Abstract
The final size of an organism, or of single organs within an organism, depends on an intricate coordination of cell proliferation and cell expansion. Although organism size is of fundamental importance, the molecular and genetic mechanisms that control it remain far from understood. Here we identify a transcription factor, KUODA1 (KUA1), which specifically controls cell expansion during leaf development in Arabidopsis thaliana. We show that KUA1 expression is circadian regulated and depends on an intact clock. Furthermore, KUA1 directly represses the expression of a set of genes encoding for peroxidases that control reactive oxygen species (ROS) homeostasis in the apoplast. Disruption of KUA1 results in increased peroxidase activity and smaller leaf cells. Chemical or genetic interference with the ROS balance or peroxidase activity affects cell size in a manner consistent with the identified KUA1 function. Thus, KUA1 modulates leaf cell expansion and final organ size by controlling ROS homeostasis.
Similar content being viewed by others
Introduction
Multicellular organisms grow through a coordinated balance between cell proliferation and expansion. Alterations to this balance typically lead to abnormal development, as in the case of cancer1. In plants, organ size is astonishingly constant within a given species and environment. However, large differences are observed when comparing one species with another, indicating that organ size is under genetic control2. Plant leaves are initially established by meristematic cell proliferation and in a second phase by cell expansion without further divisions3. Both phases are regulated by a multitude of genetic pathways, in which a fine-tuned balance between positive and negative regulators, for example, transcription factors (TFs), plays a central role4. In contrast to mammalian cells, plant cells are encased by a cell wall that gives structural support. Not surprisingly, cell expansion is affected by alterations in cell wall content and architecture5,6,7. Such alterations may be mediated by biosynthetic and/or remodelling proteins, including expansins, a class of proteins located in plant cell walls8,9, xyloglucan endotransglucosylase/hydrolases (XETs/XTHs)10, and also by peroxidases (Prxs), which modulate the level of reactive oxygen species (ROS)11. Still, the mechanistic details of their transcriptional regulation, and therefore of plant cell expansion, remain largely unknown.
ROS orchestrate downstream signalling cascades in many different organisms, from bacteria to animals, to direct developmental processes12. ROS are typified as chemically reactive compounds that contain oxygen, including oxygen ions and peroxides. These compounds play important roles in the regulation of plant growth13. Plasma membrane-located NADPH oxidases are the most abundant ROS-producing enzymes in the expanding cell wall14. The activity of three members of this family has, among others, been linked to root growth15. For example, the root hair defective2 (RHD2) protein is required for root hair initiation and expansion15. Prxs represent another class of ROS-related proteins that are widely distributed between plant tissues and have functions that involve a range of different substrates16. In the plant apoplast, Prxs may act as hydrogen peroxide (H2O2)-consuming and/or phenol-oxidizing enzymes, which typically affect lignin formation in the secondary cell wall17. Although ROS can both stimulate and inhibit cell expansion13, the genetic regulation and/or mechanistic details are often not clear. Recently, the bHLH TF UPBEAT1 (UPB1) was found to modulate the balance between cell proliferation and differentiation by repressing Prx genes in Arabidopsis thaliana roots18. Inhibition of peroxidase activity by applying the chemical inhibitor salicylhydroxamic acid (SHAM) to roots or treatment with hydrogen peroxide (H2O2) resulted in reduced meristem cell number and length of the first cortical cell of the root. In contrast, upb1-1 mutants displayed a significantly increased meristem cell number and length of the first cortical cell. This indicates that H2O2 scavenging by root peroxidases controls indeterminate root growth18.
In contrast to roots, leaf growth is determinate and the final size depends on a tight interplay between cell division and expansion. It would therefore be anticipated that overall leaf growth and size is controlled differently from root growth. Here we show that a MYB-like TF, KUODA1 (KUA1), modulates leaf organ size by controlling the expression of Prxs. We further prove causality between peroxidase activity and organ size through chemical and genetic means, thereby providing a mechanism for how final leaf organ size depends on apoplastic ROS turnover. Hence, KUA1 directly controls ROS-mediated cell expansion during leaf development to set the final size of the organ.
Results
KUODA1 positively regulates cell expansion
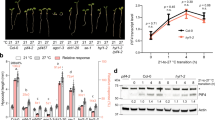
To identify gene networks controlling organ size in Arabidopsis, we performed a reverse genetics screen for genes encoding TFs that are specifically induced during leaf expansion based on transcriptome data3. We found one T-DNA insertion mutant, which affected a MYB-like gene (AT5G47390) (Supplementary Fig. 1a,b), to have reduced leaf size as compared with wild type (WT; Fig. 1a,b). Complementation of the mutant with the gene’s complementary DNA (cDNA) under control of the CaMV 35S promoter verified that the observed phenotype was caused by the loss of the TF. As overexpression of the MYB-like gene (OX) resulted in enlarged leaves, we termed the gene KUODA1 (KUA1; Chinese for ‘enlarge’ or ‘expand’). Leaf size is the sum of cell number and cell size. Most mutants known to affect leaf size act on cell proliferation, or on both cell proliferation and expansion rather than cell expansion alone19,20. To determine changes at the cellular level, mature (that is, not anymore expanding) first true leaves from soil-grown plants were harvested at 22 days after sowing (DAS). By microscopic analysis, we determined the number and area of mesophyll cells in the subepidermal layer and found that the kua1-1 leaves had reduced cell size (65% of WT), while the number of cells was unaffected (Fig. 1c,d). Moreover, the reduction in cell size was equivalent to the reduction in leaf size of the kua1-1 mutant. In contrast, overexpression of KUA1 resulted in a significantly increased leaf area (OX-1, 123% of WT; OX-2, 119% of WT) and cell size (OX-1, 118% of WT; OX-2, 114% of WT), while the number of cells were unaffected (Fig. 1b–d, Supplementary Fig. 1c–e). Consistently, the smaller cell size of the kua1-1 mutant was restored after complementation.
