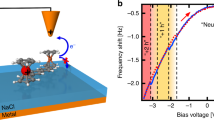
Abstract
Understanding the microscopic elementary process of chemical reactions, especially in condensed phase, is highly desirable for improvement of efficiencies in industrial chemical processes. Here we show an approach to gaining new insights into elementary reactions in condensed phase by combining quantum chemical calculations with a single-molecule analysis. Elementary chemical reactions in liquid-phase, revealed from quantum chemical calculations, are studied by tracking the fluorescence of single dye molecules undergoing a reversible redox process. Statistical analyses of single-molecule trajectories reveal molecular reaction kinetics and dynamics of elementary reactions. The reactivity dynamic fluctuations of single molecules are evidenced and probably arise from either or both of the low-frequency approach of the molecule to the internal surface of the SiO2 nanosphere or the molecule diffusion-induced memory effect. This new approach could be applied to other chemical reactions in liquid phase to gain more insight into their molecular reaction kinetics and the dynamics of elementary steps.
Similar content being viewed by others
Introduction
The kinetics of some chemical reactions has been extensively studied with ensemble-averaged methods1,2. As for the molecular reaction dynamics, only some simple (few atoms) gas-phase elementary reactions have been well studied, based on techniques, such as cross-molecular beams and infrared chemiluminescence3,4. As for the chemical reactions in condensed phase, much less has been done about the molecular reaction dynamics due to the complexity of traditional methods5,6. Recent advances in single-molecule fluorescence microscopy have allowed the observations of individual molecules, such as their translational diffusion7,8, rotational diffusion9, spectral fluctuation10, conformational motion11, the studies of single enzyme or single nanoparticle catalysis12,13,14,15,16,17, and the distinguishing of alternative reaction pathways18. Single-molecule measurements can reveal the distributions of molecular static or dynamic properties in inhomogeneous system19. These distributions cannot usually be determined by ensemble-averaged methods. Stochastic trajectories from single-molecule experiments contain detailed dynamic information extractable through statistical analysis. For chemical reactions, only the kinetics of some ‘apparent’ non-elementary chemical reactions or process has been studied through single-molecule approach15. Of particular interest, however, is the study of molecular reaction kinetics and dynamics via the real-time observation of individual elementary chemical reactions.
Here we demonstrate the use of quantum chemical calculations to examine elementary molecular reaction kinetics and dynamics in a homogeneous liquid phase. This is based on a reversible redox process on individual molecules from single-turnover level. Statistical analysis reveals molecular reaction kinetics and dynamics that provide new insight into elementary chemical reactions in condensed phase at single-molecule level. This new information is of great significance for the basic understanding of chemical reactions and their applications.
Results
Single-molecule reaction and quantum chemical calculations
We found that the fluorescent resorufin (on-state) can be reduced to non-fluorescent dihydroresorufin (off-state) by sodium borohydride (NaBH4) in water under pH 7.3, while the dihydroresorufin can be oxidized easily back to fluorescent resorufin by oxygen (Fig. 1a)20. This process can be reproduced again and again for many times due to the reversibility of this redox process (Supplementary Figs 1 and 2)21,22,23.
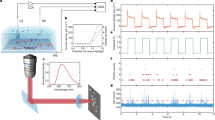
(a) Experimental design of single-molecule chemical reaction: a single-resorufin molecule locked in a porous hollow SiO2 nanosphere was reduced by NaBH4 to non-fluorescent dihydroresorufin, which was then oxidized back to fluorescent resorufin by oxygen. (b) A segment of a typical single-molecule trajectory of fluorescence for the repeated redox process of a single dye molecule. The binning is 50 ms.
The above repetitive fluorescence on-off phenomenon is the basis for observation of real-time single-molecule reaction by detecting the fluorescence of the same single dye molecule repeatedly at single-reaction single-turnover level. To monitor the individual chemical reactions in situ, first, porous hollow SiO2 nanospheres24 with inner diameter about 210 nm (Supplementary Fig. 3) were filled with diluted resorufin aqueous solution to form SiO2/Dye complex (no. of resorufin molecules ≤1 for each nanosphere, for details see Supplementary Methods and Discussion), then immobilized on quartz surface and flowed the substrate solution over (containing both reductant NaBH4 and oxidant O2; Fig. 1a). Using an inverted total internal reflection fluorescence microscope with down to 50 ms integration time and 532 nm laser excitation, we recorded movies of stochastic fluorescence bursts at some localized spots. A typical time trajectory of fluorescence intensity from these spots contains stochastic on-off digital signal that spans the entire movie (up to 1 h, Fig. 1b and Supplementary Fig. 2). The digital nature of the trajectory and the consistent height of the on-level indicate that each fluorescence burst comes from a single-resorufin molecule15.
We attributed the single-resorufin fluorescence bursts to a single-turnover of the reversible redox reaction as shown in Fig. 1. Each time when the resorufin is formed by the oxidation of the only dihydroresorufin molecule in a SiO2 nanosphere, a sudden increase in intensity is seen in the fluorescence trajectory. In contrast, each time when the only resorufin molecule in this nanosphere is reduced to nonfluorescent dihydroresorufin by NaBH4, a sudden decrease in intensity is seen. This, each on-off cycle, corresponds to a single turnover of the reversible redox reaction. All the turnovers on each trajectory are due to the repeated redox process of the same single dye molecule. This is different from that observed on single-molecule single-nanoparticle catalysis15,25, in which each burst comes from a new or different product molecule.
