Abstract
Accumulation of (R)-2-hydroxyglutarate in cells results from mutations to isocitrate dehydrogenase that correlate with cancer. A recent study reports that (R)-, but not (S)-2-hydroxyglutarate, acts as a co-substrate for the hypoxia-inducible factor prolyl hydroxylases via enzyme-catalysed oxidation to 2-oxoglutarate. Here we investigate the mechanism of 2-hydroxyglutarate-enabled activation of 2-oxoglutarate oxygenases, including prolyl hydroxylase domain 2, the most important human prolyl hydroxylase isoform. We observe that 2-hydroxyglutarate-enabled catalysis by prolyl hydroxylase domain 2 is not enantiomer-specific and is stimulated by ferrous/ferric ion and reducing agents including L-ascorbate. The results reveal that 2-hydroxyglutarate is oxidized to 2-oxoglutarate non-enzymatically, likely via iron-mediated Fenton-chemistry, at levels supporting in vitro catalysis by 2-oxoglutarate oxygenases. Succinic semialdehyde and succinate are also identified as products of 2-hydroxyglutarate oxidation. Overall, the results rationalize the reported effects of 2-hydroxyglutarate on catalysis by prolyl hydroxylases in vitro and suggest that non-enzymatic 2-hydroxyglutarate oxidation may be of biological interest.
Similar content being viewed by others
Introduction
Mutations to genes encoding for enzymes of the tricarboxylic acid (TCA) cycle, including isocitrate dehydrogenase (IDH1/2), can lead to substantial increases in cellular metabolites, as observed in both genetic diseases (germline mutations)1 and in cancer (somatic mutations)2. Elevated levels of (R)-2-hydroxyglutarate ((R)-2HG) produced by a ‘gain-of-function’ mutation to IDH1/2 result in massively increased concentrations of (R)-2HG (up to 10 mM) and decreases in 2-oxoglutarate (2OG) levels3. It is proposed that elevated levels of succinate, fumarate and (R)-2HG in humans cause pro-oncogenic effects via inhibition of Fe(II)/2OG dependent oxygenases4,5,6. 2OG oxygenases are involved in chromatin regulation by modification of both nucleic acids, for example 5-methylcytosine oxidation, and histone tails via Nε-methyl lysine residue demethylation7. 2OG oxygenases also play pivotal roles in the cellular and physiological responses to hypoxia by regulation of the levels and activity of the hypoxia-inducible transcription factor (HIF), which is an α,β-heterodimeric protein that activates transcription of an array of genes under conditions of low oxygen availability8.
The prolyl hydroxylase domain enzymes (PHDs, EGLN 1-3) catalyse hydroxylation of prolyl-residues in HIF-α isoforms, a post-translational modification that signals for the proteosomal degradation of HIF-α9. Factor inhibiting HIF (FIH) catalyses hydroxylation of an asparaginyl-residue resulting in reduction of HIF-α mediated transcription by reducing binding of HIF to transcriptional co-activators9. HIF upregulation activates the transcription of a gene array, which is proposed to promote tumour growth via increased angiogenesis. It is therefore proposed that inhibition of HIF hydroxylases by TCA cycle metabolites and (R)-2HG may be linked to tumour development10. It has been reported that elevated (R)-2HG levels caused by IDH mutations may indeed inhibit PHD action in cells resulting in HIF-activated transcription11, though our work and that of others has indicated that (R)- and (S)-2HG are only weak inhibitors for the HIF hydroxylases12,13,14. It is also possible that elevated (R)-2HG concentrations inhibit chromatin-modifying 2OG oxygenases15,16.
Although it has been widely perceived that HIF is an oncoprotein, HIF-α isoforms can act as tumour suppressors in certain tumours17,18,19,20. Thus, PHD inhibition may be of therapeutic interest for such cancers. In contrast to the paradigm that TCA cycle metabolite and (R)-2HG-mediated inhibition of 2OG dependent oxygenases is pathogenic, it has been recently reported that (R)-, but not (S)-2HG, actually stimulates PHD activity, both with isolated components and in cells, so reducing HIF levels, leading to enhanced proliferation of astrocytes21. However, the effect of (R)-2HG on PHD2 activity and its potential relation to cancer remain controversial11,13,15,21.
The consensus catalytic mechanism for 2OG dependent oxygenases proceeds via formation of an enzyme–Fe(II)–2OG–substrate complex that binds dioxygen; subsequent oxidative decarboxylation of 2OG yields succinate and carbon dioxide and a reactive Fe(IV)–oxo intermediate, which performs the two-electron substrate oxidation (Fig. 1, refs 22, 23). Koivunen et al.21 have proposed that 2OG can be formed enzymatically in the PHD2 active site as a product of (R)-2HG, but not (S)-2HG, oxidation, and suggested that this occurs via an Fe(III)–superoxide intermediate. While prime substrate C-H cleavage by such intermediates has precedent in some non-haem iron oxygenases24, C-H cleavage in a co-substrate is unprecedented in 2OG oxygenase catalysis and in earlier work we did not observe that 2HG can substitute for 2OG under our standard assay conditions14.
Here we report investigations on the mechanism of the apparent activation of PHDs by (R)-2HG and the origin of the 2HG conversion to 2OG. We show that conversion of 2HG to 2OG occurs non-enzymatically in a manner enabled by biologically relevant reducing agents including L-ascorbate and the reduced form of L-glutathione. The level of 2OG is sufficient to support catalysis by PHD2, and at least one other human 2OG oxygenase.
