Abstract
Polyploidization has provided much genetic variation for plant adaptive evolution, but the mechanisms by which the molecular evolution of polyploid genomes establishes genetic architecture underlying species differentiation are unclear. Brassica is an ideal model to increase knowledge of polyploid evolution. Here we describe a draft genome sequence of Brassica oleracea, comparing it with that of its sister species B. rapa to reveal numerous chromosome rearrangements and asymmetrical gene loss in duplicated genomic blocks, asymmetrical amplification of transposable elements, differential gene co-retention for specific pathways and variation in gene expression, including alternative splicing, among a large number of paralogous and orthologous genes. Genes related to the production of anticancer phytochemicals and morphological variations illustrate consequences of genome duplication and gene divergence, imparting biochemical and morphological variation to B. oleracea. This study provides insights into Brassica genome evolution and will underpin research into the many important crops in this genus.
Similar content being viewed by others
Introduction
Brassica oleracea comprises many important vegetable crops including cauliflower, broccoli, cabbages, Brussels sprouts, kohlrabi and kales. The species demonstrates extreme morphological diversity and crop forms, with various members grown for their leaves, flowers and stems. About 76 million tons of Brassica vegetables were produced in 2010, with a value of 14.85 billion dollars ( http://faostat.fao.org/). Most B. oleracea crops are high in protein1 and carotenoids2, and contain diverse glucosinolates (GSLs) that function as unique phytochemicals for plant defence against fungal and bacterial pathogens3 and on consumption have been shown to have potent anticancer properties4,5,6.
B. oleracea is a member of the family Brassicaceae (~\n338 genera and 3,709 species)7 and one of three diploid Brassica species in the classical triangle of U8 that also includes diploids B. rapa (AA) and B. nigra (BB) and allotetraploids B. juncea (AABB), B. napus (AACC) and B. carinata (BBCC). These allotetraploid species are important oilseed crops, accounting for 12% of world edible oil production ( http://faostat.fao.org/). As the origin and relationship between these species is clear, the timing and nature of the evolutionary events associated with Brassica divergence and speciation can be revealed by interspecific genome comparison. Each of the Brassica genomes retains evidence of recursive whole-genome duplication (WGD) events9,10 (Supplementary Fig. 1) and have undergone a Brassiceae-lineage-specific whole-genome triplication (WGT)11,12 since their divergence from the Arabidopsis lineage. These events were followed by diploidization that involved substantial genome reshuffling and gene losses11,12,13,14,15. Because of this, Brassica species are a model for the study of polyploid genome evolution (Supplementary Fig. 2), mechanisms of duplicated gene loss, neo- and sub-functionalization, and associated impact on morphological diversity and species differentiation.
We report a draft genome sequence of B. oleracea and its comprehensive genomic comparison with the genome of sister species B. rapa, which diverged from a common ancestor ~\n4 MYA. These data provide insights into the dynamics of Brassica genome evolution and divergence, and serve as important resources for Brassica vegetable and oilseed crop breeding. Furthermore, this genome will support studies of the large range of morphological variation found within B. oleracea, which includes sexually compatible crops such as cabbages, cauliflower and broccoli that are important for their economic, nutritional and potent anticancer value.
Results
B. oleracea genome assembly and annotation
Complementing the sequencing of the smaller B. rapa genome11, a draft genome assembly of B. oleracea var. capitata line 02–12 was produced by interleaving Illumina, Roche 454 and Sanger sequence data. This assembly represents 85% of the estimated 630 Mb genome, and includes >98% of the gene space (Supplementary Methods, Supplementary Tables 1–3, 7 and 8 and Supplementary Fig. 3). The assembly was anchored to a new genetic map16 to produce nine pseudo-chromosomes that account for 72% of the assembly, and validated by comparison with a B. oleracea physical map17, a high-density B. napus genetic map18 and complete BAC sequences (Supplementary Figs 4–9 and Supplementary Tables 4 and 5). For comparative analyses, identical genome annotation pipelines were used for annotation of protein-coding genes and transposable elements (TEs) for B. oleracea and B. rapa.
A total of 45,758 protein-coding genes were predicted, with a mean transcript length of 1,761 bp, a mean coding length of 1,037 bp, and a mean of 4.55 exons per gene (Table 1, Supplementary Methods, Supplementary Table 6 and Supplementary Fig. 10), similar to A. thaliana19 and B. rapa11. Publicly available ESTs, together with RNA sequencing (RNA-seq) data generated in this study, support 94% of predicted gene models (Supplementary Tables 7 and 8), and 91.6% of predicted genes have a match in at least one public protein database (Supplementary Tables 9 and 10, and Supplementary Fig. 11). Of the 45,758 predicted genes, 13,032 produce alternative splicing (AS) variants with intron retention and exon skipping (Supplementary Table 11). Genome annotation also predicted 3,756 non-coding RNAs (miRNA, tRNA, rRNA and snRNA) (Supplementary Table 12).
A combination of structure-based analyses and homology-based comparisons resulted in the identification of 13,382 TEs with clearly identified terminal boundaries, including 5,107 retrotransposons and 8,275 DNA transposons (Supplementary Methods, Supplementary Fig. 12 and Supplementary Table 13). These elements together with numerous truncated elements or TE remnants make up 38.80% of the assembled portion of the B. oleracea genome, whereas TEs account for only 21.47% of the B. rapa genome assembly. Copia (11.64%) and gypsy (7.84%) retroelements are the major constituents of the repetitive fraction, and are unevenly distributed across each chromosome, with retrotransposons predominantly found in pericentromeric or heterochromatic regions (Supplementary Fig. 13) in B. oleracea. Tentative physical positions of some of the centromeres were determined based on homologue and phylogenetic analysis of the centromere-specific 76 bp tandem repeats CentBo-1 and CentBo-2 and copia-type retrotransposon (CentCRBo) (Supplementary Table 14 and Supplementary Figs 14–17). The distribution of 45S and 5S rDNA sequences were also visualized by fluorescent in situ hybridization (Supplementary Figs 18 and 19), leading to a predicted karyotype ideogram for B. oleracea (Supplementary Fig. 20). An extra-centromeric locus with colocalized centromeric satellite repeat CentBo-1 and the centromeric retrotransposon CRBo-1 was observed on the long arm of chromosome 6 (Supplementary Figs 18–20). A comprehensive database for the genome information is accessible at http://www.ocri-genomics.org/bolbase/index.html.