Many observations support our conclusion. (1) No digital fluorescence bursts are observed in the absence of SiO2/Dye complex. (2) Only short lifetime (with average lifetime ~1 min) blinking and photobleaching are observed in the absence of NaBH4, O2 or both (Supplementary Fig. 4). (3) The fluorescence bursts are not photoinduced (Supplementary Fig. 5)15.
As shown in Fig. 1b, there are two waiting times in these single-molecule fluorescence trajectories. Resolving them enables us to probe the kinetic mechanism of this chemical reaction from single-molecule level in two parts separately: τon (approximate waiting time needed for the reduction of resorufin) and τoff (approximate waiting time needed for the oxidation of dihydroresorufin back to resorufin). Their individual values are stochastic, but their statistical properties, such as average values and distribution, are well defined by the underlying reaction kinetics. Statistically15, in this case, ‹τon›−1 and ‹τoff›−1, where ‹› denotes averaging, represent the time-averaged single redox reaction rates of reduction and oxidation processes, respectively. When averaged over the turnover trajectories from many individual redox molecules, ‹τon›−1 and ‹τoff›−1, should be expectedly dependent on reductant or oxidant concentrations, respectively, and exhibit saturation kinetics14.
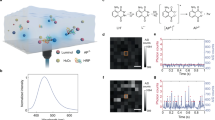
In order to tell the on- and off-process precisely, we need to know the detailed reduction mechanism and the fluorescence states of all the stable intermediates. For that goal, the NaBH4 reduction mechanism was studied in detail to find the most feasible reaction pathways and stable intermediates by quantum chemical calculation (Supplementary Methods and Discussion). NaBH4 has been extensively used as reducing agent for organic chemistry26,27,28,29. Previous work20,29,30 has shown that the reducing process of resorufin by NaBH4 should follow the attacking of BH4− to the carbonyl group to form IM1 first (Fig. 2a) and finally, the hydrolysis of IM2 to product dihydroresorufin. After investigating several possible mechanisms (Supplementary Fig. 6), the optimal one with smallest energy barrier was determined, as shown in Fig. 2a. Figure 2b is the corresponding energy profile for this reduction pathway. Here it was found that Na+ cation and water molecule have participated in the first transition state collectively via the nucleophilic addition of BH4− to the carbonyl group in resorufin, which occurs easily due to a very small activation energy. Obviously the process from substrate to IM1 is an elementary step. It was confirmed here that the proton on the C atom in the carbonyl group could be taken off easily by water from IM1 to form IM2a, with a small energy barrier of 12.0 kcal mol−1. The C atom in sp3 hybridization in IM2a tends to turn into sp2 hybridization to form another resonance structure with stable phenyl ring in IM2b. This was followed by the spontaneous fast protonation process on the N atom (inset in Fig. 2b), that is, the intermediate IM2b and then IM2. So the rate constant k2 approximately represents reactivity of the slow elementary step from IM1 to IM2a. The hydrolysis process of IM2, an elementary-reaction step, occurred via a larger activation energy to form the reduction product dihydroresorufin, where Na+ cation and water also play an important role. In other words, the hydrolysis process (k3) is the rate-determining step in the whole reduction process, which is consistent with the long τoff observed in the following single-molecule experiments. For the oxidation process, dihydroresorufin is known to be easily oxidized by oxygen back to the initial substrate resorufin20. This molecule will go to the next redox cycle again and again. The quantum chemical calculations have also been carried out to predict the properties for ground state (S0) and singlet excited state (S1) of the intermediates found in the reduction process. The results show the intermediate IM1 can still emit strong fluorescence due to the intact planar π system, and intermediate IM2 shows turned-off fluorescence due to the distorting non-planar Λ-type geometry (see Supplementary Table 1). As this calculation can reliably predict the fluorescence states of the initial substrate (resorufin) and the final product dihydroresorufin, the predictions for the two intermediates (IM1 and IM2) seem plausible.
(a) Schematic diagram of the mechanism for the redox process of single dye (resorufin) molecules. The fluorescence state (on or off) of the dye is indicated at each reaction step. (b) The energy profile for resorufin reduction by NaBH4 (kcal mol−1). Inset: IM2* represents there is one fast charge rearrangement process from IM2a to IM2b (bottom inset), and then spontaneous protonation process to IM2.
Molecular reaction kinetics and dynamics of elementary reactions
Based on the above information, we can tell that the τon process includes the steps shown in Fig. 3:
The ‘on’ represents either resorufin or IM1. Following a single-molecule kinetic analysis (Supplementary Methods and Discussion)31, we get the probability density function of τon, fon(τ) as

The reciprocal of the first moment of fon(τ), ‹τon›−1, represents the reduction rate of a dye molecule:

Equation (2) connects τon, a stochastic single-molecule quantity, with conventional kinetic parameters and predicts the saturation kinetics of ‹τon›−1. Fitting the molecule-averaged data of ‹τon›−1 at different [NaBH4], shown in Fig. 4a, gives k1=522±90 mM−1 s−1 and k2=4.93±0.05 s−1, both values here reflecting averaged properties of many dye molecules.
Reductant and oxidant concentration dependence of ‹τon›−1 (a,d) and ‹τoff ›−1 (b,c). Each data point is an average from about 80 trajectories. Solid lines are fits from Equations (2), (4) and (5). Errors are s.d. Experiments for (a,c) are in water solution with 1.3 mM O2, for (b,d) are in water solution with 0.26 mM NaBH4. Insets: the linear dependence of turnover rate on [O2]0.23 (c) and [NaBH4]0.47 (b). (e)The on-time distribution derived from redox process of a single dye molecule with 0.45 mM NaBH4 and 1.3 mM O2 (k2 being the rate-limiting step for τon). Solid line is the single exponential fit with k2=4.41±0.16 s−1. Inset is the distribution of k2 derived from about 90 dye molecules in the same sample. (f) The off-time distribution derived from redox process of a single dye molecule with 0.26 mM NaBH4 and 1.3 mM O2 (k3 being the rate-limiting step for τoff). Solid line is the single exponential fit with k3=0.5±0.2 mM−1 s−1. Inset is the distribution of k3 derived from about 90 dye molecules in the same sample. Note that k2 and k3 are time averaged over entire trajectories.