Results
Studies on the mechanisms of 2HG-promoted PHD2 catalysis
To investigate the mechanism by which PHD activity is enabled by the presence of 2HG, MALDI-TOF-MS (matrix assisted laser desorption ionization-time of flight-mass spectrometry) based assays were performed using a peptide fragment of the HIF-1α C-terminal oxygen-dependent degradation domain (HIF-1α556–574, hereafter CODD) and recombinant PHD2181–426 (hereafter PHD2). In contrast to Koivunen et al.21, when substituting 300 μM 2OG for 5 mM 2HG in our standard steady-state assay conditions (4 μM PHD2, 200 μM CODD; Supplementary Table 1), we did not observe any CODD hydroxylation after short incubation times (up to 2 h, limit of detection: ~3% CODD hydroxylation). CODD hydroxylation was observed, however, with longer incubation times (20 h) (Fig. 2c,d) in the presence of both (R)- and (S)-2HG.
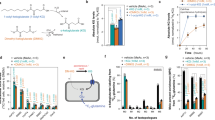
Samples containing 4 μM PHD2 (prolyl hydroxylase domain 2), 200 μM C-terminal oxygen dependent degradation domain (CODD) peptide, 5 mM (R)- or (S)-2HG, 50 μM Fe(II), 0/0.5/4/10 mM L-ascorbate in Hepes 50 mM pH 7.5 were incubated for 20 h (37 °C) and then analysed by MALDI-TOF-MS or subjected to amino acid analysis. Error bars represent s.d. of the mean of triplicate assays. 2OG (2-oxoglutarate) control incubations contained 300 μM 2OG instead of 2HG. (a) The PHD2-catalysed reaction. (b) Typical MALDI-TOF-MS spectra of CODD-OH (upper) and CODD (lower). (c) Dependence of PHD2/(R)-2HG-catalysed CODD hydroxylation on L-ascorbate and GSH (glutathione). (d) Dependence of PHD2/(S)-2HG-catalysed CODD hydroxylation reaction on L-ascorbate and GSH. (e,f) Amino acid analysis results (A: 2HG, Fe(II), L-ascorbate; B: CODD, 2HG, Fe(II), L-ascorbate; C: PHD2, 2HG, Fe(II), L-ascorbate; D: PHD2, CODD, 2HG, Fe(II), L-ascorbate; E: standard containing trans-4-hydroxyproline amino acid).
2OG oxygenase catalysis is often stimulated in vitro by L-ascorbate or other reducing agents25,26,27. L-ascorbate is required for PHD2 activity14,25,28, and is a standard component in PHD2 reaction incubations21,29 (Supplementary Table 1). We found that, under the steady-state conditions described above, the extent of PHD2/2HG-mediated CODD hydroxylation was dependent on L-ascorbate concentration; 20% CODD hydroxylation was observed in the presence of 10 mM L-ascorbate (Fig. 2c,d), ~4–7% CODD hydroxylation was observed in the presence of lower, possibly more physiologically relevant levels of L-ascorbate (0.1–0.3 mM, Supplementary Fig. 1a,b)30. L-Glutathione (thiol form, GSH) is a biologically important antioxidant that plays a role in cellular protection against oxidative stress31. We found that GSH was also able to promote PHD2/2HG-mediated CODD hydroxylation, albeit at higher concentrations than L-ascorbate (Fig. 2c,d). This observation is consistent with the reduced ability of GSH to stimulate 2OG-enabled PHD2 catalysis in vitro28.
To confirm 2HG-mediated prolyl-hydroxylation of CODD, amino acid analyses were performed; trans-4-hydroxylated proline was detected in hydrolysed PHD2/CODD assay mixtures where the assay was conducted in the presence of either (R)- or (S)-2HG (Fig. 2e,f). To test whether CODD hydroxylation was a product of catalysis, rather than a result of non specific oxidation by reactive oxidizing species (ROS) ‘leaking’ from the enzyme active site, the Fe(II)/2HG/L-ascorbate/CODD assay mixture was incubated with 0–100 μM H2O2 instead of PHD2. A +16 Da mass shift in the CODD MALDI-TOF-MS spectra was not observed under these conditions.
Studies on the binding of 2HG to PHD2
We were intrigued by the differences in our results when compared with those of Koivunen et al.21, including the incubation time required for CODD hydroxylation to be observed, the apparent dependence on the presence of reducing agent, and our observation that in the presence of L-ascorbate no significant difference in the extent of CODD hydroxylation was seen when using (S)- or (R)-2HG (Fig. 2c,d). We therefore carried out investigations to understand the nature of the 2HG/PHD2-mediated CODD hydroxylation, including the possibility of (R)-2HG oxidation being catalysed by PHD2, as proposed21.
To investigate whether (R)- and (S)-2HG bind to the catalytic domain of PHD2, biophysical techniques were employed. First, we employed an NMR assay that measures displacement of 2OG from the PHD2–Zn(II)–2OG complex (Zn(II) is a catalytically inactive substitute for Fe(II))32. Previous studies indicated very strong binding of 2OG to PHD2 (Kd=1 μM), which was not dependent on the presence of the prime substrate33. The NMR results showed that both (R)- and (S)-2HG exhibit very weak binding to PHD2 (determined Kd>1 mM and >0.6 mM respectively, Supplementary Fig. 2). Next, we used UV–vis spectroscopy to probe for the formation of a metal-to-ligand charge transfer feature: the anaerobic PHD2–Fe(II)–2OG complex absorbs at 530 nm25; however, addition of (R)- or (S)-2HG to PHD2–Fe(II) under anaerobic conditions did not result in the formation of any observable metal-to-ligand charge transfer feature (Supplementary Fig. 3). This suggests either the absence of 2HG binding within limits of detection, or significant differences in the interaction of 2HG with the active site metal compared with 2OG. We then conducted non-denaturing MS experiments to look for stable complex formation in the presence and absence of CODD prime substrate (Supplementary Fig. 4)34. (R)-2HG was not observed to bind to PHD2–Fe(II) by non-denaturing MS (in contrast with 2OG), either in the absence or presence of CODD, nor was there evidence for CODD hydroxylation on analysis of PHD2–Fe(II)–(R)-2HG–CODD as there was in the presence of 2OG. Overall, we were unable to find any evidence for efficient 2HG binding to PHD2.