(a) Images of 28-day-old wild type (WT), kua1-1, 35S:KUA1 (OX-1) and 35S:KUA1/kua1-1 plants, respectively. (b–d) Measurements of (b) leaf size, (c) mesophyll cell size and (d) cell number. Values represents means±s.d. (n=20). *P<0.05, Student’s t-test. Leaf and cell size data for a second independent overexpression line (OX-2) and the complementation line (35S:KUA1/kua1-1) is shown in Supplementary Fig. 1. (e) pKUA1::GUS expression pattern during leaf development. Activity is observed starting at the tip on day 12 and is present throughout the leaf blade at day 14 but becomes restricted towards the base at day 17, overlapping with the phase of leaf expansion.
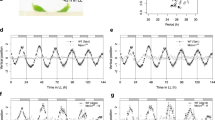
Dicot leaves exhibit diel growth patterns with maximal expansion rates in the morning that are dictated by the circadian clock21. As KUA1 regulates leaf growth, we anticipated that its expression might follow such rhythm. With the DIURNAL tool22, we found a diurnal expression pattern with maximal transcript abundance of KUA1 in the morning (Supplementary Fig. 2a). This pattern was confirmed both at the transcript and translational level in leaves of 15-day-old plants (Supplementary Fig. 2b). Morning-phased gene expression is controlled by two MYB TFs, circadian clock associated 1 (CCA1) and late-elongated hypocotyl (LHY), which affect leaf growth rate23,24. To investigate whether KUA1 expression depends on the clock, we performed a 48-h expression profiling on 16-day-old plants grown under 12-h light/12-h dark (LD) photocycles released to continuous light (LL) in cca1-1/lhy-11, CCA1-ox and WT plants25. Phase-specific expression of KUA1 is lost in both CCA1-ox and cca1-1/lhy-11 plants (one-way analysis of variance (ANOVA), P<0.01; Fig. 2), indicating that its diurnal transcript accumulation depends on an intact clock. The KUA1 promoter contains a CCA1 binding site 658 bp before the transcriptional start site (Supplementary Fig. 3a), similar to the one found in the promoter of the CCA1 direct target gene pseudo-response regulator 9 (ref. 26). Moreover, the CCA1 binding site motif was also found in upstream regulatory sequences of KUA1 orthologues from several other species, including Glyma17g15330 from Glycine max and Os01g09280 from Oryza sativa (Supplementary Fig. 3a). Together with the observation that CCA1 and LHY expression peaks several hours before that of KUA1, this suggests that KUA1 might represent a direct clock output.
Expression profile of KUA1 from 16-day-old plants entrained under 12-h light/12-h dark photocycles and transferred to constant light (LL). Sampling was performed every 4 h for three independent biological replicates. CCA1-ox is in the Col-0 background, and cca1-1/lhy-11 is in the Ler-0 background. Lines represent mean expression values in four genotypes: Col-0, CCA1-ox, Ler-0 and cca1-1/lhy-11. One-way ANOVA (effect of time in Col-0 and Ler-0) for KUA1 was significant (**P<0.01). White bars represent day, and striped bars the subjective night.
Next, we determined the spatio-temporal regulation of KUA1 expression during leaf development. Activity of a pKUA1::GUS reporter construct was not observed before 12 DAS, when staining began to appear at the tip of the leaf (Fig. 1e). Of note, cell proliferation and cell enlargement proceed from the leaf apex to the base27. Furthermore, the transition from proliferation to expansion occurs relatively abrupt28, resulting in a phase of leaf development during which almost all cells undergo expansion3. Consistently, GUS staining intensity at 14 DAS was observed throughout the leaf blade and decreased thereafter in a basipetal manner, indicating that KUA1 is specifically induced during leaf expansion. At maturity, weak GUS staining was observed in vascular tissue, which is in line with previously reported cell-specific expression data for KUA1 in companion cells (Supplementary Fig. 3b).
The KUA1 protein contains a central MYB-like DNA-binding domain and at the N-terminal side a CCHC zinc-finger domain (Supplementary Fig. 1f). Furthermore, two regulatory motifs were detected, a nuclear localization sequence and a transcriptional repression domain, R/KLFGV29. To test whether KUA1 is targeted to the nucleus in planta, the kua1-1 mutant was transformed with a KUA1-GFP fusion protein expressed from the 35S promoter. The translational fusion construct complemented the growth phenotype of the kua1-1 mutant and resulted in a nuclear localized signal of the fusion protein (Supplementary Fig. 1g).
KUODA1 is a transcriptional regulator of ROS-related genes
As KUA1 is specifically induced during leaf expansion, we envisaged that its downstream target genes have a role in setting organ size. To identify KUA1-regulated genes, we made use of an estradiol-inducible overexpression (IOX) system30 and subjected KUA1-IOX seedlings after induction to global transcriptome analysis. After 4 h of KUA1 induction, 198 genes were differentially expressed (twofold change, Student’s t-test P<0.05) of which 107 were upregulated and 91 were downregulated (Supplementary Data 1). As KUA1 contains a transcriptional repression domain, we mainly focused our analysis on genes downregulated by the TF. To investigate possible biological functions of these genes, we identified significantly enriched gene ontology (GO) categories among the changed genes31. The most significantly enriched GO category of genes negatively regulated by KUA1 was ‘peroxidase activity’ (P<10−12) (Fig. 3a, Supplementary Table 1). Other enriched categories included ‘antioxidant activity’ (P<10−6) and ‘response to oxidative stress’ (P<10−6).
