretchy substrates: the important elements of integrin-mediated rigidity sensing. Dev. Cell 19, 194–206 (2010).
Finer, J. T., Simmons, R. M. & Spudich, J. A. Single myosin molecule mechanics: piconewton forces and nanometre steps. Nature 368, 113–119 (1994).
Gao, Y., Sirinakis, G. & Zhang, Y. Highly anisotropic stability and folding kinetics of a single coiled coil protein under mechanical tension. J. Am. Chem. Soc. 133, 12749–12757 (2011).
Maillard, RA. et al. ClpX(P) Generates mechanical force to unfold and translocate its protein substrates. Cell 145, 459–469 (2011).
Junker, J. P., Ziegler, F. & Rief, M. Ligand-dependent equilibrium fluctuations of single calmodulin molecules. Science 323, 633–637 (2009).
Carrion-Vazquez, M. et al. Mechanical and chemical unfolding of a single protein: a comparison. Proc. Natl Acad. Sci. USA 96, 3694–3699 (1999).
Bakolitsa, C. et al. Structural basis for vinculin activation at sites of cell adhesion. Nature 430, 583–586 (2004).
Bois, P. R. J., O'Hara, B. P., Nietlispach, D., Kirkpatrick, J. & Izard, T. The vinculin binding sites of talin and alpha-actinin are sufficient to activate vinculin. J. Biol. Chem. 281, 7228–7236 (2006).
Barry, A. K. et al. Alpha-catenin cytomechanics - role in cadherin-dependent adhesion and mechanotransduction. J. Cell. Sci. 127, 1779–1791 (2014).
Huveneers, S. et al. Vinculin associates with endothelial VE-cadherin junctions to control force-dependent remodeling. J. Cell. Biol. 196, 641–652 (2012).
Gavard, J. et al. Lamellipodium extension and cadherin adhesion: two cell responses to cadherin activation relying on distinct signalling pathways. J. Cell. Sci. 117, 257–270 (2004).
Cohen, D. M., Chen, H., Johnson, R. P., Choudhury, B. & Craig, S. W. Two distinct head-tail interfaces cooperate to suppress activation of vinculin by talin. J. Biol. Chem. 280, 17109–17117 (2005).
Peng, X., Cuff, L. E., Lawton, C. D. & DeMali, K. A. Vinculin regulates cell-surface E-cadherin expression by binding to beta-catenin. J. Cell. Sci. 123, 567–577 (2010).
Tsao, K. L., DeBarbieri, B., Michel, H. & Waugh, D. S. A versatile plasmid expression vector for the production of biotinylated proteins by site-specific, enzymatic modification in Escherichia coli. Gene 169, 59–64 (1996).
Viterbi, A. Error bounds for convolutional codes and an asymptotically optimum decoding algorithm. IEEE Trans. Inf. Theory 13, 260–269 (1967).
Elms, P. J., Chodera, J. D., Bustamante, C. & Marqusee, S. The molten globule state is unusually deformable under mechanical force. Proc. Natl Acad. Sci. USA 109, 3796–3801 (2012).
Bustamante, C., Marko, J. F., Siggia, E. D. & Smith, S. Entropic elasticity of lambda-phage DNA. Science 265, 1599–1600 (1994).
Bell, G. I. Models for the specific adhesion of cells to cells. Science 200, 618–627 (1978).
Acknowledgements
We thank the Mechanobiology Institute Protein Expression Facility for VD1 expression and purification services. We thank Anne Houllier for her help in the construction and production of the tagged α-catenin recombinant proteins. We also thank Didier Chatenay, Michael Sheetz and André Sobel for stimulating discussions and critical reading of the manuscript. Work in Singapore was supported by the National Research Foundation of Singapore through the Mechanobiology Institute at National University of Singapore (to J.Y., B.L., C.L. and R.L.); work in France was supported by grants from CNRS, ARC, as well as from Human Frontier Science Program (HFSP grant RPG0040/2012), and Agence Nationale de la Recherche (ANR 2010 Blan1515). R.S. was supported by an Ile de France Region fellowship cNano.
Author information
Authors and Affiliations
Contributions
J.Y., R.-M.M., B.L., C.T.L. and R.L. designed the research and supervised the experiments. R.S. and M.P. expressed αCM constructs and carried out the GST-VinH pull-down experiments. M.Y. performed the magnetic tweezers experiments and W.Q. performed the photobleaching counting experiments. M.Y., W.Q., J.Y., A.K.E., P.C. and R.-M.M. analysed the data. M.Y., J.Y., R.-M.M. and B.L. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-4 (PDF 798 kb)
Rights and permissions
About this article
Cite this article
Yao, M., Qiu, W., Liu, R. et al. Force-dependent conformational switch of α-catenin controls vinculin binding. Nat Commun 5, 4525 (2014). https://doi.org/10.1038/ncomms5525
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms5525
This article is cited by
-
Talking with force at cell–cell adhesions
Nature Cell Biology (2024)
-
Cardiac fibroblasts and mechanosensation in heart development, health and disease
Nature Reviews Cardiology (2023)
-
Dynamics and functions of E-cadherin complexes in epithelial cell and tissue morphogenesis
Marine Life Science & Technology (2023)
-
The role of single-protein elasticity in mechanobiology
Nature Reviews Materials (2022)
-
Tension can directly suppress Aurora B kinase-triggered release of kinetochore-microtubule attachments
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.