We next considered the possibility of 2OG contamination of commercial batches of (R)- and (S)-2HG as a source of 2OG to elicit CODD hydroxylation. NMR and liquid chromatography-mass spectrometry (LC-MS) analyses revealed up to 0.02% (by molarity) 2OG in almost all batches (three out of four) of commercially purchased (R)-2HG, but not (S)-2HG (4 batches), which is equivalent to 1 μM 2OG in 5 mM (R)-2HG (as used in standard assay conditions) (Supplementary Fig. 5). 1 μM 2OG and 5 mM (R)-2HG support similar degrees of CODD hydroxylation, detectable under single turnover conditions (Supplementary Fig. 1c). We further subjected (R)-2HG to high-pressure liquid chromatography (HPLC) to remove 2OG contamination, then re-determined the IC50 values of both (R)- and (S)-2HG (Supplementary Fig. 6); consistent with our previous work14, we found that (R)- and (S)-2HG are both very weak inhibitors of PHD2 (IC50 values are 23 mM and 6 mM for (R)- and (S)-2HG, respectively). Thus, the low levels of CODD hydroxylation observed in the absence of L-ascorbate (Fig. 2c,d) may reflect the presence of contaminating 2OG in commercial (R)-2HG samples, and may also reflect differences in (R)- and (S)-2HG inhibition activities.
Reducing agent and iron promote non-enzymatic 2HG oxidation
Given the apparent ability of reducing agents to promote 2HG-mediated CODD hydroxylation (Fig. 2c,d), we used an antibody based assay, which gives a CODD-OH specific fluorescent signal35, to investigate the conditions required for CODD hydroxylation and to verify the sensitivity of our MS-based assay. Interestingly, we found that when PHD2/CODD were added after a 20 h preincubation of a mixture of (R)- or (S)-2HG, Fe(II) and L-ascorbate, a fluorescence signal was observed (indicative of CODD hydroxylation) after just 10 min of enzymatic activity. Fluorescence was greater in the samples containing L-ascorbate, and there was no difference between the two 2HG enantiomers (Supplementary Fig. 7). This result suggested that during the course of the preincubation 2HG was oxidized non-enzymatically to 2OG, to a level sufficient to support PHD catalysis. We therefore considered it possible that the mechanism of PHD2/2HG-mediated CODD hydroxylation was via non-enzymatic 2HG oxidation to 2OG followed by PHD2/2OG mediated CODD hydroxylation.
As reducing agents promote 2HG-mediated CODD hydroxylation (Fig. 2c,d), we investigated whether they are necessary for non-enzymatic conversion of 2HG to 2OG. Mixtures of (R)- or (S)-2HG with Fe(II) and different concentrations of L-ascorbate/GSH were incubated for 20 h (37 °C). 2OG formation was detected by LC-MS, and was observed in the samples containing L-ascorbate and GSH, without discrimination between (R)- or (S)-2HG (Fig. 3). Substituting with oxidized forms of L-ascorbate and glutathione (dehydroascorbate and GSSG) resulted in little or no 2OG formation (limited 2OG formation observed in the presence of dehydroascorbate could be due to its partial reduction to L-ascorbate under the assay conditions) (Fig. 3c,d). Reducing agents can therefore promote the non-enzymatic oxidation of 2HG to 2OG.
(a,b) LC-MS chromatograms showing 2OG formation under non-enzymatic conditions in the presence of GSH (glutathione). (c,d) Amounts of 2OG formed with different reducing agents. Error bars represent s.d. of the mean of triplicate assays. The reaction mixture was incubated for 20 h (37 °C) and contained 5 mM (R)- or (S)-2HG, 50 μM Fe(II), reducing agent (from bottom to top in (a,b): no reducing agent; 0.5 mM; 4 mM; 10 mM reducing agent). All panels in one stack in (a,b) have the same range of ion intensity (0–6.5·105).
We investigated the Fe(II)-dependence of non-enzymatic conversion of 2HG to 2OG and found that ascorbate- and glutathione-mediated 2HG oxidation requires the presence of either Fe(II) or Fe(III) (Supplementary Figs 8 and 9). The level of 2OG produced (detected by LC-MS) was dependent on the concentration of iron. We also investigated whether other transition metals could substitute for Fe(II)/Fe(III) in this reaction, and found that metals such as Zn(II), Mn(II), Co(II), Ni(II) and Cu(II) did not promote 2OG formation. NMR studies were performed to confirm that non-enzymatic oxidation of 2HG to 2OG is Fe(II)/(III)- and reducing agent-dependent (Supplementary Fig. 9). Consistent with the LC-MS results, the non-enzymatic conversion of 2HG to 2OG was observed, which was promoted by the presence of both L-ascorbate and Fe(II) (Fig. 4a). The time course of the reaction was investigated by NMR and showed a linear increase in 2OG generation over 20 h (Fig. 4b). Although, an excess of L-ascorbate/GSH relative to iron was found to be required to effectively enable the non-enzymatic reaction (Fig. 3c,d), physiologically relevant levels of these reducing agents30,36 are able to promote 2OG formation to levels that sustain CODD hydroxylation (Fig. 2c,d). Notably, low amounts of 2OG were also observed after incubation of 2HG and 10 mM L-ascorbate when no Fe(II) was added (Fig. 4a), which could be due to trace Fe(II) contamination present in reaction components including ascorbate and H2O, as revealed by inductively coupled plasma-mass spectrometry (ICP-MS) analyses (at least ~1 μM Fe(II)/Fe(III), Supplementary Table 2).