(a) Enriched gene ontology (GO) categories among genes that are negatively regulated by KUA1. (b) Expression profile of Prx genes on alteration of KUA1 expression levels as determined by qRT-PCR. Data represent mean ΔΔCT from three independent biological replicates (see Supplementary Table 2). Yellow indicates an increase in expression, blue indicates a decrease in expression; scale bar shows Log2 fold changes (FC). Naming of peroxidase genes after Valério et al.36
For the genes positively regulated by KUA1, the most enriched GO category was ‘glutathione transferase activity’ (P<10−6) (Supplementary Fig. 4a, Supplementary Table 1), indicating that KUA1 affects ROS homeostasis. In addition, ‘cell wall’ (P<10−4) and ‘endomembrane system’ (P<10−6) GO categories were over-represented among the upregulated genes, suggesting a role of KUA1 in cell wall remodelling5.
We verified the expression of a select number of differentially expressed genes by qRT–PCR (Supplementary Table 2). Glutathione S-transferase (GSTU) genes were upregulated on inducible overexpression of KUA1, but a mild effect was observed on the expression of these genes in OX-1 plants (Supplementary Fig. 4b). Still, GSTU genes were downregulated in kua1-1 together with several TF genes, including ZAT12, REVEILLE8 (RVE8) and a bHLH (AT1G10585). In contrast, inducible overexpression of KUA1 resulted in the downregulation of more than 10 Prx genes (Fig. 3b). Consistently, constitutive overexpression of KUA1 represses these genes, while in the kua1-1 mutant several showed enhanced expression, most prominently Prx7, Prx10, Prx23 and Prx57. As KUA1 appears to act as a transcriptional repressor, we used a transient luciferase reporter gene assay to validate this. To this end, the promoter of Prx7 was cloned directly upstream of the firefly luciferase open reading frame and transformed into Arabidopsis protoplasts (Supplementary Fig. 5a). Co-transformation of the reporter construct with a 35S:KUA1 construct resulted in a significant downregulation of the luciferase signal, indicating that KUA1 in vivo acts as a transcriptional repressor of Prx7. Fusion of a GAL4 activation domain with the KUA1 protein overcomes the repressive effect of KUA1 and causes a significant induction of the Prx7 reporter. Taken together, KUA1 acts as a transcriptional repressor and modulates the expression of ROS-related genes.
KUODA1 is a direct regulator of peroxidase genes
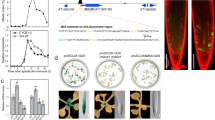
The upstream promoter sequences of the differentially expressed Prxs were screened for over-represented 6-mer motifs to identify a potential DNA-binding site for KUA1 using the MEME Suite32 and TAIR motif finder ( www.arabidopsis.org). This analysis resulted in the identification of several abundant motifs (Fig. 4a). These include ATCACA (Motif1), which represents an unknown motif, and AAACGT (Motif2), which is identical to the SORLREP2 motif that is present in the promoters of light-regulated genes33. Furthermore, an element similar to the RY/G-box was found, C(A/G)TGCA (Motif3), which renders seed-specific expression34, and the (A/C)CTTGC (Motif4), which is similar to the iron-deficiency response element35. We used an electrophoretic mobility shift assay (EMSA) to assess whether KUA1 binds to any of these motifs. We found that KUA1 can bind a DNA probe spanning the unknown ATCACA motif (Motif1) in the promoters of both Prx44 and Prx60, while no binding was found with probes containing the AAACGT motif (Motif2) from Prx60 (Fig. 4b). In addition, a probe with a point mutation in Motif1 from Prx44 causing the exchange of the first base into a G (GTCACA; Motif1*) was less strongly retained in the gel as compared with probes with the original motif. Subsequently, we tested whether Prxs are direct KUA1 targets by performing a chromatin immunoprecipitation (ChIP) assay with the functional GFP-tagged KUA1 protein (35S:KUA1-GFP/kua1-1). To this end, primers spanning genomic regions containing the identified cis-element were used (Fig. 4c). We found that KUA1 associated with the promoters of Prx7, Prx8, Prx10, Prx30, Prx35, Prx44 and Prx57 (Fig. 4d). In addition, no enrichment was detected for three negative controls (promoter segments of three ROS-related genes whose expression levels were unaffected by KUA1) or in the further upstream regions in the promoter of Prx30 (Supplementary Fig. 5b). Thus, KUA1 directly interacts with seven Prx loci causing repression of their expression.
(a) Identified 6-mer motifs (Motif1-4) enriched in upstream promoter sequences of the differentially expressed Prx genes in KUA1-IOX microarray experiment, as detected by the MEME Suite and TAIR motif finder. Indicated are the E-value, representing the statistical significance of the motif, and the number of sites contributing to the construction of the motif32. (b) EMSA performed with probes for Motif1 and Motif2 (A). Incubation of KUA1 protein with oligonucleotides spanning Motif1 from the promoters of Prx44 or Prx60 causes a band shift (arrow), which disappears on competition (‘comp’) with unlabelled probe (100-fold molar excess). A mutation in Motif1 from Prx44 (Prx44 Motif1*) from ATCACA to GTCACA reduced binding affinity. No retention was observed with probes based on Motif2 from Prx60. (c) KUA1-binding sites (M; as based on Motif1) in upstream promoter regions of selected Prx genes. Blue arrows indicate amplified regions in ChIP-qPCR experiments. Positions of the transcriptional start sites are indicated by arrows with turning lines. (d) ChIP-qPCR results from 35S:KUA1-GFP/kua1-1 plants for seven Prx promoters and three negative controls. M1 and M2 represent KUA1 binding sites as shown in c; Prx genes are indicated by numbers. N1, N2 and N3 represent genomic control regions lacking KUA1 binding sites. Values represent average enrichment (FC) of three independent biological replicates. The amounts of immunoprecipitated genomic DNA were normalized to the input fraction. The fold enrichments for immunoprecipitation of the KUA1-GFP DNA complex by an anti-GFP antibody were calculated over control (IgG precipitated) samples for each analysed region. *P<0.05, Student’s t-test. Error bars indicate s.e. (n=3).