The above model also predicts that the τon process has nothing to do with [O2]. In the single-molecule experiments with a wide range of different [O2] and the same [NaBH4], ‹τon›−1 roughly showed no change, as shown in Fig. 4d.
For the τoff process, it contains the steps shown in Fig. 5.
It clearly shows the τoff is not a pure oxidation process of dihydroresorufin. It contains the hydrolysis process of IM2 and the oxidation process of dihydroresorufin by O2. As the quantum chemical calculation (Fig. 2) shows the Na+ cation, which mainly comes from NaBH4, participates in the hydrolysis process of IM2, the ‹τoff›−1 should depend on both [NaBH4] and [O2]. Following a similar single-molecule kinetic analysis, we get the probability density function of τoff (Supplementary Methods and Discussion):

When we vary the concentration of NaBH4 with constant [O2], Equation (3) can be simplified as:

with  .
.
Then

Fitting the molecule-averaged data of ‹τoff›−1 at different [NaBH4] with fixed O2 concentration, as shown in Fig. 4c, gives k3=1.1±0.1 mM−1 s−1 and k40=0.23±0.02 s−1. As the O2 concentration is about 1.3 mM in this case, then k4 is about 0.22 mM−1/2 s−1.
Similarly, when we vary the O2 concentration with constant [NaBH4], the probability density function of τoff, foff(τ) is

with  .
.
and

Here, the rate constant k3 contains water contribution as its concentration is very high and keeps constant during the whole reaction process. Fitting the molecule-averaged data at different [O2], shown in Fig. 4b, gives k30=0.21±0.02 s−1 and k4=0.4±0.1 mM−1/2 s−1. Based on [NaBH4]=0.26 mM, k3 is estimated to be about 0.81 mM−1 s−1, which reflects the rate of the hydrolyzation of intermediate IM2. Compared with k1 and k2 for τon, the much smaller k3 and k4 indicate the τoff is a much slower process than τon. That is why ‹τoff› observed here is much longer than ‹τon›.
As shown above, two independent experiments that had varying reductant and oxidant concentrations were used to determine k3 and k4, respectively. Values obtained for k3 and k4 are within the same order of magnitude and approximately consistent within error range. The differences in values of k3 and k4 are about 26 and 45%, respectively. The big differences probably come from the estimation error of NaBH4 concentration due to its slow hydrolysis process.
The reduction process of resorufin by NaBH4 was studied precisely based on elementary steps discriminated from quantum chemical calculation. This provided detailed kinetic information for each elementary step, helping us to understand elementary chemical reactions more deeply. This has not been shown in previous reports. Obviously the fine discrimination and the kinetic study of elementary chemical reactions usually cannot be done with traditional ensemble-averaged methods. The new knowledge obtained in this approach, such as the rate constants for some rate-limiting elementary steps, is of great significance for the optimization of chemical reactions.
On the other hand, from the above we can see the apparent net reaction is:

The total turnover rate rtotal=kapp[NaBH4]n[O2]k, where n, k are the real reaction orders with respect to NaBH4 and O2, respectively.
For single dye molecule-based reaction occurred in individual SiO2 hollow nanospheres,  , in the concentration range of NaBH4 and O2 used in the present work,
, in the concentration range of NaBH4 and O2 used in the present work,  , so
, so  . As the τoff process is related to [NaBH4] according to Equation (4),
. As the τoff process is related to [NaBH4] according to Equation (4),  when [O2] is fixed, by fitting the data shown in the inset of Fig. 4c,
when [O2] is fixed, by fitting the data shown in the inset of Fig. 4c,  and n with respect to NaBH4 were obtained to be about 0.225±0.004 mM−0.47 s−1 and 0.47±0.01, respectively. Similarly for O2, when [NaBH4] is fixed,
and n with respect to NaBH4 were obtained to be about 0.225±0.004 mM−0.47 s−1 and 0.47±0.01, respectively. Similarly for O2, when [NaBH4] is fixed,  , as shown by the fit in the inset of Fig. 4b,
, as shown by the fit in the inset of Fig. 4b,  and k were obtained to be 0.15±0.01 mM−0.23 s−1 and 0.23±0.01, respectively. Obviously in this special case the real reaction orders (n=0.47, k=0.23) with respect to NaBH4 and O2 are different from the stoichiometric coefficients (1 and 1/2). This phenomenon has been extensively observed before based on ensemble-averaged methods32. It is interesting to see that the ratio between the reaction order n and k is 2, the same as the ratio between the stoichiometric coefficients. Furthermore, based on the obtained n and k, kapp is obtained to be about 0.25 mM−0.7 s−1. The disclosure of the real reaction orders and apparent rate constant for single-molecule chemical reactions above exemplifies a new power of single-molecule method in revealing the molecular reaction dynamics.
and k were obtained to be 0.15±0.01 mM−0.23 s−1 and 0.23±0.01, respectively. Obviously in this special case the real reaction orders (n=0.47, k=0.23) with respect to NaBH4 and O2 are different from the stoichiometric coefficients (1 and 1/2). This phenomenon has been extensively observed before based on ensemble-averaged methods32. It is interesting to see that the ratio between the reaction order n and k is 2, the same as the ratio between the stoichiometric coefficients. Furthermore, based on the obtained n and k, kapp is obtained to be about 0.25 mM−0.7 s−1. The disclosure of the real reaction orders and apparent rate constant for single-molecule chemical reactions above exemplifies a new power of single-molecule method in revealing the molecular reaction dynamics.