(a) 1H NMR spectra showing conversion of (S)-2HG (2-hydroxyglutarate) to 2OG (2-oxoglutarate). The 1H spectra show the 2OG CH2COCO2H resonance. Conditions (from top to bottom): 50 μM Fe(II) and 10 mM L-ascorbate; 5 mM (S)-2HG and 50 μM Fe(II); 5 mM (S)-2HG and 10 mM L-ascorbate; 5 mM (S)-2HG; 5 mM (S)-2HG, 50 μM Fe(II) and 10 mM L-ascorbate. Reaction mixtures were incubated for 20 h at 25 °C. All solutions were buffered with 50 mM Tris-d11 (pH 7.5) and 0.02% NaN3 in 90% H2O and 10% D2O. (b) Time course of non-enzymatic 2OG production as analysed by 1H NMR. Conditions: 5 mM (S)-2HG, 5 μM Fe(II), 2 mM L-ascorbate. (c) Succinate and 2OG production (total concentration) in the assay mixtures containing (R)-2HG. Conditions: 5 mM (R)-2HG, 5 μM Fe(II), 2 mM L-ascorbate, 4 μM PHD2 (prolyl hydroxylase domain 2), 128 μM CODD (C-terminal oxygen dependent degradation domain). Reaction mixtures were incubated for 20 h at 37 °C, PHD2 was precipitated by incubation in boiling water for 30 s before NMR analyses. All solutions were buffered with 50 mM Tris-d11 (pH 7.5) and 0.02% NaN3 in 90% H2O and 10% D2O. Error bars represent s.d. of the mean of triplicate assays. (d) Scheme for 2HG conversion to 2OG. SA, succinic acid; SSA, succinic semialdehyde.
H2O2 and iron promote non-enzymatic 2HG oxidation
In addition to 2OG, the NMR assays revealed formation of succinate and another oxidation product, which on the basis of NMR and MS studies was assigned as succinic semialdehyde (SSA) by comparison with a synthetic standard (Fig. 4d, Supplementary Figs 10 and 11) and LC-MS assays (Supplementary Fig. 12). Fenton type chemistry is viable for α-hydroxyacid oxidation37; therefore, the possibility of H2O2-mediated 2HG oxidation was studied. It was found that 2HG oxidation is enabled by H2O2 to give 2OG, SSA and succinate, and that Fe(II) or Fe(III) are required for this transformation (Fig. 5, Supplementary Fig. 13). Interestingly, while 2HG oxidation is relatively slow under the conditions of the Fe(II)/L-ascorbate system (16–20 h or more), the H2O2-driven reaction is substantially faster (2OG formation was apparent by NMR after 1 h of incubation) (Supplementary Fig. 14). The observed ratio between 2OG and SSA was found to be dependent on the amount of H2O2 added (Supplementary Figs 15 and 16). To investigate whether SSA is the product of further oxidation of 2OG, 2OG oxidation under Fe(II)/L-ascorbate and Fe(II)/H2O2 conditions was also studied. 2OG was found to be oxidized to succinate, but not to detectable levels of SSA (Supplementary Figs 17 and 18), indicating that SSA formation occurs via an alternative 2HG oxidative pathway. Detectable SSA oxidation to succinate was not observed under these conditions (Supplementary Fig. 14). Therefore the 2HG oxidation reaction most likely proceeds via iron-complex formation. Overall, the results reveal that 2HG is oxidized to 2OG and SSA via (at least) two alternative oxidative pathways, and the ratio of the products depends on the amount of H2O2 present.
Reaction of (R)-2HG with H2O2 in a 90% water 10% D2O mixture ((a) 5 mM (R)-2HG, (b) 5 mM (R)-2HG, 50 μM Fe(II); (c) 5 mM (R)-2HG, 10 mM L-ascorbate; (d) 5 mM (R)-2HG, 5 mM H2O2, (e) 5 mM (R)-2HG, 50 μM Fe(II), 5 mM H2O2; (f) 5 mM (R)-2HG, 10 mM L-ascorbate, 5 mM H2O2; (g) 5 mM (R)-2HG, 50 μM Fe(II), 10 mM L-ascorbate; (h) 5 mM (R)-2HG, 50 μM Fe(II), 10 mM L-ascorbate, 5 mM H2O2). 2HG, 2-hydroxyglutarate; 2OG, 2-oxoglutarate; SA, succinic acid; SSA, succinic semialdehyde. Addition of both Fe(II)/L-ascorbate/H2O2 leads to impaired 2OG and SSA formation, likely due to competition between different oxidative pathways and/or further oxidation of 2OG and SSA.
To investigate the role of iron in the observed peroxide driven reaction, we performed stopped-flow UV–vis spectroscopy studies. Transient species absorbing at 240–340 nm, which likely represent different iron oxidation states38, were observed during the H2O2-mediated 2HG oxidation (where the Fe(II)/(R)-2HG mixture was rapidly mixed with H2O2, Supplementary Figs 19 and 20). The formation and degradation of these species occurred over 5 min with the maximum time of their accumulation dependent on the H2O2 concentration, consistent with the NMR 2HG oxidation time course studies (Supplementary Fig. 14), which supports the proposal of iron-mediated conversion of 2HG to 2OG.
2HG-enabled promotion of catalysis by other 2OG oxygenases
2OG oxygenases can produce ROS as a result of incomplete catalytic cycles, which we envisaged could stimulate the observed H2O2-dependent 2HG oxidation27. To investigate this possibility, we studied formation of succinate and 2OG (from 2HG) in the presence of PHD2, with and without CODD substrate (Fig. 4c). The total combined concentrations of 2OG and succinate were not increased in the presence of PHD2, suggesting against 2HG oxidation due to ROS production from the PHD2 active site. Owing to consumption of 2OG (produced from 2HG) in the enzymatic reaction, levels of succinate were higher in the presence of PHD2 supporting the proposal that the non-enzymatic oxidation of 2HG occurs at a level sufficient to support PHD2 catalysis.