Peroxidase activity is linked to cell size
Prxs can stimulate as well as restrict plant growth11. The enhanced expression of Prxs in kua1-1 and their repression by KUA1 suggest that increased peroxidase activity might be symptomatic for the observed phenotype. To test this, we treated 14-day-old plants with SHAM, a peroxidase inhibitor, and measured the size of the first leaf pair at 22 DAS. SHAM treatment of the kua1-1 mutant resulted in an increased plant size in a concentration-dependent manner (Fig. 5a). Similar, but less prominent, effects were observed for WT. A single application of 20 μM SHAM fully restored expansion growth of the first leaf pair in kua1-1, while 5 μM showed a partial recovery as compared with WT (Fig. 5b,c). Since Class III peroxidases have been implicated in the production of H2O2 in the apoplast11,36, we stained plants with the 3,3′-diaminobenzidine and found an increased intensity in the kua1-1 mutant as compared with WT (Supplementary Fig. 6a). Quantitative determination of the H2O2 concentration revealed a significant increase in kua1-1 leaves, while significant lower levels were detected in OX-1 plants, as compared with wild type (Supplementary Fig. 6b). In contrast, no difference in the levels of superoxide was found between the different lines (Supplementary Fig. 6c). Moreover, treatment of kua1-1 with the H2O2 scavenger KI reverted the mutant phenotype (Supplementary Fig. 6d–g). These results indicate that increased peroxidase activity and apoplastic H2O2 production are causative for the reduced growth.
(a) Images of 22-day-old wild-type (WT) and kua1-1 plants treated at day 14 with SHAM. (b–c) Measurements of (b) leaf size and (c) cell size. Values represents means±s.d. (n=20). *P<0.05, Student’s t-test. (d) Peroxidase activities of ionically bound and soluble protein fractions were determined for wild type (WT), kua1-1 and 35S:KUA1 at ZT4 and ZT9. Values represent means±s.d. (n=4). *P<0.05, Student’s t-test. FW, fresh weight. (e) Images of 22-day-old wild-type (WT) and 35S:Prx57 plants. (f–h) Measurements of (f) leaf size, (g) mesophyll cell size and (h) cell number. Values represent means±s.d. (n=20). *P<0.05, Student’s t-test.
As leaf growth is under control of the diel cycle, we measured cell wall peroxidase activity at Zeitgeber times ZT4 and ZT9. KUA1 had the highest expression level at ZT4 (Supplementary Fig. 2). Consistent with a negative regulation of Prx expression, we found lower peroxidase activity in the WT at ZT4 as compared with ZT9 for both the ionically bound and soluble fraction (Fig. 5d). Furthermore, kua1-1 showed significantly higher peroxidase activity at both time points, while overexpressing plants showed significantly lower activity at ZT9. To further confirm that Prxs regulate leaf growth, we obtained an overexpression line for Prx57 (ref. 18), which showed increased peroxidase activity (Supplementary Fig. 6h). Interestingly, leaf and cell size in plants with increased Prx57 activity was significantly reduced, while cell number was not affected (Fig. 5e–h). These results imply that KUA1 modulates cell size through the action of cell wall-located Prxs.
Prxs are mainly known to control extensibility of the cell wall by causing crosslinking or cleaving of cell wall polysaccharides11,36. Still, it is possible that the changes in Prx expression cause other cell wall alterations. We therefore determined the cell wall compositions in the different genotypes, including the 35S:Prx57 line. Neither cellulose, the major component of the cell wall, was affected by altering the expression level of KUA1 (Table 1), nor was that of uronic acids. In dicots, the hemicellulose network mainly consists of xyloglucan37, which is known to be enzymatically loosened by the action of expansins, endoglucanases and endotransglycosylases to allow for turgor-driven growth38,39. Among the neutral sugars measured, we only found an increase in the level of glucose for the KUA1 overexpressing line (Table 1). These data indicate that changes in KUA1 expression have only limited effect on the composition of the cell wall.
Discussion
Plant tissue growth depends on cell proliferation and directed cell expansion. Although several gene regulatory networks that control leaf primordia initiation and proliferation have been uncovered4, those controlling cell expansion remain largely elusive. Here, we identified kua1-1, which has smaller leaves due to a decrease in cell size but not cell number. In contrast, KUA1 overexpression resulted in increased cell size and larger leaves, while the number of cells was unaffected. Thus, KUA1 is a specific regulator of leaf cell size in plants (Fig. 6). Through expression profiling on inducible overexpression lines, we found that KUA1 negatively regulates the expression of Prx genes. Furthermore, by ChIP and EMSA assays, we demonstrated that KUA1 directly regulates the expression of seven Prx genes. This repression alters the apoplastic ROS balance, establishing the means to physically control cell expansion in leaves (Fig. 6).
In this model, the expression level of KUA1 is positively correlated with whole-leaf and leaf cell size, but negatively with peroxidase activity. A high expression of KUA1 (red) results in the repression of seven peroxidase Prx genes, which allows for cell expansion due to lower H2O2 levels and subsequently reduced crosslinking of the cell wall. In contrast, restriction of leaf size can be achieved by downregulation of KUA1 (blue), which results in an increased peroxidase activity and a stiffening of the cell wall, inhibiting cell expansion.
Apoplastic Prxs are known to either restrict or promote cell expansion11,17,18. These apparent contradictory effects are linked to the regulatory modes under which the Prxs are working. On one hand, the Prxs may favour elongation by generation of oxygen radicals or restrict growth by the generation of H2O2. In the first scenario, cell wall polymers may be cleaved by the radicals and thus promote cell wall loosening and therefore growth. In contrast, an increase in H2O2 typically leads to crosslinking phenolics and promotes rigid crosslinks between extensins making the cell wall stiffer11. KUA1 effectively represses Prx expression and activity in leaves, which promotes cell expansion. While this promotion is clearly linked to changed levels of apoplastic H2O2, it is important to note that changes in H2O2 also can affect the pool of oxygen radicals40. Notably, at least in the case of KUA1, the inhibitory effect of Prx, mediated via H2O2, appears to be the causal link for regulation of leaf cell expansion.