The probability density function of τon, as shown in Equation (1), is the convolution of two exponentials (k1[NaBH4] and k2)33. In this case, k1=522±90 mM−1 s−1 and k2=4.93±0.05 s−1. The much larger k10 (=k1[NaBH4]) than k2 makes the convolution in τon invisible even at the lowest concentration of NaBH4 (0.065 mM), at which we can only see the monotonic decay, as is the case observed before for single nanoparticle catalysis15. On the other hand, when [NaBH4]→∞,  . It predicts that the waiting time (τon) distribution for each single molecule should be a single exponential when the reductant concentration is high enough. As an example shown in Fig. 4e at high reductant concentration, from the decay constant we obtained the rate constant of k2 for this single molecule to be about 4.41±0.16 s−1. To see the static heterogeneity of this type of molecule, we obtained the values of k2 for many individual molecules. From the distribution (insert in Fig. 4e with ‹k2›=4.62±0.89 s−1) we can see the difference is very big among these individual molecules, although they are all exactly the same in chemical composition and physical structure.
. It predicts that the waiting time (τon) distribution for each single molecule should be a single exponential when the reductant concentration is high enough. As an example shown in Fig. 4e at high reductant concentration, from the decay constant we obtained the rate constant of k2 for this single molecule to be about 4.41±0.16 s−1. To see the static heterogeneity of this type of molecule, we obtained the values of k2 for many individual molecules. From the distribution (insert in Fig. 4e with ‹k2›=4.62±0.89 s−1) we can see the difference is very big among these individual molecules, although they are all exactly the same in chemical composition and physical structure.
Similarly, the probability density function of τoff (Equation (3)) is the convolution of two exponentials (k3[NaBH4] and k4[O2]1/2). We checked the τoff distributions for all the trajectories at different [NaBH4] or [O2] concentrations; some distributions from single-molecule trajectories (about 1%) show the obvious convolution of two exponentials (Supplementary Fig. 7). However, from these distributions we cannot get precise values for the rate constants from the double-exponential fitting (Supplementary Methods and Discussion). On the other hand, for Equation (3b), when[O2]→∞,  . Thus, we can get the value of k3 from
. Thus, we can get the value of k3 from  for each individual molecule from single exponential distribution of waiting time (τoff) when the oxidant concentration is high enough. Indeed, as shown in Fig. 4f, we obtained the rate constant of k3 for a single molecule to be about 0.5±0.2 mM−1 s−1. The values of k3 for many individual molecules (insert in Fig. 4f) also show big static heterogeneity of this type of molecule with ‹k3›=0.54±0.15 mM−1 s−1. These static heterogeneities could be partially attributed to the heterogeneous microenvironments around each molecule. The different microenvironments probably could be attributed to the static heterogeneity of individual SiO2 nanospheres, such as different sizes, porosities and surface patterns in nanospheres.
for each individual molecule from single exponential distribution of waiting time (τoff) when the oxidant concentration is high enough. Indeed, as shown in Fig. 4f, we obtained the rate constant of k3 for a single molecule to be about 0.5±0.2 mM−1 s−1. The values of k3 for many individual molecules (insert in Fig. 4f) also show big static heterogeneity of this type of molecule with ‹k3›=0.54±0.15 mM−1 s−1. These static heterogeneities could be partially attributed to the heterogeneous microenvironments around each molecule. The different microenvironments probably could be attributed to the static heterogeneity of individual SiO2 nanospheres, such as different sizes, porosities and surface patterns in nanospheres.
The reactivity dynamic fluctuations of single molecules
From each single-turnover trajectory, we further determined the time dependence of the turnover rates (the number of off-on cycles per second; Fig. 6a). Large temporal variations were observed, indicating dynamic reactivity fluctuations of individual dye molecules, consistent with the revelations from the Poisson indicator (P)34, which is related to the second moment of f(t) (Supplementary Figs 8 and 9). The reactivity fluctuations could be related to reaction rate changes in the τon or τoff process. To assess the contribution of τon and τoff processes to the reactivity fluctuations, we extracted the sequence of individual τon and τoff from each turnover trajectory, and then calculated their autocorrelation functions Cτ(m)=‹Δτ(0)Δτ(m)›/‹Δτ2› (refs 14, 15, 35). Here, τ is either τon or τoff, m is the turnover index number in the sequence and Δτ(m)=τ(m)−‹τ›, the delay time. In the presence of reactivity fluctuations, Cτ(m)>0 shows a decay behaviour with the decay time constant being the fluctuation correlation time14,35.
(a) Trajectory of turnover rate (s−1) for a single dye molecule at 0.45 mM NaBH4 and 1.3 mM O2. Data points are calculated every five turnovers. (b,c) Autocorrelation functions of τon and τoff derived from the same trajectory as that in a. Solid lines are the exponential fits with decay constants of mon=2.3±0.7 and moff=13.3±5.5 turnovers. Insets: Histograms of fluctuation correlation times for τon (k2) and τoff (k3). (d) Mapping of Brownian motion of a single dye molecule inside a hollow SiO2 nanosphere with optical super-resolution imaging. The dotted circle indicates the inner cavity of the nanosphere; the solid one indicates the centre area of the cavity. Top left: the distribution of the dye in x-direction; Bottom right: the distribution of the dye in y-direction; Top right: a typical transmission electron microscopy image of a hollow SiO2 nanosphere, the scale bar, 200 nm.