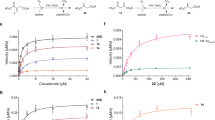
Given the reported ability of (R)-2HG to activate the PHDs but not other 2OG oxygenases21, we investigated whether the observed L-ascorbate-dependent 2HG to 2OG conversion is sufficient to support catalysis by oxygenases other than PHD2. We carried out activity assays with 2HG and L-ascorbate for other human 2OG oxygenases including BBOX1 (γ-butyrobetaine oxygenase, which catalyses the final step in L-carnitine biosynthesis), FIH and the catalytic domain of KDM4A (JMJD2A), an Nε-methyl lysine demethylase. NMR studies on BBOX1 showed that in the absence of 2OG, 2HG addition enables carnitine formation from γ-butyrobetaine (GBB) (Fig. 6). The observed carnitine levels were higher in the case of (R)-2HG, which is consistent with its lower inhibition potency compared with (S)-2HG for BBOX1 (IC50 values are 13.2 mM and 142 μM, for (R)-2HG and (S)-2HG, respectively)14. Consistent with the fluorescence-based assays on PHD2, addition of BBOX1 to a 2HG/Fe(II)/L-ascorbate mixture, reacted prior to the enzyme-incubation at 37 °C for 20 h, leads to higher levels of carnitine formation (Fig. 6). As for PHD2 this observation implies that the conversion of 2HG to 2OG is non-enzymatic, with 2OG being formed under the assay conditions. The assays with FIH and KDM4A did not reveal any 2HG-enabled enzyme catalysis at the same concentrations of 2HG/Fe(II)/L-ascorbate (for which activity was observed for PHD2 and BBOX1), as determined by MS assays. These observations can be rationalized by the 2HG-mediated inhibition of FIH and KDM4A catalysis, and/or by the increased sensitivity of FIH (relative to PHD2) to ROS, including H2O2, as reported14,21,39.
(a) Standards of (S)-2HG (2-hydroxyglutarate) and (e) (R)-2HG; (b) (S)-2HG or (f) (R)-2HG incubated with the assay mixture for 24 h prior to analysis; (c) (S)-2HG or (g) (R)-2HG incubated in the assay mixture with BBOX1 for 24 h prior to analysis; (d) (S)-2HG or (h) (R)-2HG incubated in the assay mixture for 24 h, then treated with BBOX1 and Fe(II) (1:1) and incubated for a further 2 h prior to analysis. The standard assay mixture contained 100 μM GBB (γ-butyrobetaine), 4 mM L-ascorbate, 50 μM Fe(II), 80 mM KCl in 50 mM Tris-d11 pH 7.5 containing 10% D2O. BBOX1 concentration was 100 nM. (S)-2HG and (R)-2HG were used at a final concentration of 5 mM.
As SSA was observed as a co-product of 2HG oxidation, we studied its influence on 2OG-dependent oxygenases such as PHD2, FIH, KDM4A and BBOX1 (to the extent of our knowledge no data on SSA inhibition of 2OG oxygenases have been reported). SSA caused weak inhibition of all these enzymes (IC50 values are 4.5 mM, 2.9 mM, 2.8 mM and 267 μM for FIH, PHD2, KDM4A and BBOX1, respectively (Supplementary Fig. 21)).
Discussion
Overall, our results clearly demonstrate that the non-enzymatic conversion of 2HG to 2OG can occur. The reaction is promoted by the combination of Fe(II)/Fe(III) and biologically relevant reducing agents such as L-ascorbate and GSH as well as by H2O2. The 2OG levels produced by non-enzymatic oxidation under our standard steady-state assay conditions are sufficient to support in vitro catalysis by some 2OG oxygenases (PHD2 and BBOX1), but not others (FIH and KDM4A). Although it is not possible to isolate and characterize these reactions within cells, the non-enzymatic conversion of 2HG to 2OG can occur in vitro with physiologically relevant concentrations of Fe(II/III)/reducing agents. We propose that the conditions used by Koivunen et al.21 likely resulted in non-enzymatic 2HG conversion to 2OG, and that contamination of 2HG with 2OG in commercial samples could also contribute to the 2HG-mediated CODD hydroxylation observed previously and in this study (Supplementary Fig. 1c). Although we cannot rule out that it occurs at levels below our limits of detection, we have accrued no evidence for direct 2OG oxygenase catalysed 2HG oxidation; indeed, biophysical studies imply that 2HG binds very weakly to PHD2. In addition, non-enzymatic oxidation should occur without discrimination between enantiomers, which is in accord with our observations.
The apparent preference for PHD2 catalysis to proceed in the presence of (R)- rather than (S)-2HG observed by Koivunen et al. can be rationalized by the observation that the (S) enantiomer is a better PHD2 inhibitor14 and/or by contamination of some commercial (R)-2HG batches by 2OG. Similarly with BBOX1, the increased activity with (R)-, rather than (S)-2HG, is consistent with the higher inhibition potency for the (S)-enantiomer14. The lack of observed 2HG catalysis with KDM4A and FIH may be due to the relatively stronger inhibition of the former by both (R)- and (S)-2HG, and the sensitivity of FIH to ROS, including H2O2 (refs 14, 39). Thus, even though the conversion of 2HG to 2OG is, at least predominantly, non-enzymatic and not enantioselective, the results of the incubation of 2HG with enzymes can appear enantioselective in a condition-dependent manner.