Organ size is typically underpinned by a balance between cell proliferation and growth3. However, this balance is largely depending on tissue and cell type. This may readily be seen in plant organ growth, where leaf growth is typically restricted to a set number of expanding cells, whereas root growth is a result of continuous cell proliferation and cell expansion18,28. Hence, the determinate and indeterminate growth of leaves and roots, respectively, should be regulated by different molecular and genetic means. Interestingly, Prxs were recently shown to affect Arabidopsis root growth18. Here, a superoxide/H2O2 equilibrium regulates the size of proliferation and differentiation zones. An enhanced expression of Prx genes in roots lacking the TF UPB1 was associated with an increased number of meristematic cells and an increase in length of the first mature cortical cells18. Thus, enhanced Prx activity promotes cellular proliferation and differentiation in roots. In our study, we show that repression of Prx expression by KUA1 enhanced leaf cell expansion, but did not cause an increase in leaf cell proliferation. Furthermore, as both SHAM and H2O2 treatments reduce the size of the first cortical root cell18, the root-localized Prxs are presumably needed to maintain low H2O2 levels to promote root cell expansion. In contrast, here we show that inhibition of leaf peroxidase activity increases leaf size, suggesting that in leaves the apoplastic Prxs mainly produce H2O2, which potentially results in cell wall crosslinking11. Hence, the impact of Prxs, or more precisely their effect on the H2O2 level, appears to have largely opposing effects on organ growth in plants.
Leaf expansion rates in dicots follow a diel rhythm21, and this correlates with the accumulation of KUA1 transcript during the morning. Moreover, KUA1 transcript cycling depends on a functional clock, as in cca1-1/lhy-11 and CCA1-ox plants the time-of-day-specific expression of KUA1 is lost. Interestingly, the promoter of KUA1 and its orthologues contain a CCA1 binding site (Supplementary Fig. 3a), indicating that the expansion growth controlled by KUA1 could be directly regulated by the clock. Previously, we demonstrated that ROS homeostasis and ROS-responsive gene expression are under the control of CCA1 (ref. 25). Here we demonstrate that KUA1 controls apoplastic ROS homeostasis to mediate cell expansion. Of note, leaf expansion rates in dicots are the highest in the morning24, which correlates with the increased expression level of KUA1 during this time of the day. Thus, to obtain high expansion rates during the morning, KUA1 is induced to repress Prx gene expression. In addition, the promoter of KUA1 contains several abiotic stress-related cis-elements (Supplementary Fig. 3a). As expression of KUA1 is affected by various abiotic stresses (Supplementary Fig. 3c) it is likely that it functions to modulate growth in a changing environment.
In conclusion, we have identified a mechanism in which KUA1 regulates Prx activity to control final leaf cell size. Despite that Prxs are commonly required for the modulation of H2O2 levels in the apoplastic space, there are remarkable differences in the molecular implementation of their action in the regulation of leaf and root growth.
Methods
Plant material and growth conditions
Arabidopsis thaliana Columbia-0 (Col-0) was used as a wild type. The T-DNA insertion line kua1-1 (GK-783B02) was obtained from the Nottingham Arabidopsis Stock Centre. Plant growth and β-estradiol treatments were done as previously described41. In short, Arabidopsis plants were grown in soil (Einheitserde GS90; Gebrüder Patzer, Sinntal-Jossa, Germany) in a growth chamber with a 16-h day (120 μmol m−2 s−1) and a day/night temperature of 22/16 °C and 60/75% relative humidity. In tissue culture, seedlings were grown in half-strength Murashige and Skoog medium (0.5 MS) supplemented with 1% sucrose and solidified with 0.7% agar under a 16-h day (140 μmol m−2 s−1)/8-h night regime (22 °C). SHAM and KI treatments were done by applying a foliar spray to 14-day-old soil-grown plants. GUS staining was performed according to Jefferson42.
Construction of plasmids
To generate overexpression constructs and C-terminal GFP fusions of KUA1, the coding sequence (CDS) was amplified from leaf cDNA by PCR and cloned into pENTR/D (Invitrogen) with the primers listed in Supplementary Table 3. Sequence-confirmed ENTRY clones were used for recombination with pK7FWG2 vector to construct a fusion with the GFP coding region at the C-terminal part of the TF ( http://gateway.psb.ugent.be)43. The clones containing the stop codon were used for generating a CaMV 35S construct by recombination with vector pK7WG2 (ref. 43).
The promoter region directly upstream of the start codon of KUA1 (1,050 bp) was amplified from genomic DNA and cloned into pENTR/D and verified by sequencing. Thereafter, the clone was recombined with the GATEWAY compatible vector pKGWFS7 that contains a fused reporter gene consisting of eGFP and the E. coli β-glucuronidase (GUS)43. End clones were selected by restriction analysis and confirmed by sequencing before plant transformation.
For the establishment of an inducible overexpression vector the KUA1 CDS was amplified by PCR with primers that contain an XhoI or SpeI site (Supplementary Table 3). PCR products were ligated overnight by TA cloning into pGEM-T Easy vector (Promega). After sequence confirmation, the vector was digested and the XhoI-SpeI fragment containing the CDS was ligated into the pER8 vector30.
Plant transformation and selection of transgenic plants
Constructs were transformed into Arabidopsis Col-0 wild type or the kua1-1 mutant using Agrobacterium tumefaciens (strain GV3101)-mediated transformation employing a modified floral dip method44. T0 seeds were selected on MS medium containing the appropriate antibiotic. Subsequently, stable transgenic T3 lines were established based on a segregation analysis. Transgenic lines showing either an increased expression level by qPCR or reporter gene signal were used for detailed analysis in this study.