Conserved syntenic blocks and genome rearrangement after WGT
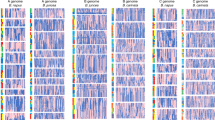
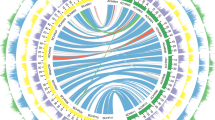
The relatively complete triplicated regions in B. oleracea and B. rapa were constructed and they relate to the 24 ancestral crucifer blocks (A–X) in A. thaliana20. Further the triplicated blocks resulting from WGT in the two Brassica species were partitioned into three subgenomes: LF (Least-fractionated), MF1 (Medium-fractionated) and MF2 (Most-fractionated)11 (Fig. 1a, Supplementary Methods, Supplementary Tables 15 and 16, and Supplementary Figs 21–26). These syntenic blocks occupy the majority of the genome assemblies of A. thaliana (19,628 genes, 72.24% of 27,169 genes), B. oleracea (26,485 genes, 57.88%) and B. rapa (26,698 genes, 64.84%), and provide a foundation for comparative analyses of chromosomal rearrangement, gene loss and divergence of retained paralogues after WGT. Massive gene loss occurred in an asymmetrical and reciprocal fashion in the three subgenomes of each species and was largely completed before the B. oleracea–B. rapa divergence (Fig. 1c, Supplementary Tables 17–19 and Supplementary Figs 25–27). The timing of this evolutionary process was supported by the estimated timing of WGT ~\n15.9 million years ago (MYA), and species divergence ~\n4.6 MYA, based on synonymous substitution (Ks) rates of genes located in the blocks (Fig. 1b and Supplementary Table 20). Gene loss occurred mainly through small deletions that may be caused by illegitimate recombination21,22 (Supplementary Fig. 27), consistent with observations in other plant genomes.
(a) Segmental colinearity of the genomes of B. oleracea, B. rapa and A. thaliana. Syntenic blocks are defined and labelled from A to X (coloured) previously reported in A. thaliana20. (b) Time estimate of WGD and subsequent two Brassica species divergence. (c) Pattern of retention/loss of orthologous genes on each set of three subgenomic (LF, MF1 and MF2) blocks of B. oleracea and B. rapa corresponding to A. thaliana A to X blocks. The x axis denotes the physical position of each A. thaliana gene locus. The y axis denotes the proportion of orthologous genes retained in the B. oleracea and B. rapa subgenomic blocks around each A. thaliana gene, where 500 genes flanking each side of a certain gene locus were analysed, giving a total window size of 1,001 genes.
Abundant genome rearrangement following WGT and subsequent Brassica species divergence resulted in complex mosaics of triplicated ancestral genomic blocks in the A and C genomes (Fig. 1a and Supplementary Fig. 28). At least 19 major, and numerous fine-scale, chromosome rearrangements occurred, which differentiate the two Brassica species (Supplementary Fig. 29). This is in agreement with previous comparative studies based on chromosome painting12,23 and genetic mapping24,25. The extensive chromosome reshuffling in Brassica is in contrast to that observed in other taxa, such as the highly syntenic tomato–potato and pear–apple genomes, each with longer divergence times and less genome rearrangement26,27. This difference may be a consequence of mesopolyploidy in Brassica.
Greater TEs accumulation in B. oleracea than B. rapa
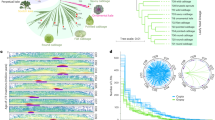
Both retro- (22.13%) and DNA (16.67%) TEs appear to be greater amplified in B. oleracea relative to B. rapa (9.43 and 12.04%) (Fig. 2a and Supplementary Table 13). We constructed 1,362 gap-free contig-contig syntenic regions by clustering orthologous B. rapa—B. oleracea genes using MCscan (Supplementary Figs 29 and 30). The B. oleracea TE length (34.03% of the 259.6M) is 3.4 times greater than that of the syntenic B. rapa regions (16.73% of the 155.0M) (Fig. 2c, Supplementary Tables 21 and 22, and Supplementary Fig. 31). Phylogenetic analysis revealed that B. oleracea has both more LTR retrotransposon (LTR-RT) families, and more members in most families than B. rapa (Fig. 2d and Supplementary Figs 12, 32 and 33). Furthermore, two new lineages of LTR-RTs, Brassica Copia Retrotransposon and Brassica Gypsy Retrotransposon, were defined in both Brassica species (Supplementary Fig. 33). Analysis of LTR insertion time revealed that ~\n98% of B. oleracea intact LTR-RTs amplified continuously over the ~\n4 million years (MY) since the B. oleracea–B. rapa split, whereas ~\n68% of B. rapa intact LTR-RTs amplified rapidly within the last 1 MY, predominantly in the recent 0.2 MY (Fig. 2b and Supplementary Fig. 34). Hence, LTR-RTs expanded more in the intergenic space of euchromatic regions in B. oleracea than B. rapa. This agrees with previous observations based on comparison of BAC sequences between the A and C genomes28. As a consequence of continuous TE amplification over the last 4 MY, the genome size of B. oleracea is ~\n30% larger than that of B. rapa although the two genomes share the same ploidy and are largely collinear.
(a) TE copy number and total length in each assembly and B. oleracea–B. rapa syntenic blocks. (b) The number of intact LTR (Copia-like and Gypsy-like) birthed at different times (million years ago, MYA) in the syntenic regions of B. oleracea and B. rapa. (c) The comparison of TE distribution and composition in B. oleracea–B. rapa syntenic blocks along B. oleracea chromosomes. We divided B. oleracea–B. rapa syntenic region into non-overlapping sliding 200 kb windows to compare TE contents. For each window, the ratio log10(B. oleracea/B. rapa) was calculated for total syntenic block length (blue line), LTR length (purple line), gene length (yellow point), exons length (red point) and intron length (green point). If B. oleracea > B. rapa in absolute length of TE composition in a compared window, the dot or line is above the line y=0. The corresponding B. rapa chromosome segments along B. oleracea C08 were indicated by coloured bars. All other B. oleracea chromosomes are showed in Supplementary Fig. 31. (d) Phylogeny of the Copia-like elements as an example of LTR-RTs of the syntenic regions in B. rapa and B. oleracea. The neighbor-joining (NJ) trees were generated based on the conserved RT domain nucleotide sequences using the Kimura two-parameter method68 in MEGA4 (ref. 69).
Species-specific genes and tandemly duplicated genes
While the genomes of B. oleracea and B. rapa are highly similar in terms of total gene clusters/sequences and the gene number in each cluster, there are also a large number of species-specific genes in the two species. A total of 66.5% (34,237 genes) of B. oleracea genes and 74.9% (34,324) of B. rapa genes were clustered into OrthoMCL groups (Supplementary Table 23 and Supplementary Fig. 35). We identified 9,832 B. oleracea-specific and 5,735 B. rapa-specific genes, of which 77% were supported by gene expression and/or a clear Arabidopsis homologue (Supplementary Table 24). Of them, >90% of these specific genes were validated for their absence in the counterpart genomes by reciprocal mapping of raw clean reads (Supplementary Tables 25 and 26). Most Brassica-specific genes are randomly distributed along the chromosomes (Supplementary Figs 36 and 37). More than 80% of the species-specific genes were surrounded by non-specific genes (Supplementary Fig. 38), suggesting that deletion of individual genes may be the major mechanism underlying gene loss and the difference in gene numbers between B. oleracea and B. rapa.