The exact mechanisms (likely more than one) of the non-enzymatic oxidation of 2HG are unclear, but probably proceed via iron-complexed ROS40,41,42. Higher rates of reaction were observed with H2O2 replacing L-ascorbate, suggesting the role of L-ascorbate/GSH is to generate ROS, though the exact mechanisms in operation are likely condition-dependent40. Further work can focus on whether the non-enzymatic conversion of 2HG to 2OG is of physiological relevance. Given the complexity of 2HG/2OG/TCA cycle metabolism, it is difficult to distinguish between non-enzyme and enzyme-catalysed reactions in cells (Fig. 7)1. However, it is well-established that many proteins undergo non-enzymatic oxidative modifications43. As indicated in Fig. 7, it is possible that non-enzymatic reactions may augment the network of identified enzyme-catalysed reactions related to 2HG. The concentrations of the reducing agents at which we observed significant conversion in vitro are probably above those for individual reducing agents in cells30. However, taking into account the total cellular concentration of reducing agents capable to promote 2HG oxidation we cannot exclude the possibility of them being sufficient to promote the reaction. Although our in vitro observations indicate that supra-physiological levels of free Fe(II)/Fe(III) (>5 μM)36 are required for significant levels of 2OG to be formed from 2HG, it is possible that 2HG oxidation may be mediated by complexed iron. Given that non-enzymatic 2HG oxidation is promoted by H2O2 and likely some other ROS, this reaction may be relatively more important under conditions of oxidative stress, mediated by elevated ROS, such as that occurring in hypoxia44,45,46,47 or in environments of high H2O2 concentration, for example, as occurring in peroxisomes.
Red arrows represent non-enzymatic transformations observed in this study, black arrows represent identified metabolic pathways.1 L-maIDH, L-malate dehydrogenase; L-2-HGDH, L-2-hydroxyglutarate dehydrogenase; IDH2, isocitrate dehydrogenase 2; D-2HGDH, D-2-hydroxyglutarate dehydrogenase; HOT, hydroxyacid-oxoacid transhydrogenase enzyme; GBH, γ-hydroxybutyrate; SSA reductase, succinic semialdehyde reductase enzyme; SSADH, succinic semialdehyde dehydrogenase; GABA, γ-aminobutyric acid.
Interestingly, recent work indicates reduction of antioxidant concentrations in IDH-mutated mice12. It is thus possible that non-enzymatic Fenton-type chemistry including 2HG/2OG and related metabolites may be involved in the regulation of ROS levels. Indeed, although IDH1/2 mutations have been identified in many cancers5, the mechanisms by which they relate to tumorigenesis are unclear. Although, the balance of current evidence suggests that direct inhibition of the HIF hydroxylases by (R)-2HG is probably not physiologically important12,14,15, the inhibition of chromatin-modifying (or other) 2OG oxygenases by (R)-2HG with pathophysical consequences remains plausible14,15,16,48. It is also proposed that ‘gain-of-function’ IDH mutations may result in depletion of antioxidants including L-ascorbate and GSH, together with perturbation of the NADP+/NADPH ratio, leading to physiologically relevant oxidative stresses5. The non-enzymatic chemistry identified here may contribute to the development of such stresses.
SSA is a metabolite produced by oxidation of γ-aminobutyric acid (GABA)49 (Fig. 7). A deficiency in SSA dehydrogenase results from genetic disorders leading to accumulation of SSA in cells where it cannot be further oxidized to succinic acid and subsequently utilized in the TCA cycle50. The observation that SSA and succinic acid can be concomitantly produced with 2OG by 2HG oxidation is of interest. The reactive nature of SSA suggests that a potential role for it in cancer metabolism merits investigation. Given that we found that the ratio of 2OG to SSA observed was condition-dependent, it will also be of interest to investigate whether SSA can be detected or its levels vary in cells under conditions of redox stress such as in tumour cells or in highly oxidizing environments such as peroxisomes. It is also notable that in some cases the inhibition potency of SSA for the studied enzymes was higher than for (R)-2HG, raising the possibility that it might inhibit enzymes in cells.
Our results have consequence for the interpretation of the important cellular observations of Koivunen et al.21 Both HIF-1α and HIF-2α levels were reported to be lower in mid-passage astrocyte cells, in which mutant IDH1 R132H protein was present, compared with wild-type cells. Our results imply that this is unlikely to be due to direct use of (R)-2HG as a PHD2 co-substrate. Although it does not rationalize the reported resistance of the IDH1 mutant cells to the NOG (N-oxalyl glycine) precursor DMOG (dimethyloxalyl glycine)21, it is possible that the observed effects on HIF-α levels in the astrocyte cells were in part a manifestation of the effect of increased 2HG levels on chromatin-modifying enzymes, including the JmjC demethylases and TET enzymes. The proposal that some of the observed effects on the IDH1 mutant on HIF-α might be mediated via ‘epigenetic processes’ may relate to the observation that HIF-α levels were reported to normalize in later passages (more than passage 20) despite continued production of the mutant IDH1 enzyme and (R)-2HG. We hope that our results will stimulate further work on the cellular mechanism by which 2HG stimulates PHD activity.
Methods
Materials
(R)- and (S)-2-hydroxyglutaric acid disodium salts were from Sigma-Aldrich, UK. The HIF-1α(556–574) CODD peptide, where Met residues in the initial HIF-1α sequence were replaced with Ala, (DLDLEALAPYIPADDDFQL) and HIF-1α(788–822) CAD peptide (DESGLPQLTSYDCEVNAPIQGSRNLLQGEELLRAL) were from GL Biochem (Shanghai) Ltd, China. H3K9me3 (ARK(me3)STGGK) was synthesized as described51. Recombinant constructs of catalytic domain of KDM4A (residues 1–359), PHD2 (residues 181–426) and full-length FIH were produced as N-terminally His6-tagged protein in Escherichia coli BL21(DE3) cells and purified by Ni-affinity, size-exclusion and cation exchange chromatography as described52,53,54. BBOX1 was expressed and purified as described by Leung et al.55
Enzymatic assays
PHD2, FIH and KDM4A assays were performed at 37 °C as reported14. The substrate for PHD2 was the HIF-1α(556–574) CODD 19-mer peptide fragment, wherein the Met residues of the human HIF-1α sequence were substituted for Ala residues to avoid non specific Met oxidation to give a sulfoxide (+16 Da). The substrate for FIH was the HIF-1α(788–822) CAD 35-mer peptide fragment. The substrate for KDM4A was the H3K9me3 peptide. The conditions are described in the Supplementary Table 3. Samples were analysed using a Waters Micromass MALDI micro MX mass spectrometer (Micromass UK Ltd., Manchester, UK) in reflectron mode with flight tube voltage at 12 kV and reflectron voltage at 5.2 kV. Data were analysed using MassLynx version 4.1. BBOX1 activity assays were performed by NMR. Unless otherwise stated the standard assay mixture contained 100 nM BBOX1, 100 μM GBB (γ-butyrobetaine), 4 mM L-ascorbate, 50 μM (NH4)2Fe(SO4)2 and 80 mM KCl in 50 mM Tris-d11 (pH 7.5) containing 10% D2O.