For a dye molecule in a hollow SiO2 nanosphere at saturating reductant (0.45 mM of NaBH4) or oxidant (1.3 mM of O2) concentrations, both  and
and  show exponential decay, indicating both reduction and hydrolysis rates fluctuate or a memory effect14. For instance, as shown in Fig. 6b, c, the decay constants are mon=2.3±0.7 and moff=13.3±5.5 turnovers obtained from two different trajectories with [NaBH4] and [O2] saturated, respectively. These two trajectories were from NaBH4- and O2-titration experiments, respectively, as shown above. With average turnover time of 9.3 and 6.2 s for these two turnover trajectories, the fluctuation correlation times for τon and τoff related to the delay process are ~21.4 and ~72.5 s, respectively. These two correlation times reflect the fluctuation or delay timescales of k2 and k3, which are rate-limiting in τon and τoff processes at saturating reductant and oxidant concentrations. These two steps also are the key elementary steps for the reduction process of dye by NaBH4 and could never be resolved and studied separately by any traditional method. The insets in Fig. 6b,c show the histograms of fluctuation correlation time of τon (k2) and τoff (k3) processes from many individual dye molecules; their broad distributions indicate that the reactivity of these resorufin molecules fluctuates in very different timescales. To further confirm the existence of dynamic disorder or memory effect, we presented a statistical analysis of two-dimensional conditional probability distributions for a pair of adjacent on-times and off-times, as shown in Supplementary Fig. 10, no diagonal features could be seen clearly, indicating that the two-dimensional histograms of waiting times are not sensitive to the dynamic disorder or memory effect14, similar to that previously observed for single nanoparticle catalysis15.
show exponential decay, indicating both reduction and hydrolysis rates fluctuate or a memory effect14. For instance, as shown in Fig. 6b, c, the decay constants are mon=2.3±0.7 and moff=13.3±5.5 turnovers obtained from two different trajectories with [NaBH4] and [O2] saturated, respectively. These two trajectories were from NaBH4- and O2-titration experiments, respectively, as shown above. With average turnover time of 9.3 and 6.2 s for these two turnover trajectories, the fluctuation correlation times for τon and τoff related to the delay process are ~21.4 and ~72.5 s, respectively. These two correlation times reflect the fluctuation or delay timescales of k2 and k3, which are rate-limiting in τon and τoff processes at saturating reductant and oxidant concentrations. These two steps also are the key elementary steps for the reduction process of dye by NaBH4 and could never be resolved and studied separately by any traditional method. The insets in Fig. 6b,c show the histograms of fluctuation correlation time of τon (k2) and τoff (k3) processes from many individual dye molecules; their broad distributions indicate that the reactivity of these resorufin molecules fluctuates in very different timescales. To further confirm the existence of dynamic disorder or memory effect, we presented a statistical analysis of two-dimensional conditional probability distributions for a pair of adjacent on-times and off-times, as shown in Supplementary Fig. 10, no diagonal features could be seen clearly, indicating that the two-dimensional histograms of waiting times are not sensitive to the dynamic disorder or memory effect14, similar to that previously observed for single nanoparticle catalysis15.
Discussion
Interestingly, here a long timescale from seconds to tens of seconds has been observed from the on-time and off-time autocorrelation functions. The long autocorrelation time indicates a slow fluctuation of the microenvironment. Obviously, solvent rearrangements and vibrational or rotational motion of the dye molecule occur on a tremendously faster timescale than the observed decays. So what kind of fluctuations of the microenvironment around the dye molecules occur on such a slow timescale? The first reason probably could be due to the low-frequency approaching or contact of single dye molecule to the internal surface of SiO2 nanosphere. Each time when the free dye molecule approaches to the Si or O atoms on the internal surface of SiO2 nanosphere, its reactivity with reductant or oxidant probably will be affected due to the asymmetric microenvironment (one side is solid wall, the other side is liquid)36,37. In order to confirm above analysis, we applied a Monte-Carlo simulation to mimic the Brownian motion of a single dye molecule confined in the nanosphere (Supplementary Methods and Discussion). The Monte-Carlo time with unit mcs (Monte-Carlo step) and the times of the contact between dye molecule and internal surface of the hollow nanosphere were recorded (Supplementary Table 2). It shows, when the sphere diameter is increased from 20 to 200 (10 times), the average waiting time for a contact between the dye molecule and the internal surface of the nanosphere is increased from 43 to 6,510 mcs (more than 150 times). According to previous results for particle Brownian motion38,39, the long waiting time of 6,510 mcs could approximately correspond to a real-time scale of up to second. So, the low-frequency approaching of dye molecule to the inner surface of SiO2 nanosphere probably contributes in part to the slow fluctuation of microenvironment around the single dye molecule. Another possible reason for the slow fluctuation of microenvironment was single-molecule diffusion-induced memory effect, which was observed unexpectedly in the following single-molecule tracking analysis (see more discussions there). Both the τoff and τon processes probably depend on the same fluctuations mentioned above, but maybe in different ways, which then leads to different decays of τoff and τon.
To get more clues about reactivity dynamic fluctuation of single dye molecules, we used the fitting of the emission point spread function40,41, the basis for optical super-resolution imaging methods23,42,43, to ‘track’ the movement of single dye molecules in SiO2 nanospheres. In this way, we can map the location variation of a free single dye molecule in situ inside the cavity of a hollow SiO2 nanosphere during the repeated redox process (Supplementary Methods and Discussion; Supplementary Figs 11 and 12). Centre locations of single dye molecule at different time were obtained from many τon periods in one fluorescence trajectory with better than 10 nm precision. As shown in Fig. 6d, all the centres of the same dye molecule in the on-state from different τon appear in a circle with diameter about 150 nm, which is close to the averaged inner diameter of hollow SiO2 nanospheres used here (top right: transmission electron microscopy image).