Tandem duplication produces clusters of duplicated genes and contributes to the expansion of gene families29. We identified 1,825, 2,111 and 1,554 gene clusters containing 4,365, 5,181 and 4,170 tandemly duplicated genes in B. oleracea, B. rapa and A. thaliana, respectively (Fig. 3a, Supplementary Tables 27 and 28 and Supplementary Fig. 39). The wide range of sequence divergence of tandem gene pairs in each species suggests that tandem gene duplication occurred continuously throughout the evolutionary history of these species, rather than in discrete bursts (Supplementary Figs 40 and 41). Their continuous and asymmetrical occurrence after species divergence resulted in 522, 697 and 815 species-specific tandem clusters in the three genomes. The frequency of tandem duplication is independent of the total gene content, suggesting that genome triplication has not inhibited its occurrence. Tandemly duplicated genes are preferentially enriched for gene ontology (GO) categories related to defence response and pathways related to secondary metabolism such as indole alkaloid biosynthesis and tropane, piperidine and pyridine alkaloid biosynthesis (Fig. 3b, Supplementary Tables 29–32 and Supplementary Fig. 42). Over 44.0 and 51.9% of the NBS-encoding resistance genes are tandemly duplicated in B. oleracea and B. rapa, respectively (Supplementary Table 33).
(a) A Venn diagram showing shared and specific tandem duplication events in A. thaliana, B. rapa and B. oleracea. (b,c) Distribution of tandem genes and WGT/WGD-derived paralogues in the KEGG pathway maps in B. oleracea (bol), B. rapa (bra) and A. thaliana (ath). For each KEGG pathway map, the proportion of the number of duplicated genes or paralogues to the total genes was calculated (x axis) and the number of maps whose tandem gene proportion fell in a range was shown on the y axis. (d) Oxidative phosphorylation pathway enriched by WGT-derived paralogous genes in the Brassica genomes. The gene copy number for each KO enzyme in B. oleracea, B. rapa and A. thaliana were shown (dash-connected) under the KO enzyme number.
Biased loss and retention of genes after WGT/WGD
Following polyploidization, reversion of gene numbers towards diploid levels through gene loss has been widely observed in plants30. However, in Brassica this only appears to be true for collinear genes in the conserved syntenic regions, with a loss of ~\n60% of the predicted post-triplication gene set, nearly restoring the pre-triplication gene number. This is reflected in an overall retention rate of 1.2-fold of A. thaliana orthologous genes in corresponding syntenic regions (Fig. 1c and Supplementary Table 18). In contrast, in terms of genes that have no collinear gene in A. thaliana and either Brassica species (hereafter called non-collinear genes), gene retention rates is 2.5-fold the A. thaliana gene number in B. oleracea and 1.9-fold in B. rapa, both significantly higher than the expected rates (P value <2.2e–16;Supplementary Table 34). For these retained genes, the numbers of the genes that are common in the two Brassica species are 11,746 in B. oleracea and 10,411 in B. rapa. Most of these genes are supported by expression and/or the presence of an Arabidopsis homologue (Supplementary Table 35). More than 61% of these genes have homologues present as collinear genes and 16% also are homologous to other non-collinear genes, indicating gene movement from triplicated syntenic regions and being similar to observations in A. thaliana, where half of the genes are nonsyntenic within rosids31. This suggests that the breakdown of the triplicated syntenic relationship has not only prevented gene loss and a move towards pre-triplication gene numbers but has also maintained a higher gene density, and thus maintained WGT-derived genes for species evolution.
The presence of a large number of the retained paralogous genes in the syntenic regions led us to examine whether genes in some functional categories have preferentially been over-retained, as observed in other plants29. The results indicate that WGT-produced paralogous genes are over-retained in GO categories associated with regulation of metabolic and biosynthetic processes, RNA metabolism and transcription factors (Supplementary Table 36 and Supplementary Figs 43–45), and the two Brassica species exhibit similar patterns of gene category retention. From a study of KEGG pathways, we also found that WGT-produced Brassica paralogous genes contribute 40–60% of total genes for 90% of KEGG pathways (Fig. 3c and Supplementary Fig. 43), and are functionally enriched in primary or core metabolic processes such as oxidative phosphorylation, carbon fixation, photosynthesis, circadian rhythm32 and lipid metabolism (Supplementary Tables 36 and 37 and Supplementary Figs 43–45). Notably, the pathways associated with energy metabolism have been enhanced in both Brassica species. For instance, in the oxidative phosphorylation pathway, there are 161 genes in A. thaliana, but 241 in B. oleracea and 208 in B. rapa. The majority (143/241 and 142/208) of these Brassica genes are multiple paralogues residing in the triplicated syntenic regions, and more than half of these paralogues have been retained as three copies, significantly higher than observed for other genes in the triplication regions (Fig. 3d and Supplementary Fig. 43).
Phylogenetic analyses show that WGT led to an expansion of genes involved in auxin functioning (AUX, IAA, GH3, PIN, SAUR, TAA, TIR, TPL and YUCCA), morphology specification (TCP), and flowering time control (FLC, CO, VRN1, LFY, AP1 and GI) (Supplementary Table 38 and Supplementary Figs 46–61), and that most Arabidopsis genes in these families have two or three orthologs in Brassica species. These WGT-produced duplicated genes may provide important sources of evolutionary innovation33 and contribute to the extreme morphological diversity in Brassica species.
Divergence of duplicated genes in the Brassica genomes
The largest genetic foundation for plant genome evolution and new species formation is the differentiation of retained paralogous and orthologous genes. Around 38% (4,302/11,493) of all paralogous gene pairs in B. oleracea and ~\n36% (4,089/11,448) in B. rapa have different predicted exon numbers (Supplementary Data 1, Supplementary Tables 39 and 40 and Supplementary Fig. 62). There are 6,571 orthologous gene pairs with different exon numbers, accounting for 27.6% of total gene pairs (23,823). Some paralogous or orthologous pairs have high Ks values and low sequence similarity (Supplementary Fig. 63), indicating sequence differentiation. Of these paralogous genes, some offer appreciable opportunity for non-reciprocal DNA exchanges (gene conversion). About 8% of the 4,296 homologous quartets in B. rapa and B. oleracea have been affected by gene conversion (Fig. 4a, Supplementary Table 41 and Supplementary Fig. 64) and about one-sixth (53) of converted genes were inferred to have experienced independent conversion events in both Brassica species, a parallelism sometimes observed in other plants11,34. Around 40–44% of conversion events involved paralogues in the less-fractionated subgenomes LF in both species, substantially higher than the other two subgenomes (Supplementary Table 41). This finding suggests that gene conversion is related to homologous gene density, which determines the likelihood of illegitimate recombination.
(a) Genome-wide gene conversion in B. oleracea. The conversion in B. rapa is showed in Supplementary Fig. 64. (b) The ratio of differentially expressed duplicated gene pairs derived from different duplications: alpha whole-genome duplication (α-WGD), Brassiceae-lineage WGT, tandem duplication (TD). Bol, B. oleracea; Bra, B. rapa. C: callus; R: root; St: stem; L: leaf; F: flower; Si: silique. The differentially expressed duplicated gene pairs were defined as fold change >2 and false discovery rate (FDR) <0.05 or gene pair where expression was detected for only one gene within gene pairs (FDR <0.05). (c) Box and whisker plots for differentiated expression for three subgenomes (LF, MF1 and MF2) in flower tissue of B. oleracea and B. rapa. For the other tissues, see Supplementary Fig. 67. (d) The duplicated gene pairs belonging to transcription factors (TFs) and its related GO terms contain a significantly lower ratio of differentially expressed duplicated gene pairs than the average at the genome-wide level in leaf (values given) and other tissues (values not presented) (Supplementary Table 45). (e) The GO terms (left) in which the duplicated gene pairs contain a significantly higher ratio of differentially expressed duplicated gene pairs than the average ratio at the genome-wide level in leaf and other tissues (Supplementary Table 46). Values from one tissue were presented and the other tissues were indicated with abbreviated letters to the right if expression in these tissues is significantly higher. (f) Expression variation caused by divergence (either different variants or differential expression of the same variants) of alternative splicing (AS) variants in WGT paralogous gene pairs with identical numbers of exons and in Bol–Bra orthologous gene pairs. IRES denotes types of intron retention and exon skipping.