SSA inhibition studies were performed by MALDI-TOF-MS and NMR (human BBOX1) using the described conditions14. Inhibition of human BBOX1 by SSA was measured by observing carnitine formation by 1H NMR (Bruker Avance 700). Control reaction mixtures contained 40 μM 2-oxoglutarate disodium salt, 40 μM γ-butyrobetaine hydrochloride, 40 μM ammonium (NH4)2Fe(SO4)2, 500 μM L-ascorbate, 40 mM KCl and 10% D2O in 50 mM pH 7.5 Tris-d11. Reactions were initiated by addition of human BBOX1 (final concentration 100 nM), the first 1H NMR spectrum was acquired after 10 min.
LC-MS experiments
For LC-MS assays, reaction mixtures typically contained 5 mM (R)- or (S)-2HG, 50 μM (NH4)2Fe(SO4)2 and variable concentrations (0–10 mM) L-ascorbate/GSH prepared in 50 mM Hepes (pH 7.5), and were incubated for 20 h (37 °C). 2OG production was subsequently measured. Chromatographic separation was performed at 50 °C using a Waters ACQUITY BEH Amide 1.7 μm 2.1 × 100 mm column and a Waters ACQUITYTM ultra-performance liquid chromatography system (Waters Corp., Milford, MA, USA). The following eluents were used: mobile phase A: 10% H2O, 90% acetonitrile (v/v), 10 mM ammonium formate; mobile phase B: 50% H2O, 50% acetonitrile (v/v), 10 mM ammonium formate. The elution gradient was 0–10.0 min linear from 10–30% B, and 10.0–12.0 min at 10% B for re-equilibration of the column. A constant flow rate of 0.4 ml min−1 was used. This was directly coupled to a Waters Quattro micro (Micromass UK Ltd) triple quadrupole mass spectrometer in negative ionization mode. The desolvation temperature was set to 250 °C and the source temperature to 120 °C.
NMR experiments
NMR experiments were conducted at 298 K using a Bruker Avance III 700 MHz spectrometer equipped with a 5 mm inverse TCI cryoprobe and 3 mm diameter MATCH NMR tubes (Bruker). Water suppression was achieved by the excitation sculpting method using a 2 ms 180 degree selective Sinc1.1000 pulse at the H2O frequency. Typical experimental parameters were as follows: number of transients 256, relaxation delay 2 s. Unless otherwise stated all samples were 400 μl final volume, consisting of 90% H2O and 10% D2O with the following concentrations in the reaction mixture: 5 mM 2HG, 50 μM (NH4)2Fe(SO4)2 and 10 mM L-ascorbate. The non-enzymatic reaction was started by addition of hydrogen peroxide with 5 mM as standard final concentration and then monitored for 2 h by 1H NMR at 298 K.
The CPMG (Carr–Purcell–Meiboom–Gill) displacement experiments were conducted using the AV700 instrument and 3 mm MATCH NMR tubes32. The PROJECT-CPMG sequence (90°x–[τ–180°y–τ–90°y–τ–180°y–τ]n–acq) was applied as described by Aguilar et al.56 Typical experimental parameters were as follows: total echo time 48 ms (τ=2 ms, n=6), acquisition time 2.94 s, relaxation delay 2 s and number of transients 128. Water suppression was achieved by presaturation.
UV–vis spectroscopy experiments
Absorption spectra were acquired using a Cary Varian 4000 UV–vis spectrophotometer at 22 °C with 0.1 mM PHD2, 0.1 mM Fe(II) and 0.5 mM 2OG or (R)/(S)-2HG in deoxygenated 50 mM Hepes (pH 7.5). The buffer and enzyme were deoxygenated according to the described procedure22. All other solutions were prepared in an anaerobic glove box (Belle Ltd., UK).
Non-denaturing ESI-MS experiments
Non-denaturing ESI-MS experiments were performed as described57. PHD2 was buffer-exchanged into 15 mM ammonium acetate (pH 7.5). Data were analysed using MassLynx 4.0.
Amino acid analysis
Amino acid analysis was performed as previously described58. An extracted ion chromatogram was created using m/z value corresponding to AQC-derivatized hydroxyproline (304±0.2 Da). Three chromatograms were acquired for every sample. The limit of detection for the method was determined to be ~1 pmol μl−1.
Stopped-flow UV–vis spectroscopy experiments
Solutions of 10 mM (R)-2HG, 0.2 mM Fe(II) were prepared in Milli-Q water and rapidly mixed with 2 mM H2O2 at 1:1 ratio in a stopped-flow apparatus (Applied Photophysics, Leatherhead, UK). Subsequent analysis used an SX20 stopped-flow spectrometer and spectra were recorded over time up to 500 s using a photodiode array detector. All assays were performed at least in triplicate.