Microscopic analysis
Cleared first true leaves of 22-day-old plants were used to measure leaf area, cell size and cell number according to Nguyen et al.41
Total and polysomal RNA isolation
Total RNA was extracted from leaves or seedlings using the RNeasy kit (Qiagen). Before cDNA synthesis with RevertAid First-strand cDNA Synthesis kit (Fermentas), the isolated RNA was treated with DNase I (Ambion). Quantitative real-time PCR was done using Power SYBR Green PCR Master Mix (Applied Biosystems). ACTIN2 (AT3G18780) was used as reference gene (Supplementary Table 3). Oligonucleotides for qPCR were designed with the webtool QUANTPRIME45. Relative transcript abundance was determined by the comparative CT method46. Polysomes from 35S::HF-RPL18 plants (15 DAS) were immunoprecipitated using powdered tissue and extracted in polysome extraction buffer as previously described47,48.
Gene expression analysis
For microarray experiments total RNA was isolated from 4-h mock or 15 μM estradiol-treated KUA1-IOX plants. Two independent biological replicates for each treatment were used for hybridization to Arabidopsis whole-genome Affymetrix ATH1 GeneChip (ATLAS Biolabs). Expression data were submitted to the NCBI Gene Expression Omnibus (GEO) repository ( www.ncbi.nlm.nih.gov/geo/) under accession number GSE50520.
ChIP assay
35S:KUA1-GFP/kua1-1 plants were grown for 12 days on MS media. Whole-shoot tissue was fixed with formaldehyde and for the ChIP assay the EpiQuick kit (Epigentek Group Inc, NY, USA) was used as described in Lai et al.25
Motif analysis and EMSA
To identify conserved promoter motifs among the differentially expressed Prx genes, 1,000-bp upstream sequences were extracted using Phytozome49. Promoter analysis was performed using the MEME program with the following settings: one or more occurrences per sequence, a 6-base motif length, and 15 as the minimum number of sites to be detected in all promoters tested32. Next to that, the AGI codes of the Prx genes were used as a query set for TAIR motif analysis ( www.arabidopsis.org). Motifs reported by both tools were compared with known cis-regulatory elements using the webtool ATCOECIS50. Subsequently, several motifs were tested empirically by using an EMSA as previously described51.
In short, the CDS of KUA1 was cloned into the pF3A WG Flexi vector (Promega), which contains translation enhancer sequences from the barley yellow dwarf virus (BYDV), an RNA plant virus, that facilitates in vitro expression of proteins in wheat germ extracts. For the in vitro expression, 5.0 μg of plasmid DNA was added to TNT SP6 High Yield Wheat Germ Mastermix (Promega) containing 1 μl FluoroTec Green Lys (Promega). After 2 h of incubation at 30 °C, synthesized protein was analysed by SDS-PAGE and detection was performed with a Typhoon Scanner (GE Healthcare).
For gel-shift assays, Cy5-labelled probes were generated based on 35–40 bp long regions derived from the Prx promoters (Supplementary Table 3). DNA probes were established by annealing forward and reverse oligonucleotides in equal-molar ratios and slowly cooling them down from 95 to 30 °C. Probes were diluted to a final concentration of 500 fmol and kept in amber tubes until use. Binding reactions were performed using the LightShift Chemiluminescent Assay Kit (Pierce, Rockford, USA). The incubated protein-DNA mixture was separated on a 5% native polyacrylamide gel after which the Cy5 signal was captured using a Typhoon Scanner.
Peroxidase activity assay and ROS quantification
Proteins were extracted from leaves according to Bindschedler et al.52 Peroxidase activity was determined spectrophotometrically at 405 nm using 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) diammonium salt as a substrate. Determination of H2O2 was done by using the Amplex Red Hydrogen Peroxide/Peroxidase Assay Kit (Invitrogen) as described previously25. NBT staining and measurement of superoxide levels were done according to Tsukagoshi et al.18
Cell wall biochemical analysis
For cell wall analysis, the first leaf pair of soil-grown plants at 15 DAS was extracted, and crude cell wall material of six independent biological replicates was used for neutral sugar analysis, uronic acid and cellulose measurements as described53.
Transactivation assay
A 1-kb promoter fragment of Prx7 was amplified and cloned into the p2GWL7.0 vector43. Furthermore, a 35S:KUA1 and 35S:GAD:KUA1 fusion constructs (the latter containing the GAL4 activation domain) were created as described before54. Subsequently, protoplast transformation and measurement of luciferase signal were done as reported previously41.
Additional information
Accession codes: The microarray expression data have been deposited in the NCBI Gene Expression Omnibus (GEO) database under accession code GSE50520.
How to cite this article: Lu, D. et al. Transcriptional control of ROS homeostasis by KUODA1 regulates cell expansion during leaf development. Nat. Commun. 5:3767 doi: 10.1038/ncomms4767 (2013).
Accession codes
References
Zhang, L. et al. The TEAD/TEF family of transcription factor Scalloped mediates Hippo signaling in organ size control. Dev. Cell 14, 377–387 (2008).
Powell, A. E. & Lenhard, M. Control of organ size in plants. Curr. Biol. 22, R360–R367 (2012).
Beemster, G. T. et al. Genome-wide analysis of gene expression profiles associated with cell cycle transitions in growing organs of Arabidopsis. Plant Physiol. 138, 734–743 (2005).
Townsley, B. T. & Sinha, N. R. A new development: evolving concepts in leaf ontogeny. Annu. Rev. Plant Biol. 63, 535–562 (2012).
Geisler, D. A., Sampathkumar, A., Mutwil, M. & Persson, S. Laying down the bricks: logistic aspects of cell wall biosynthesis. Curr. Opin. Plant Biol. 11, 647–652 (2008).
Bringmann, M. et al. POM-POM2/cellulose synthase interacting1 is essential for the functional association of cellulose synthase and microtubules in Arabidopsis. Plant Cell 24, 163–177 (2012).