Analysis of RNA-seq data generated from callus, root, leaf, stem, flower and silique of B. oleracea and B. rapa suggests that >40% of WGT paralogous gene pairs are differentially expressed in these species (Fig. 4b and Supplementary Fig. 65), suggesting potential subfunctionalization of these genes. In both species, a general trend of expression differentiation was alpha-WGD paralogous genes (~\n46%) > WGT paralogous genes (~\n42%) > tandemly duplicated genes (~\n35%) (Fig. 4b, Supplementary Fig. 66 and Supplementary Tables 42 and 43). Different tissues harbour approximately the same number of differentially expressed duplicates, but this number was slightly higher in flower tissue. The expression level of genes in the LF subgenome was significantly higher than corresponding syntenic genes in the more fractionated subgenomes (MF1 and MF2) while no expression dominance relationship was observed between the subgenomes MF1 and MF2 (Fig. 4c, Supplementary Table 44 and Supplementary Fig. 67). Duplicated transcription factor gene pairs showed less differentiated expression (~\n38%) than the expected ratio at the genome-wide level (Fig. 4d and Supplementary Table 45), while paralogues with GO categories related to membrane, catalytic activity and defence response exhibited a higher ratio of differentiated expression (Fig. 4e and Supplementary Table 46). Of B. oleracea–B. rapa orthologous gene pairs (23,823 in total), ~\n42% were differentially expressed across all tissues (Supplementary Tables 42 and 43).
Furthermore, many paralogues generate different transcripts, resulting in expression differentiation. Analysis of AS variants of paralogous gene pairs that have identical numbers of exons demonstrated that these variants (either different variants or differential expression of the same variants) cause >20% and >44% of such paralogous genes to be differentially expressed in B. oleracea and B. rapa, respectively (Fig. 4f and Supplementary Table 47). For orthologous gene pairs of B. oleracea and B. rapa, 35.5% (8,467) of gene pairs showed differential expression due to AS variation. When only counting intron retention and exon skipping, 9.3% (2,215) of gene pairs differ. Divergence in AS variants of gene pairs presents an important layer of gene regulation, as reported35,36,37,38, and thus provides a genetic basis for species evolution and new species formation.
Unique GSLs metabolism pathways
GSLs and hydrolysis products have been of long-standing interest due to their role in plant defence and anticancer properties. Compared with B. rapa and B. napus, B. oleracea has the greatest GSL profile diversity, with wide qualitative and quantitative variation39,40. We identified 101 and 105 GSL biosynthesis genes in B. rapa and B. oleracea, respectively, and 22 GSL catabolism genes in each species (Fig. 5a, Supplementary Table 48 and Supplementary Data 2). In the GSL biosynthesis and catabolism pathways, tandem genes (41.4%, 40.7% and 33.9% in A. thaliana, B. oleracea and B. rapa, respectively) were present in a much higher proportion than the genome-wide average (Supplementary Table 32). The observed variation of GSL profiles is mainly attributed to the duplication of two genes, methylthioalkylmalate (MAM) synthase and 2-oxoglutarate-dependent dioxygenase (AOP).
(a) Aliphatic and indolic GSL biosynthesis and catabolism pathways in A. thaliana, B. oleracea and B. rapa. The copy number of GSL biosynthetic genes in A. thaliana, B. rapa and B. oleracea are listed in square brackets, respectively. Potential anticancer substances/precursors are highlighted in blue bold. Two important amino acid chain elongation and side-chain modification loci MAMs and AOP2 are highlighted in red bold, within the number in the green bracket representing the number of non-functional genes. (b,c) The neighbour-joining (NJ) trees of MAM and AOP genes were generated based on the aligned coding sequences and 100 bootstrap repeats. The silenced genes are indicated by red hollow circle, expressed functional genes are represented by red solid disc and green rectangle. In A. thaliana ecotype Columbia there are just MAM1 and MAM3. (d) Three B. oleracea AOP2 loci among which are one functional AOP2 and two mutated AOP2. 1MOI3M: 1-methoxyindol-3-ylmethyl GSL; 1OHI3M: 1-hydroxyindol-3-ylmethyl GSL; 3MSOP: 3-methylsulfinylpropyl GSL; 3MTP: 3-methylthiopropyl GSL; 3PREY: 2-Propenyl GSL; 4BTEY: 3-butenyl GSL; 4MOI3M: 4-methoxyindol-3-ylmethyl GSL; 4OHB, 4-hydroxybutyl GSL; 4OHI3M: 4-hydroxyindol-3-ylmethyl GSL; 4MSOB: 4-methylsulfinylbutyl GSL; 4MTB, 4-methylthiobutyl GSL; AITC: allyl isothiocyanate; I3C: indole-3-carbinol; I3M: indolyl-3-methyl GSL; DIM: 3,3′-diindolymethane; MAM: methylthioalkylmalate; AOP: 2-oxoglutarate-dependent dioxygenase.
In Arabidopsis, the MAM family contains three tandemly duplicated and functionally diverse members (MAM1, MAM2 and MAM3), and functional analysis demonstrated that MAM2 (absent in ecotype Columbia) and MAM1 catalyses the condensation reaction of the first and the first two elongation cycles for the synthesis of dominant 3 and 4 carbon (C) side-chain aliphatic GSLs, respectively40,41, while MAM3 is assumed to contribute to the production of all GSL chain lengths42. In B. rapa and B. oleracea, MAM1/MAM2 genes experienced independent tandem duplication to produce 6 and 5 orthologs respectively (Fig. 5b,c). The main GSLs in B. oleracea are 4C and 3C GSLs (progoitrin, gluconapin, glucoraphanin and sinigrin)43, while those in B. rapa are 4C and 5C GSLs (gluconapin and glucobrassicanapin)39 (Fig. 5a). Based on the results of expression and phylogenetic analyses, we found a pair of genes Bol017070 and Bra013007, which are the only orthologous genes showing high expression in B. oleracea but silenced in B. rapa (Fig. 5a). This expression difference most likely leads to greater accumulation of the 3C GSL anticancer precursor sinigrin in B. oleracea. Meanwhile, the expression level of MAM3 in B. rapa is much higher than in B. oleracea, explaining the accumulation of 5C GSL glucobrassicanapin in B. rapa. Other genes affecting specific anticancer GLS products are AOPs. Previously, research has reported four gene loci involved in the side-chain modifications of aliphatic GSLs in Arabidopsis. Two tandemly duplicated genes AOP2 and AOP3 catalyse the formation of alkenyl and hydroxyalkyl GSLs, respectively. When both AOPs are non-functional, the plant accumulates the precursor methylsulfinyl alkyl GSL. We identified three AOP2 genes in B. oleracea (Fig. 5d), but two are non-functional due to the presence of premature stop codons. In contrast, all three AOP2 copies are functional in B. rapa44. No AOP3 homologue has been identified in Brassica. This analysis supports GSL content surveys and explains why glucoraphanin is abundant in B. oleracea, but not in B. rapa.