To get more information from the Brownian motion of single dye molecule inside the SiO2 nanosphere, the position of dye molecule on each frame of τon was obtained and analysed deeply. According to the scheme shown in Fig. 7a, we calculated two sequences of distances from each trajectory: D 1 {a1a2, a2a3,…an−1an, b1b2, b2b3,..bn−1bn,…} sequence for the molecule on two adjacent frames in the same τon, such as between ai (or bi, ci…) and ai+1 (or bi+1, ci+1…) (i=1, 2…n); Dk {anb1, bnc1,…dne1…} sequence for the molecule on two frames exactly separated by a τoff, such as between an (or bn, cn…) and b1 (or c1, d1…). It was found that the averaged Dk (36±6 nm), the distance for the molecule before/after multiple steps diffusion, is much longer than the averaged D 1 (26±4 nm), the distance for the molecule on two adjacent frames or the diffusion step-size, indicating a divergent diffusion pattern of the molecule. Furthermore, we calculated the autocorrelation coefficients for these two sets of distances (D 1 and Dk), respectively. Interestingly, the D 1 sequence shows strong positive autocorrelation with coefficient of ρ=0.425. While the Dk sequence shows no autocorrelation (ρ=−0.05). These data unambiguously show a dynamic disorder or a memory effect related to the diffusion of dye molecule, indicating a short diffusion-step tends to be followed by another short step, and a long step tends to be followed by another long step. The molecule diffusion-induced memory effect has been realized to be the basis of molecular communication44,45. In it, the messages can be conveyed via the variation in the concentration of molecules in the small-scale medium. However, due to the delay of the variation of concentration of molecules, the dynamic disorder or memory effect appears during the molecule diffusion process44.
(a) The scheme to show how the distance sequences (D 1 and Dk) were calculated from τon on a fluorescence trajectory. D 1:{a1a2, a2a3,…an−1an, b1b2, b2b3,..bn−1bn,…} for the molecule on two adjacent frames in the same τon; Dk:{anb1, bnc1,…dne1…} for the molecule on two frames exactly separated by a τoff (n is the total number of frames in one τon, and the value should be different for different τon). (b,c) The autocorrelation functions of the distance sequences (D 1 (b) and Dk (c)). The solid line in b is the exponential fit with decay constant of m=1.7. The m is the index number in sequence.
To evaluate the timescale of this diffusion-induced memory effect in this case, we further calculated the autocorrelation function for the data sets (D 1 and Dk). Interesting, as an example shown in Fig. 7b, the decay constant for the successive diffusion steps (D 1 sequence) is m=1.7, corresponding to the autocorrelation time of about 0.1 s, further confirming a dynamic disorder or a memory effect (Supplementary Fig. 13a)14. While as shown in Fig. 7c, distances (Dk sequence) for the points separated by multiple (⩾2) frames show no autocorrelation or memory effect (Supplementary Fig. 13b). These facts are consistent with previous observation due to a molecule diffusion-based memory effect44. From multiple individual molecules, we found the averaged diffusion-induced autocorrelation time is about 0.5 s. The slow timescale observed here is approximately in the range of that observed from τon and τoff autocorrelation shown in Fig. 6. So the diffusion-induced dynamic fluctuation observed here may also contribute in part to the slow dynamic fluctuation of the microenvironment or the reactivity of single dye molecules.
Here we examined the kinetics and dynamics of three tandem homogeneous elementary chemical reactions in real time from single-molecule level at single-turnover resolution. This study exemplifies a new powerful tool of quantum chemical calculation-based single-molecule approach in revealing new insight of elementary molecular reaction kinetics and dynamics. By this approach, the molecular reaction kinetics, dynamics and the reactivity dynamic fluctuation of a complex chemical process in a homogeneous liquid phase were studied thoroughly from an elementary-reaction level. In principle, this new approach may be applicable for some other chemical reactions in liquid phase to get more insight about their molecular reaction kinetics and dynamics for elementary steps.
Methods
Hollow SiO2 nanoparticle synthesis and characterization
The hollow SiO2 nanoparticles were synthesized in two steps. Briefly, solid SiO2 spheres were made first, and then etched to hollow mesoporous silica nanospheres with wormhole-like shell. These SiO2 nanospheres are hollow and porous in structure with inner diameters about 210 nm averagely.
Single-molecule chemical reaction experiments
Single-molecule fluorescence measurements were performed on a homebuilt prism-type total internal reflection fluorescence microscope based on an Olympus IX71 inverted microscope. A continuous wave circularly polarized 532 nm laser beam (CrystaLaser, GCL-025-L-0.5%) of 4–5 mW was focused onto an area of ~80 × 80 μm2 on the sample to directly excite the fluorescence of resorufin. The fluorescence of resorufin was collected by a × 60 NA1.2 water-immersion objective (UPLSAPO60XW, Olympus), filtered by two filters (HQ550LP, HQ580m60), and projected onto a camera (Andor iXon EMCCD, DU-897U-CS0-#BV), which is controlled by an Andor IQ software. All optical filters are from Chroma Technology. The movies are analysed using a home-written IDL program, which extracts the fluorescence intensity trajectories from localized fluorescence spots individually across the entire movie. The intensity of each bright spot in an image is obtained by integrating the signal counts over an area of ~1 × 1 μm2. A flow cell, 100 μm (height) × 2 cm (length) × 5 mm (width), formed by double-sided tapes sandwiched between a quartz slide (Technical Glass or Finkenbeiner) and a borosilicate coverslip (Gold Seal), was used to hold aqueous sample solutions for single-molecule fluorescence measurements. A volume of 100 μl of 1 nM colloidal hollow SiO2 or SiO2/Dye complex solution was spin-coated onto quartz slide, and then rinsed several times with MilliQ water to remove unbound nanoparticles. On the quartz slide, two holes were drilled to connect to polyethylene tubing and a syringe pump for continuous solution flow at 10 μl min−1.