Rubio-Díaz, S. et al. Cell expansion-mediated organ growth is affected by mutations in three EXIGUA genes. PLoS ONE 7, e36500 (2012).
Cosgrove, D. J. Growth of the plant cell wall. Nat. Rev. Mol. Cell Biol. 6, 850–861 (2005).
Schmidt, R. et al. Multipass, a rice R2R3-type MYB transcription factor, regulates adaptive growth by integrating multiple hormonal pathways. Plant J. 76, 258–273 (2013).
Sasidharan, R. et al. Light quality-mediated petiole elongation in Arabidopsis during shade avoidance involves cell wall modification by xyloglucan endotransglucosylase/hydrolases. Plant Physiol. 154, 978–990 (2010).
Passardi, F., Penel, C. & Dunand, C. Performing the paradoxical: how plant peroxidases modify the cell wall. Trends Plant Sci. 9, 534–540 (2005).
Schippers, J. H. M., Nguyen, H. M., Lu, D., Schmidt, R. & Mueller-Roeber, B. ROS homeostasis during development: an evolutionary conserved strategy. Cell Mol. Life Sci. 69, 3245–3257 (2012).
Schopfer, P. & Liszkay, A. Plasma membrane-generated reactive oxygen intermediates and their role in cell growth of plants. Biofactors 28, 73–81 (2006).
Keller, T. et al. A plant homolog of the neutrophil NADPH oxidase gp91phox subunit gene encodes a plasma membrane protein with Ca2+ binding motifs. Plant Cell 10, 255–266 (1998).
Foreman, J. et al. Reactive oxygen species produced by NADPH oxidase regulate plant cell growth. Nature 422, 442–446 (2003).
Hiraga, S., Sasaki, K., Ito, H., Ohashi, Y. & Matsui, H. A large family of class III plant peroxidases. Plant Cell Physiol. 42, 462–468 (2001).
Lee, Y., Rubio, M. C., Alassimone, J. & Geldner, N. A mechanism for localized lignin deposition in the endodermis. Cell 153, 402–412 (2013).
Tsukagoshi, H., Busch, W. & Benfey, P. N. Transcriptional regulation of ROS controls transition from proliferation to differentiation in the root. Cell 143, 606–616 (2010).
Horiguchi, G., Fujikura, U., Ferjani, A., Ishikawa, N. & Tsukaya, H. Large-scale histological analysis of leaf mutants using two simple leaf observation methods: identification of novel genetic pathways governing the size and shape of leaves. Plant J. 48, 638–644 (2006).
Pérez-Pérez, J. M. et al. Lessons from a search for leaf mutants in Arabidopsis thaliana. Int. J. Dev. Biol. 53, 1623–1634 (2009).
Ruts, T., Matsubara, S., Wiese-Klinkenberg, A. & Walter, A. Diel patterns of leaf and root growth: endogenous rhythmicity or environmental response? J. Exp. Bot. 63, 3339–3351 (2012).
Mockler, T. C. et al. The DIURNAL project: DIURNAL and circadian expression profiling, model-based pattern matching, and promoter analysis. Cold Spring Harb. Symp. Quant. Biol. 72, 353–363 (2007).
Wang, Z. Y. & Tobin, E. M. Constitutive expression of the circadian clock associated 1 (CCA1) gene disrupts circadian rhythms and suppresses its own expression. Cell 26, 1207–1217 (1998).
Ruts, T., Matsubara, S., Wiese-Klinkenberg, A. & Walter, A. Aberrant temporal growth pattern and morphology of root and shoot caused by a defective circadian clock in Arabidopsis thaliana. Plant J. 72, 154–161 (2012).
Lai, A. G. et al. Circadian clock-associated 1 regulates ROS homeostasis and oxidative stress responses. Proc. Natl Acad. Sci. USA 109, 17129–17134 (2012).
Farré, E. M., Harmer, S. L., Harmon, F. G., Yanovsky, M. J. & Kay, S. A. Overlapping and distinct roles of PRR7 and PRR9 in the Arabidopsis circadian clock. Curr. Biol. 15, 47–54 (2005).
Donnelly, P. M., Bonetta, D., Tsukaya, H., Dengler, R. E. & Dengler, N. G. Cell cycling and cell enlargement in developing leaves of Arabidopsis. Dev. Biol. 215, 407–419 (1999).
Andriankaja, M. et al. Exit from proliferation during leaf development in Arabidopsis thaliana: a not-so-gradual process. Dev. Cell 22, 64–78 (2012).
Ikeda, M. & Ohme-Takagi, M. A novel group of transcriptional repressors in Arabidopsis. Plant Cell Physiol. 50, 970–975 (2009).
Zuo, J., Niu, Q. W. & Chua, N. H. (2000) Technical advance: An estrogen receptor-based transactivator XVE mediates highly inducible gene expression in transgenic plants. Plant J. 24, 265–273 (2000).
Proost, S. et al. PLAZA: a comparative genomics resource to study gene and genome evolution in plants. Plant Cell. 21, 3718–3731 (2009).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Hudson, M. E. & Quail, P. H. Identification of promoter motifs involved in the network of phytochrome A-regulated gene expression by combined analysis of genomic sequence and microarray data. Plant Physiol. 133, 1605–1616 (2003).
Ezcurra, I., Wycliffe, P., Nehlin, L., Ellerström, M. & Rask, L. Transactivation of the Brassica napus napin promoter by ABI3 requires interaction of the conserved B2 and B3 domains of ABI3 with different cis-elements: B2 mediates activation through an ABRE, whereas B3 interacts with an RY/G-box. Plant J. 24, 57–66 (2000).
Ogo, Y. et al. A novel NAC transcription factor, IDEF2, that recognizes the iron deficiency-responsive element 2 regulates the genes involved in iron homeostasis in plants. J. Biol. Chem. 283, 13407–13417 (2008).