Discussion
The Brassica genomes experienced WGT11,12,25 followed by massive gene loss and frequent reshuffling of triplicated genomic blocks. Analysis of retained or lost genes following triplication identified over-retention of genes for metabolic pathways such as oxidative phosphorylation, carbon fixation, photosynthesis and circadian rhythm32, which may contribute to polyploid vigour45. Fewer lost genes were observed in the less-fractionated subgenome, possibly due to expression dominance as reported in maize46.
Gene expression analysis revealed extensive divergence and AS variants between duplicate genes. This subfunctionalization or neofunctionalization of duplicated genes provides genetic novelty and a basis for species evolution and new species formation. For example, TF genes that are considered to be conserved still have more than 38% of paralogous pairs showing differential expression across tissues although this percentage is lower than the average from all duplicated genes. Gene expression variation may contribute to an increased complexity of regulatory networks after polyploidization.
The multi-layered asymmetrical evolution of the Brassica genomes revealed in this study suggests mechanisms of polyploid genome evolution underlying speciation. Asymmetrical gene loss between the Brassica subgenomes, the asymmetrical amplification of TEs and tandem duplications, preferential enrichment of genes for certain pathways or functional categories, and divergence in DNA sequence and expression, including alternative splicing among a large number of paralogous and orthologous genes, together shape a route for genome evolution after polyploidization. A molecular model of polyploid genome evolution through these asymmetrical mechanisms is summarized in Supplementary Fig. 2. The additional information of accessible large datasets and resource was provided in Supplementary Table 49.
In summary, the B. oleracea genomic sequence, its features in comparison with its relatives, and the genome evolution mechanisms revealed, provide a fundamental resource for the genetic improvement of important traits, including components of GSLs for anticancer pharmaceuticals. The genome sequence has also laid a foundation for investigation of the tremendous range of morphological variation in B. oleracea as well as supporting genome analysis of the important allotetraploid crop B. napus (canola or rapeseed).
Methods
Sample preparation and genome sequencing
A B. oleracea sp. capitata homozygous line 02–12 with elite agronomic characters and widely used as a parent in hybrid breeding was used for the reference genome sequencing (Supplementary Methods). The seedlings of plants were collected and genomic DNA was extracted from leaves with a standard CTAB extraction method. Illumina Genome Analyser whole-genome shotgun sequencing combined with GS FLX Titanium sequencing technology was used to achieve a B. oleracea draft genome. We constructed a total of 35 paired-end sequencing libraries with insertion sizes of 180 base pairs (bp), 200 bp, 350 bp, 500 bp, 650 bp, 800 bp, 2 kb, 5 kb, 10 kb and 20 kb following a standard protocol provided by Illumina (Supplementary Methods). Sequencing was performed using Illumina Genome Analyser II according to the manufacturer’s standard protocol.
Genome assembly and validation
We took a series of checking and filtering measures on reads following the Illumina-Pipeline, and low-quality reads, adaptor sequences and duplicates were removed (Supplementary Methods). The reads after the above filtering and correction steps were used to perform assembly including contig construction, scaffold construction and gap filling using SOAPdenovo1.04 ( http://soap.genomics.org.cn/) (Supplementary Methods). Finally, we used 20-kb-span paired-end data generated from the 454 platform and 105-kb-span BAC-end data downloaded from NCBI ( http://www.ncbi.nlm.nih.gov/nucgss?term=BOT01) to extend scaffold length (Supplementary Methods). The B. oleracea genome size was estimated using the distribution curve of 17-mer frequency (Supplementary Methods).
To anchor the assembled scaffolds onto pseudo-chromosomes, we developed a genetic map using a double haploid population with 165 lines derived from a F1 cross between two homozygous lines 02–12 (sequenced) and 0188 (re-sequenced). The genetic map contains 1,227 simple sequence repeat markers and single nucleotide polymorphism markers in nine linkage groups, which span a total of 1,180.2 cM with an average of 0.96 cM between the adjacent loci16. To position these markers to the scaffolds, marker primers were compared with the scaffold sequences using e-PCR (parameters -n2 -g1 –d 400–800), with the best-scoring match chosen in case of multiple matches.
We validated the B. oleracea genome assembly by comparing it with the published physical map constructed using 73,728 BAC clones ( http://lulu.pgml.uga.edu/fpc/WebAGCoL/brassica/WebFPC/)17 and a genetic map from B. napus18 (Supplementary Methods). Eleven Sanger-sequenced B. oleracea BAC sequences were used to assess the assembled genome using MUMmer-3.22 ( http://mummer.sourceforge.net/) (Supplementary Methods).
Gene prediction and annotation
Gene prediction was performed on the genome sequence after pre-masking for TEs (Supplementary Methods). Gene prediction was processed with the following steps: (i) De novo gene prediction used AUGUSTUS47 and GlimmerHMM48 with parameters trained from A. thaliana genes. (ii) For homologue prediction, we mapped the protein sequences from A. thaliana, O. sativa, C. papaya, V. vinifera and P. trichocarpa to the B. oleracea genome using tblastn with an E-value cutoff of 10−5, and used GeneWise (Version 2.2.0)49 for gene annotation. (iii) For EST-aided annotation, the Brassica ESTs from NCBI were aligned to the B. oleracea genome using BLAT (identity ⩾0.95, coverage ⩾0.90) and further assembled using PASA50. Finally, all the predictions were combined using GLEAN51 to produce the consensus gene sets.
Functional annotation of B. oleracea genes was based on comparison with SwissProt, TrEMBL, Interproscan and KEGG proteins databases. The tRNA genes were identified by tRNAscan-SE using default parameters52. Then rRNAs were compared with the genome using blastn. Other non-coding RNAs, including miRNA, snRNA, were identified using INFERNAL53 by comparison with the Rfam database.
TE annotation
LTR-RTs were initially identified using the LTR_STRUC54 programme, and then manually annotated and checked based on structure characteristics and sequence homology. Refined intact elements were then used to identify other intact elements and solo LTRs55. All the LTR-RTs with clear boundaries and insertion sites were classified into superfamilies (Copia-like, Gypsy-like and Unclassified retroelements) and families relying on the internal protein sequence, 5′, 3′ LTRs, primer-binding site and polypurine tracts. Non-LTR-RTs (Long interspersed nuclear element, LINE and Short interspersed nuclear element, SINE) and DNA transposons (Tc1-Mariner, hAT, Mutator, Pong, PIF-Harbinger, CACTA and miniature inverted repeat TE) were identified using conserved protein domains of reverse transposase or transposase as queries to search against the assembled genome using tblastn. Further upstream and downstream sequences of the candidate matches were compared with each other to define their boundaries and structure56. Helitron elements were identified by the HelSearch 1.0 programme57 and manually inspected. All the TE categories were identified according to the criteria described previously58. Typical elements of each category were selected and mixed together as a database for RepeatMasker59 analysis. Around 20 × coverage of shotgun reads randomly sampled from the two Brassica genomes were masked by the same TE data set to confirm the different accumulation of TEs between the two genomes.