Quantum chemical calculations
Density functional theory (DFT) method at M062X/6-31+G* level has been carried out to calculate the stationary point with frequency computations. The total energy has been corrected by higher accurate second-order Møller-Plesset perturbation theory (MP2) level using 6-311++G** basis set with solvation effect. The solvent effect in water was employed by using SMD solvent method, which can be more accurate for charged species than other solvent models. All the reactant, intermediates and product have been confirmed to be the minimum with no imaginary frequencies, and each transition state had only one imaginary frequency.
Additional information
How to cite this article: Zhang, Y. et al. Single-molecule chemical reaction reveals molecular reaction kinetics and dynamics. Nat. Commun. 5:4238 doi: 10.1038/ncomms5238 (2014).
References
Somorjai, G. A. & Li, Y. Introduction to Surface Chemistry and Catalysis Wiley-Interscience (2010).
Ovchinnikov, A. A. & Zeldovich, Y. B. Role of density fluctuations in bimolecular reaction kinetics. Chem. Phys. 28, 215–218 (1978).
Dai, D. et al. Interference of quantized transition-state pathways in the H+D2→D+HD chemical reaction. Science 300, 1730–1734 (2003).
Strazisar, B., Lin, C. & Davis, H. F. Mode specific energy disposal in the 4-atom reaction OH+D2 -> HOD+D. Science 290, 958–961 (2000).
Nuernberger, P., Vogt, G., Brixner, T. & Gerber, G. Femtosecond quantum control of molecular dynamics in the condensed phase. Phys. Chem. Chem. Phys. 9, 2470 (2007).
Burnham, C. J., Petersen, M. K. & Voth, G. A. Modeling condensed-phase chemistry through molecular dynamics simulation. Comput. Sci. Eng. 5, 31–35 (2003).
Dickson, R. M., Norris, D. J., Tzeng, Y.-L. & Moerner, W. E. Three-dimensional imaging of single molecules solvated in pores of poly(acrylamide) gels. Science 274, 966–968 (1996).
Xu, X.-H. & Yeung, E. S. Direct measurement of single-molecule diffusion and photodecomposition in free solution. Science 275, 1106–1109 (1997).
Ha, T., Glass, J., Enderle, T., Chemla, D. S. & Weiss, S. Hindered rotational diffusion and rotational jumps of single molecules. Phys. Rev. Lett. 80, 2093–2096 (1998).
Lu, H. P. & Xie, X. S. Single-molecule spectral fluctuations at room temperature. Nature 143, 143–146 (1998).
Wennmalm, S., Edman, L. & Rigler, R. Conformational fluctuations in single DNA molecules. Proc. Natl Acad. Sci. USA 94, 10641–10646 (1997).
Cao, J. Michaelis-Menten equation and detailed balance in enzymatic networks. J. Phys. Chem. B 115, 5493–5498 (2011).
Cao, J. Event-averaged measurements of single-molecule kinetics. Chem. Phys. Lett. 327, 38–44 (2000).
Lu, H. P., Xun, L. Y. & Xie, X. S. Single-molecule enzymatic dynamics. Science 282, 1877–1882 (1998).
Xu, W., Kong, J. S., Yeh, Y.-T. E. & Chen, P. Single-molecule nanocatalysis reveals heterogeneous reaction pathways and catalytic dynamics. Nat. Mater. 7, 992–996 (2008).
Wang, Y., Wang, X., Ghosh, S. K. & Lu, H. P. Probing single-molecule interfacial electron transfer dynamics of porphyrin on TiO2 nanoparticles. J. Am. Chem. Soc. 131, 1479–1487 (2009).
Tachikawa, T., Wang, N., Yamashita, S., Cui, S.-C. & Majima, T. Design of a highly sensitive fluorescent probe for interfacial electron transfer on a TiO2 surface. Angew. Chem. Int. Ed. 49, 8593–8597 (2010).
Rybina, A. et al. Distinguishing alternative reaction pathways by single-molecule fluorescence spectroscopy. Angew. Chem. Int. Ed. 52, 6322–6325 (2013).
Macklin, J. J., Trautman, J. K., Harris, T. D. & Brus, L. E. Imaging and time-resolved spectroscopy of single molecules at an interface. Science 272, 255–258 (1996).
Zhao, B., Ranguelova, K., Jiang, J. J. & Mason, R. P. Studies on the photosensitized reduction of resorufin and implications for the detection of oxidative stress with Amplex Red. Free Radic. Biol. Med. 51, 153–159 (2011).
Vogelsang, J. et al. A reducing and oxidizing system minimizes photobleaching and blinking of fluorescent dyes. Angew. Chem. Int. Ed. 47, 5465–5469 (2008).
Kundu, K. et al. Hydrocyanines: a class of fluorescent sensors that can image reactive oxygen species in cell culture, tissue, and in vivo. Angew. Chem. 121, 305–309 (2009).
Schwering, M. et al. Far-field nanoscopy with reversible chemical reactions. Angew. Chem. Int. Ed. 50, 2940–2945 (2011).