Valério, L., De Meyer, M., Penel, C. & Dunand, C. Expression analysis of the Arabidopsis peroxidase multigenic family. Phytochemistry 65, 1331–1342 (2004).
Pauly, M., Albersheim, P., Darvill, A. & York, W. S. Molecular domains of the cellulose/xyloglucan network in the cell walls of higher plants. Plant J. 20, 629–639 (1999).
Marga, F., Grandbois, M., Cosgrove, D. J. & Baskin, T. I. Cell wall extension results in the coordinate separation of parallel microfibrils: evidence from scanning electron microscopy and atomic force microscopy. Plant J. 43, 181–190 (2005).
Rose, J. K., Braam, J., Fry, S. C. & Nishitani, K. The XTH family of enzymes involved in xyloglucan endotransglucosylation and endohydrolysis: current perspectives and a new unifying nomenclature. Plant Cell Physiol. 43, 1421–1435 (2002).
Liszkay, A., Kenk, B. & Schopfer, P. Evidence for the involvement of cell wall peroxidase in the generation of hydroxyl radicals mediating extension growth. Planta 217, 658–667 (2003).
Nguyen, H. M. et al. An upstream regulator of the 26S proteasome modulates organ size in Arabidopsis thaliana. Plant J. 74, 25–36 (2013).
Jefferson, R. A. Assaying chimeric genes in plants: The GUS gene fusion system. Plant Mol. Biol. Rep. 5, 387–405 (1987).
Karimi, M., Inzé, D. & Depicker, A. Gateway vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci. 7, 193–195 (2002).
Logemann, E., Birkenbihl, R. P., Ulker, B. & Somssich, I. E. An improved method for preparing Agrobacterium cells that simplifies the Arabidopsis transformation protocol. Plant Methods 2, 16 (2006).
Arvidsson, S., Kwasniewski, M., Riaño-Pachón, D. M. & Mueller-Roeber, B. QuantPrime - a flexible tool for reliable high-throughput primer design for quantitative PCR. BMC Bioinformatics. 9, 465 (2008).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108 (2008).
Mustroph, A., Juntawong, P. & Bailey-Serres, J. Isolation of plant polysomal mRNA by differential centrifugation and ribosome immunopurification methods. Meth. Mol. Biol. 553, 109–126 (2009).
Ribeiro, D. M., Araújo, W. L., Fernie, A. R., Schippers, J. H. M. & Mueller-Roeber, B. Translatome and metabolome effects triggered by gibberellins during rosette growth in Arabidopsis. J. Exp. Bot. 63, 2769–2786 (2012).
Goodstein, D. M. et al. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178–D1186 (2012).
Vandepoele, K., Quimbaya, M., Casneuf, T., De Veylder, L. & Van de Peer, Y. Unraveling transcriptional control in Arabidopsis using cis-regulatory elements and coexpression networks. Plant Physiol. 150, 535–546 (2009).
Schmidt, R. et al. Salt-responsive ERF1 regulates reactive oxygen species-dependent signaling during the initial response to salt stress in rice. Plant Cell 25, 2115–2131 (2013b).
Bindschedler, L. V. et al. Peroxidase-dependent apoplastic oxidative burst in Arabidopsis required for pathogen resistance. Plant J. 47, 851–863 (2006).
Persson, S. et al. The Arabidopsis irregular xylem8 mutant is deficient in glucuronoxylan and homogalacturonan, which are essential for secondary cell wall integrity. Plant Cell 19, 237–255 (2007).
Schmidt, R. et al. SALT-RESPONSIVE ERF1 is a negative regulator of grain filling and gibberellin-mediated seedling establishment in rice. Mol. Plant 7, 404–421 (2014).
Acknowledgements
T.W., S.P. and B.M.-R. thank the Max Planck Society and the International Max Planck Research School ‘Plant Growth and Primary Metabolism’ for funding. D.L. thanks the China Scholarship Council for financial support. We thank Alvina Lai for support with circadian experiments and Professor Dr Graham Noctor for useful advice regarding apoplastic redox pools. We are also very grateful to Norma Funke, Ines Fehrle and Alexander Erban, as well as Dr Joachim Kopka for assistance during cell wall analyses.
Author information
Authors and Affiliations
Contributions
J.H.M.S. and B.M.-R. designed the experiments. D.L. and J.H.M.S. performed most experiments and data analyses, T.W. and S.P. performed cell wall analysis, D.L., B.M.-R. and J.H.M.S. wrote the manuscript; J.H.M.S. and S.P. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures, Tables and Reference
Supplementary Figures 1-6, Supplementary Tables 1-3 and Supplementary Reference. (PDF 1063 kb)
Supplementary Data 1
Indentification of KUA1 regulon. Genes differentially expressed upon induction of KUA1. Results are based on two biological replicates. (XLSX 21 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/3.0/
About this article
Cite this article
Lu, D., Wang, T., Persson, S. et al. Transcriptional control of ROS homeostasis by KUODA1 regulates cell expansion during leaf development. Nat Commun 5, 3767 (2014). https://doi.org/10.1038/ncomms4767
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms4767
This article is cited by
-
OsUGE2 Regulates Plant Growth through Affecting ROS Homeostasis and Iron Level in Rice
Rice (2024)
-
Periderm differentiation: a cellular and molecular approach to cork oak
Trees (2023)
-
The effects of exogenously applied antioxidants on plant growth and resilience
Phytochemistry Reviews (2023)
-
Quaternary ammonium iminofullerenes improve root growth of oxidative-stress maize through ASA-GSH cycle modulating redox homeostasis of roots and ROS-mediated root-hair elongation
Journal of Nanobiotechnology (2022)
-
Mapping of the plant SnRK1 kinase signalling network reveals a key regulatory role for the class II T6P synthase-like proteins
Nature Plants (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.