Syntenic block construction of B. oleracea and its relatives
We used the same strategy as described in the B. rapa genome paper11 to construct syntenic blocks between species (Supplementary Methods). The all-against-all blastp comparison (E-value ≤ 1e–5) provided the gene pairs for syntenic clustering determined by MCScan (MATCH_SCORE: 50, MATCH_SIZE: 5, GAP_SCORE: –3, E_VALUE: 1E–05). As applied in B. rapa11, we assigned and partitioned multiple B. oleracea or B. rapa chromosomal segments that matched the same A. thaliana segment (‘A to X’ numbering system in A. thaliana22) into three subgenomes: LF, MF1 and MF2.
OrthoMCL clustering
To identify and estimate the number of potential orthologous gene families between B. oleracea, B. rapa, A. thaliana, C. papaya, P. trichocarpa, V. vinifera, S. bicolor and O. sativa, and also between B. oleracea and B. rapa, we applied the OrthoMCL pipeline60 using standard settings (blastp E value <1 × 10−5 and inflation factor =1.5) to compute the all-against-all similarities.
Phylogenetic analysis of gene families
We performed comparative analysis of trait-related gene families. Genes from grape, papaya and Arabidopsis were downloaded from the GenoScope database ( http://www.genoscope.cns.fr/externe/GenomeBrowser/Vitis/), the Hawaii Papaya Genome Project ( http://asgpb.mhpcc.hawaii.edu/papaya/), and the Arabidopsis Information Resource ( http://www.arabidopsis.org/). Previously reported Arabidopsis and Brassica gene sequences were downloaded from TAIR ( http://www.arabidopsis.org/) and BRAD ( http://brassicadb.org/brad/). The protein sequences of the genes were used to determine homologues in grape, papaya, Arabidopsis, B. oleracea and B. rapa by performing blast comparisons with an E-value 1e–10. The Clustal61 programs were used for multiple sequence alignment. Alignment of the small family of GI genes was performed using MEGA562 to conduct neighbour-joining analysis with default parameters and subjected to careful manual checks to remove highly divergent sequences from further analysis. While for other genes, often found in families of tens of genes, the phylogenetic analysis were performed by PhyML63, which can accommodate quite divergent sequences by implementing a maximal likelihood approach with initial analysis based on neighbour-joining method. During these analyses, we constructed trees using both CDS and protein sequence, and the protein-derived tree was used to show the phylogeny if not much incongruity was found. Bootstrapping was performed using 100 repetitive samplings for each gene family. All the inferred trees were displayed using MEGA5 (ref. 62). The multiple sequence alignment of these families was provided as Supplementary Data 3.
Differential expression of duplicated genes across tissues
RNA-seq reads were mapped to their respective locations on the reference genome using Tophat64. Uniquely aligned read counts were calculated for each gene for each tissue sample. We performed the exact conditional test of two Poisson rates on read counts of duplicated genes to test the differential expression of duplicated genes, according to the method applied in soybean65,66. For each duplicated gene pair (for example, genes A and B), read counts and gene length were denoted as Ea and La for gene A, and Eb and Lb for gene B, respectively. The read counts of the genes A and B were assumed to follow the Poisson distributions with rates λA=Ra × La and λB=Rb × Lb. Under the null hypothesis of equal expression of the genes A and B, that is, Ra=Rb, the conditional distribution of Ea given Ea+Eb=k follows a binomial distribution with success probability P=λa/(λa+λb)=La/(La+Lb). The P values were computed and further adjusted to maintain the false discovery rate at 0.05 across gene pairs using the Benjamini–Hochberg method67.
Statistical analysis
The average number of all retained orthologues in the three subgenomes was used to estimate the expected retained gene number in each block, and used together with the observed retained gene number, for the gene retention disparity statistics using the χ2 test. In the GO, IPR (Interproscan) or KEGG enrichment analyses of WGT or tandem genes, the χ2 test (N>5) or the Fisher’s exact test (N≤5) was used to detect significant differences between the proportion of (WGT or tandem) genes observed in each child GO, IPR or KEGG categories, and the expected overall proportion of (WGT or tandem) genes in the whole genome. Correlation of the gene numbers of WGT-derived paralogous genes with tandem genes in 938 GO terms was tested by Pearson correlation coefficients (Supplementary Figure 68). The Benjamini–Hochberg false discovery rate was performed to adjust the P values67.
Additional information
Accession codes: Genome sequence data for B. oleracea have been deposited in the DDBJ/EMBL/GenBank nucleotide core database under the accession code AOIX00000000. Transcriptome sequence data for B. rapa and B. oleracea have been deposited in the DDBJ/EMBL/GenBank Sequence Read Archive (SRA) under the accession codes GSEhttp://www.ncbi.nlm.nih.gov/nuccore/?term=43245 and GSE42891 respectively.
How to cite this article: Liu, S. et al. The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat. Commun. 5:3930 doi: 10.1038/ncomms4930 (2014).
Accession codes
References
U.S. Department of Agriculture, Agricultural Research Service. USDA National Nutrient Database for Standard Reference, Release 26-Vegetables and Vegetable Products. (2013).
Kopsell, D. A. & Kopsell, D. E. Accumulation and bioavailability of dietary carotenoids in vegetable crops. Trends Plant Sci. 11, 499–507 (2006).
Halkier, B. A. & Gershenzon, J. Biology and biochemistry of glucosinolates. Annu. Rev. Plant Biol. 57, 303–333 (2006).
Khwaja, F. S., Wynne, S., Posey, I. & Djakiew, D. 3,3'-diindolylmethane induction of p75NTR-dependent cell death via the p38 mitogen-activated protein kinase pathway in prostate cancer cells. Cancer Prev. Res. (Phila) 2, 566–571 (2009).
Li, Y. et al. Sulforaphane, a dietary component of broccoli/broccoli sprouts, inhibits breast cancer stem cells. Clin. Cancer Res. 16, 2580–2590 (2010).
Higdon, J. V., Delage, B., Williams, D. E. & Dashwood, R. H. Cruciferous vegetables and human cancer risk: epidemiologic evidence and mechanistic basis. Pharmacol Res. 55, 224–236 (2007).
Warwick, S. I., Francis, A. & Al-Shehbaz, I. A. Brassicaceae: species checklist and database on CD-Rom. Pl. Syst. Evol. 259, 249–258 (2006).
Nagaharu, U. Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilication. Jap. J. Bot. 7, 389–452 (1935).
Bowers, J. E., Chapman, B. A., Rong, J. & Paterson, A. H. Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events. Nature 422, 433–438 (2003).