Fang, X., Chen, C., Liu, Z., Liu, P. & Zheng, N. A cationic surfactant assisted selective etching strategy to hollow mesoporous silica spheres. Nanoscale 3, 1632–1639 (2011).
Xu, W., Jain, P., Beberwyck, B. & Alivisatos, A. P. Redox photocatalysis of trapped electrons and holes on single Sb-doped Titania nanorod surfaces. J. Am. Chem. Soc. 134, 3946–3949 (2012).
Blackburn, L. & Taylor, R. J. K. In situ oxidation−imine formation−reduction routes from alcohols to amines. Org. Lett. 3, 1637–1639 (2001).
Allen, A. E. & MacMillan, D. W. C. Enantioselective α-arylation of aldehydes via the productive merger of iodonium salts and organocatalysis. J. Am. Chem. Soc. 133, 4260–4263 (2011).
Ibrahem, I. & Córdova, A. Direct catalytic intermolecular α-allylic alkylation of aldehydes by combination of transition-metal and organocatalysis. Angew. Chem. Int. Ed. 45, 1952–1956 (2006).
Suzuki, Y., Kaneno, D. & Tomoda, S. Theoretical study on the mechanism and diastereoselectivity of NaBH4 reduction. J. Phys. Chem. A 113, 2578–2583 (2009).
Balvers, W. G., Boersma, M. G., Vervoort, J. & Rietjens, I. M. C. M. Experimental and theoretical study on the eedox cycling of resorufin by solubilized and membrane-bound NADPH-cytochrome reductase. Chem. Res. Toxicol. 5, 268–273 (1992).
Xu, W., Kong, J. S. & Chen, P. Single-molecule kinetic theory of heterogeneous and enzyme catalysis. J. Phys. Chem. C 113, 2393–2404 (2009).
Petrucci, R. H., Harwood, W. S. & Herring, F. G. General Chemistry 8th edn Prentice-Hall (2002).
Cao, J. & Silbey, R. J. Generic Schemes for single-molecule kinetics. 1: self-consistent pathway solutions for renewal processes. J. Phys. Chem. B 112, 12867–12880 (2008).
Chaudhury, S., Cao, J. & Sinitsyn, N. A. Universality of Poisson indicator and fano factor of transport event statistics in ion channels and enzyme kinetics. J. Phys. Chem. B 117, 503–509 (2013).
Witkoskie, J. B. & Cao, J. Single molecule kinetics. I. Theoretical analysis of indicators. J. Chem. Phys. 121, 6361–6372 (2004).
Pophristic, V. & Goodman, L. Hyperconjugation not steric repulsion leads to the staggered structure of ethane. Nature 411, 565–568 (2001).
Weinhold, F. Chemistry. A new twist on molecular shape. Nature 411, 539–541 (2001).
Chubykalo, O. et al. Monte Carlo technique with a quantified time step: application to the motion of magnetic moments. Phys. Rev. B 67, 064422–064431 (2003).
Smirnov-Rueda, R., Chubykalo, O., Nowak, U., Chantrell, E. W. & Gonzalez, J. M. Real time quantification of Monte Carlo steps for different time scales. J. Appl. Phys. 87, 4798–4800 (2000).
Zhou, X. et al. Quantitative super-resolution imaging uncovers reactivity patterns on single nanocatalysts. Nat. Nanotechnol 7, 237–241 (2012).
Roeffaers, M. B. J. et al. Super-resolution reactivity mapping of nanostructured catalyst particles. Angew. Chem. Int. Ed. 48, 9285–9289 (2009).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Bates, M., Huang, B., Dempsey, G. T. & Zhuang, X. Multicolor super-resolution imaging with photo-switchable fluorescent probes. Science 317, 1749–1753 (2007).
Einolghozati, A., Sardari, M., Beirami, A. & Fekri, F. Capacity of discrete molecular diffusion channels. arXiv:1105.1969v1.
Pierobon, M. & Akyildiz, I. F. Capacity of a diffusion-based molecular communication system with channel memory and molecular noise. IEEE Trans. Inf. Theory 59, 942–954 (2013).
Acknowledgements
This study was funded by the National Natural Science Foundation of China (21273220 and 21303180), the National Basic Research Program of China (973 Program, 2012CB932800 and 2014CB932700), the Strategic Priority Research Program of CAS (XDA0903104), the National High Technology Research and Development Program of China (863 Program, 2012AA053401), and ‘the Recruitment Program of Global youth Experts’ of China. The computational support is appreciated from Dr Ke-Li Han from Dalian institute of Chemical Physics, Chinese Academy of Sciences. Dr Yutian Zhu is appreciated for the help of Monte-Carlo simulation.
Author information
Authors and Affiliations
Contributions
W.X. conceived and coordinated the research. Y.Z. contributed to ensemble and single-molecule experiments and data analysis. P.S. contributed to the theoretical calculation and data analysis. Q.F. and M.R. contributed to the synthesis of nanoparticles. The manuscript was primarily written by W.X, Y.Z. and P.S. All authors contributed to discussions and manuscript review.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-13, Supplementary Tables 1-2, Supplementary Methods and Supplementary References (PDF 1889 kb)
Rights and permissions
About this article
Cite this article
Zhang, Y., Song, P., Fu, Q. et al. Single-molecule chemical reaction reveals molecular reaction kinetics and dynamics. Nat Commun 5, 4238 (2014). https://doi.org/10.1038/ncomms5238
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms5238
This article is cited by
-
Reactions in single-molecule junctions
Nature Reviews Materials (2022)
-
Evolution of hidden localized flow during glass-to-liquid transition in metallic glass
Nature Communications (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.