Jiao, Y. et al. Ancestral polyploidy in seed plants and angiosperms. Nature 473, 97–100 (2011).
Wang, X. et al. The genome of the mesopolyploid crop species Brassica rapa. Nat. Genet. 43, 1035–1039 (2011).
Lysak, M. A., Koch, M. A., Pecinka, A. & Schubert, I. Chromosome triplication found across the tribe Brassiceae. Genome Res. 15, 516–525 (2005).
Cheng, F. et al. Deciphering the diploid ancestral genome of the Mesohexaploid Brassica rapa. Plant Cell 25, 1541–1554 (2013).
Town, C. D. et al. Comparative genomics of Brassica oleracea and Arabidopsis thaliana reveal gene loss, fragmentation, and dispersal after polyploidy. Plant Cell 18, 1348–1359 (2006).
Mun, J. H. et al. Genome-wide comparative analysis of the Brassica rapa gene space reveals genome shrinkage and differential loss of duplicated genes after whole genome triplication. Genome Biol. 10, R111 (2009).
Wang, W. et al. Construction and analysis of a high-density genetic linkage map in cabbage (Brassica oleracea L. var. capitata). BMC Genomics 13, 523 (2012).
Wang, X. et al. A physical map of Brassica oleracea shows complexity of chromosomal changes following recursive paleopolyploidizations. BMC Genomics 12, 470 (2011).
Bancroft, I. et al. Dissecting the genome of the polyploid crop oilseed rape by transcriptome sequencing. Nat. Biotechnol. 29, 762–766 (2011).
Arabidopsis Genome and Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Schranz, M. E., Lysak, M. A. & Mitchell-Olds, T. The ABC’s of comparative genomics in the Brassicaceae: building blocks of crucifer genomes. Trends Plant Sci. 11, 535–542 (2006).
Woodhouse, M. R. et al. Following tetraploidy in maize, a short deletion mechanism removed genes preferentially from one of the two homologs. PLoS Biol. 8, e1000409 (2010).
Devos, K. M., Brown, J. K. & Bennetzen, J. L. Genome size reduction through illegitimate recombination counteracts genome expansion in Arabidopsis. Genome Res. 12, 1075–1079 (2002).
Lysak, M. A., Cheung, K., Kitschke, M. & Bures, P. Ancestral chromosomal blocks are triplicated in Brassiceae species with varying chromosome number and genome size. Plant Physiol. 145, 402–410 (2007).
Panjabi, P. et al. Comparative mapping of Brassica juncea and Arabidopsis thaliana using Intron Polymorphism (IP) markers: homoeologous relationships, diversification and evolution of the A, B and C Brassica genomes. BMC Genomics 9, 113 (2008).
Parkin, I. A. et al. Segmental structure of the Brassica napus genome based on comparative analysis with Arabidopsis thaliana. Genetics 171, 765–781 (2005).
Wu, J. et al. The genome of the pear (Pyrus bretschneideri Rehd.). Genome Res. 23, 396–408 (2012).
The Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485, 635–641 (2012).
Cheung, F. et al. Comparative analysis between homoeologous genome segments of Brassica napus and its progenitor species reveals extensive sequence-level divergence. Plant Cell 21, 1912–1928 (2009).
Freeling, M. Bias in plant gene content following different sorts of duplication: tandem, whole-genome, segmental, or by transposition. Annu. Rev. Plant Biol. 60, 433–453 (2009).
Sankoff, D., Zheng, C. & Zhu, Q. The collapse of gene complement following whole genome duplication. BMC Genomics 11, 313 (2010).
Woodhouse, M. R., Tang, H. & Freeling, M. Different gene families in Arabidopsis thaliana transposed in different epochs and at different frequencies throughout the rosids. Plant Cell 23, 4241–4253 (2011).
Lou, P. et al. Preferential retention of circadian clock genes during diploidization following whole genome triplication in Brassica rapa. Plant Cell 24, 2415–2426 (2012).
Doyle, J. J. et al. Evolutionary genetics of genome merger and doubling in plants. Annu. Rev. Genet. 42, 443–461 (2008).
Wang, X., Tang, H. & Paterson, A. H. Seventy million years of concerted evolution of a homoeologous chromosome pair, in parallel, in major Poaceae lineages. Plant Cell 23, 27–37 (2011).
Syed, N. H., Kalyna, M., Marquez, Y., Barta, A. & Brown, J. W. Alternative splicing in plants--coming of age. Trends Plant Sci. 17, 616–623 (2012).
Gabut, M. et al. An alternative splicing switch regulates embryonic stem cell pluripotency and reprogramming. Cell 147, 132–146 (2011).
Zhang, P. G., Huang, S. Z., Pin, A. L. & Adams, K. L. Extensive divergence in alternative splicing patterns after gene and genome duplication during the evolutionary history of Arabidopsis. Mol. Biol. Evol. 27, 1686–1697 (2010).
Filichkin, S. A. et al. Genome-wide mapping of alternative splicing in Arabidopsis thaliana. Genome Res. 20, 45–58 (2010).
Yang, B. & Quiros, C. F. Survey of glucosinolate variation in leaves of Brassica rapa crops. Genet. Res. Crop Evol. 57, 1079–1089 (2010).
Benderoth, M., Pfalz, M. & Kroymann, J. Methylthioalkylmalate synthases: genetics, ecology and evolution. Phytochem. Rev. 8, 255–268 (2009).
Benderoth, M. et al. Positive selection driving diversification in plant secondary metabolism. Proc. Natl. Acad. Sci. USA 103, 9118–9123 (2006).
Textor, S., de Kraker, J. W., Hause, B., Gershenzon, J. & Tokuhisa, J. G. MAM3 catalyses the formation of all aliphatic glucosinolate chain lengths in Arabidopsis. Plant Physiol. 144, 60–71 (2007).
Volden, J. et al. Processing (blanching, boiling, steaming) effects on the content of glucosinolates and antioxidant related parameters in cauliflower (Brassica oleracea L. ssp. botrytis). LWT Food Sci. Technol. 42, 63–73 (2009).
Wang, H. et al. Glucosinolate biosynthetic genes in Brassica rapa. Gene 487, 135–142 (2011).
Chen, Z. J. Molecular mechanisms of polyploidy and hybrid vigour. Trends Plant Sci. 15, 57–71 (2010).
Schnable, J. C., Springer, N. M. & Freeling, M. Differentiation of the maize subgenomes by genome dominance and both ancient and ongoing gene loss. Proc. Natl. Acad. Sci. USA 108, 4069–4074 (2011).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–W312 (2004).
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 20, 2878–2879 (2004).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004).
Xu, Y., Wang, X., Yang, J., Vaynberg, J. & Qin, J. PASA—a program for automated protein NMR backbone signal assignment by pattern-filtering approach. J. Biomol. NMR 34, 41–56 (2006).
Elsik, C. G. et al. Creating a honey bee consensus gene set. Genome Biol. 8, R13 (2007).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
Nawrocki, E. P., Kolbe, D. L. & Eddy, S. R. Infernal 1.0: inference of RNA alignments. Bioinformatics 25, 1335–1337 (2009).
McCarthy, E. M. & McDonald, J. F. LTR_STRUC: a novel search and identification program for LTR retrotransposons. Bioinformatics 19, 362–367 (2003).
Ma, J., Devos, K. M. & Bennetzen, J. L. Analyses of LTR-retrotransposon structures reveal recent and rapid genomic DNA loss in rice. Genome Res. 14, 860–869 (2004).
Holligan, D., Zhang, X., Jiang, N., Pritham, E. J. & Wessler, S. R. The transposable element landscape of the model legume Lotus japonicus. Genetics 174, 2215–2228 (2006).
Yang, L. & Bennetzen, J. L. Structure-based discovery and description of plant and animal Helitrons. Proc. Natl Acad. Sci. USA 106, 12832–12837 (2009).
Wicker, T. et al. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 8, 973–982 (2007).
Smit, A., Hubley, R. & Green, P. RepeatMasker. http://www.repeatmasker.org.
Li, L., Stoeckert, C. J. Jr & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Guindon, S., Delsuc, F., Dufayard, J. F. & Gascuel, O. Estimating maximum likelihood phylogenies with PhyML. Methods Mol. Biol. 537, 113–137 (2009).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Roulin, A. et al. The fate of duplicated genes in a polyploid plant genome. Plant J. 73, 143–153 (2012).
Gu, K., Ng, H. K., Tang, M. L. & Schucany, W. R. Testing the ratio of two poisson rates. Biom. J. 50, 283–298 (2008).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy. Statist. Soc. Ser. 57, 289–300 (1995).
Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120 (1980).
Tamura, K., Dudley, J., Nei, M. & Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596–1599 (2007).
Acknowledgements
This work was supported by the National Basic Research Program of China (2011CB109300, 2012CB113906, 2012CB723007 and 2006CB101600), the National Natural Science Foundation of China (3067134, 30671119 and 31301039), the National High Technology Research and Development Program (2013AA102602, 2012AA100105 and 2012AA100104), the China Agriculture Research System (CARS-13 and CARS-25-A), the Core Research Budget of the Non-profit Governmental Research Institution (1610172010005), the Special Fund for Agro-scientific Research in the Public Interest (201103016), China–Australia collaboration project (2010DFA31730), UK Biotechnology and Biological Sciences Research Council (BB/E017363/1), the Australian Research Council (LP0882095, LP0883462, DP0985953 and LP110100200), the Next-Generation BioGreen 21 Program (PJ008944 and PJ008202), and the US National Science Foundation (IOS 0638418, DBI 0849896, MCB 1021718).
Author information
Authors and Affiliations
Contributions
I.B., B.C., D.E., Q.H., W.H., G.J.K., S.L., Y.L., J. Ma, A.H.P., J.C.P., I.A.P.P., JunW., XiaowuW., XiyinW. and T.-J.Y. are principal investigators (alphabetic order). B.C., W.H., A.H.P., JunW. and XiaowuW. are equally contributing senior authors. S.L., J.W., W.H., X.X. and Z.Y. planned and managed the project. S.L., C.T., A.H.P. and D.E., X.Y. and M.Z. wrote this manuscript and I.B., J. Ma., G.J.K., J.C.P., B.C., T.-J.Y., I.A.P.P., XiyinW., XiaowuW., K.L., Y.L., J.B. and A.G.S. made revision or edits or comments. J.W. (leader), W.H. (co-leader), JunW., L.Y., and Z.Y. performed DNA sequencing. L.Y. (leader), W.H. (co-leader), S.H., J.W., S.L. and J.Y. conducted genomic sequence assembly. S.H. (leader), XiyinW. (co-leader), J.Min, I.B., W.H., J.B., D.E., P.R., S.L., J.S., Y.L. and W.W. conducted scaffold anchoring to linkage maps and assembly validation. X.Y. (leader), J.Y. (co-leader), S.L., Q.Z., S.H. and J. Min performed annotation. C.T. (leader), Wanshun L., W.H., Y.L., C.L., W.W., J. Wu, S.L., C.D. and M.Z. performed transcriptome sequencing. S.L. conceived analysis of comparison and evolution. S.L. (leader), C.T., X.Y., ZhangyanW., C.L., S.H., J. Ma, J.Y., M.Z., Zhuo W., Q.Z., S.P., I.A.P.P., A.G.S., L.Y., I.B., G.J.K., J.C.P., XiaowuW., B.C., F.C., YinH., WenbinL. and X.Liang performed analysis of comparative genomics and evolution. J. Ma (leader), M.Z., Q.Z., C.T., S.L., B.C., S.H., H.B., C.L. and JianaL. conducted TE analysis. XiyinW. (leader), J.Y., T.-J.Y., ZhangyanW., L.W., J. Li, T.-H.L., JinpengW., H.J., X.T., X.L., M.G. and L.J. conducted gene family analysis. K.L. (leader), J.Y., S.L., C.T., H.L., H.G., S.P., D.Z., Z.F., Q.H., Xnfa W., C.Q., D.D., Z.H., Y.H., J.H., D.M., J.L., Z. Li, J.Z., L.X., Y.Zhou., Z.L. and Y.Zhang conducted trait-related gene analysis. A.H.P. (leader), XiyinW., D.J., Y.W. and T.-H.L. conducted gene conversion analysis. T.-J. Y. (leader), M.Z., P.S., B.-S.P., J.Ma, N.E.W., R.Q., X.L., J.Lee and H.H.K. conducted centromere analysis. C.T. (leader), S.L., X.Y., S.H., C.L., Zhangyan W., Q.Z., J.Y., J.T. and J.B. conducted tandemly duplicated gene analysis. ZhangyanW. and J.Y. performed data submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures, Tables, Methods and References
Supplementary Figures 1-68, Supplementary Tables 1-49, Supplementary Methods and Supplementary References (PDF 12359 kb)
Supplementary Data 1
The 23,823 Brassica oleracea-B. rapa orthologous gene pairs and those with different exon numbers (XLS 3106 kb)
Supplementary Data 2
The genes for biosynthesis and breakdown of glucosinolates (GSL) in B. rapa and B. oleracea. (XLS 36 kb)
Supplementary Data 3
The multiple sequence alignment of gene families corresponding to Figure 5 and Supplementary Figures 46-61. (XLS 9526 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Liu, S., Liu, Y., Yang, X. et al. The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat Commun 5, 3930 (2014). https://doi.org/10.1038/ncomms4930
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms4930
This article is cited by
-
Large-scale gene expression alterations introduced by structural variation drive morphotype diversification in Brassica oleracea
Nature Genetics (2024)
-
Macroevolutionary dynamics of gene family gain and loss along multicellular eukaryotic lineages
Nature Communications (2024)
-
Genomic insights into biased allele loss and increased gene numbers after genome duplication in autotetraploid Cyclocarya paliurus
BMC Biology (2023)
-
Functional and evolutionary study of MLO gene family in the regulation of Sclerotinia stem rot resistance in Brassica napus L.
Biotechnology for Biofuels and Bioproducts (2023)
-
Different evolutionary patterns of TIR1/AFBs and AUX/IAAs and their implications for the morphogenesis of land plants
BMC Plant Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.