Synthesis of SSA
SSA was prepared from methyl 4,4-dimethoxybutyrate (Sigma-Aldrich). The starting material (100 mg) was dissolved in 5 ml of tetrahydrofurane and treated with 3 ml of 1 M lithium hydroxide. The reaction mixture was stirred overnight and then tetrahydrofurane was removed in vacuo. The water phase was acidified with concentrated HCl and stirred for a further 6 h. The solution was concentrated to 1 ml in vacuo and purified on HPLC (reverse phase column, gradient 10% of B in 15 min where A: 0.05% TFA in H2O, B: 0.1% TFA in acetonitrile, retention time 9 min). Fractions containing the product were freeze-dried to yield SSA as an off-white oil.
1H NMR (500 MHz, D2O) δ p.p.m. 5.00 (t, J=5.5), 2.40 (t, J=7.5), 1.82 (dt, J1=7.5, J2=5.5); 13C NMR (125 MHz, D2O) δ p.p.m. 177.5, 89.1, 31.9, 28.8. LC-MS analysis retention time is 1.15 min (Waters ACQUITY BEH Amide 1.7 μm 2.1 × 100 mm column, mobile phase A: 10% H2O, 90% acetonitrile (v/v), 10 mM ammonium formate; mobile phase B: 50% H2O, 50% acetonitrile (v/v), 10 mM ammonium formate), mass—101.2 (negative ion mode).
Additional information
How to cite this article: Tarhonskaya, H. et al. Non-enzymatic chemistry enables 2-hydroxyglutarate-mediated activation of 2-oxoglutarate oxygenases. Nat. Commun. 5:3423 doi: 10.1038/ncomms4423 (2014).
References
Kranendijk, M., Struys, E., Salomons, G., van der Knaap, M. & Jakobs, C. Progress in understanding 2-hydroxyglutaric acidurias. J. Inherited. Metab. Dis. 35, 571–587 (2012).
Matsunaga, H. et al. IDH1 and IDH2 have critical roles in 2-hydroxyglutarate production in D-2-hydroxyglutarate dehydrogenase depleted cells. Biochem. Biophys. Res. Commun. 423, 553–556 (2012).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 465, 966–966 (2010).
Yang, H., Ye, D., Guan, K.-L. & Xiong, Y. IDH1 and IDH2 mutations in tumorigenesis: mechanistic insights and clinical perspectives. Clin. Cancer Res. 18, 5562–5571 (2012).
Ichimura, K. Molecular pathogenesis of IDH1/2 mutations in gliomas. Brain Tumor Pathol. 29, 131–139 (2012).
Losman, J.-A. et al. (R)-2-hydroxyglutarate is sufficient to promote leukemogenesis and its effects are reversible. Science 339, 1621–1625 (2013).
Walport, L. J., Hopkinson, R. J. & Schofield, C. J. Mechanisms of human histone and nucleic acid demethylases. Curr. Opin. Chem. Biol. 16, 525–534 (2012).
Semenza, G. L. HIF-1 and mechanisms of hypoxia sensing. Curr. Opin. Cell Biol. 13, 167–171 (2001).
Schofield, C. J. & Ratcliffe, P. J. Oxygen sensing by HIF hydroxylases. Nat. Rev. Mol. Cell Biol. 5, 343–354 (2004).
Hewitson, K. S. et al. Structural and mechanistic studies on the inhibition of the hypoxia-inducible transcription factor hydroxylases by tricarboxylic acid cycle intermediates. J. Biol. Chem. 282, 3293–3301 (2007).
Zhao, S. et al. Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1α. Science 324, 261–265 (2009).
Sasaki, M. et al. D-2-hydroxyglutarate produced by mutant IDH1 perturbs collagen maturation and basement membrane function. Genes Dev. 26, 2038–2049 (2012).
Williams, S. et al. R132H-mutation of isocitrate dehydrogenase-1 is not sufficient for HIF-1α upregulation in adult glioma. Acta Neuropathol. (Berl.) 121, 279–281 (2011).
Chowdhury, R. et al. The oncometabolite 2-hydroxyglutarate inhibits histone lysine demethylases. EMBO Rep. 12, 463–469 (2011).
Xu, W. et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate-dependent dioxygenases. Cancer Cell 19, 17–30 (2011).
Lu, C. et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483, 474–478 (2012).
Carmeliet, P. et al. Role of HIF-1a in hypoxia-mediated apoptosis, cell proliferation and tumour angiogenesis. Nature 394, 485–490 (1998).
Mack, F. A. et al. Loss of pVHL is sufficient to cause HIF dysregulation in primary cells but does not promote tumor growth. Cancer Cell 3, 75–88 (2003).
Song, L. P. et al. Hypoxia-inducible factor-1α-induced differentiation of myeloid leukemic cells is its transcriptional activity independent. Oncogene 27, 519–527 (2007).
Acker, T. et al. Genetic evidence for a tumor suppressor role of HIF-2α. Cancer Cell 8, 131–141 (2005).
Koivunen, P. et al. Transformation by the (R)-enantiomer of 2-hydroxyglutarate linked to EGLN activation. Nature 483, 484–488 (2012).
Price, J. C., Barr, E. W., Tirupati, B., Bollinger, J. M. & Krebs, C. The first direct characterization of a high-valent iron intermediate in the reaction of an α-ketoglutarate-dependent dioxygenase: a high-spin Fe(IV) complex in taurine/α-ketoglutarate dioxygenase (TauD) from Escherichia coli. Biochemistry 42, 7497–7508 (2003).
Clifton, I. J. et al. Structural studies on 2-oxoglutarate oxygenases and related double-stranded β-helix fold proteins. J. Inorg. Biochem. 100, 644–669 (2006).
van der Donk, W. A., Krebs, C. & Bollinger, J. M. Jr Substrate activation by iron superoxo intermediates. Curr. Opin. Struct. Biol. 20, 673–683 (2010).
Flashman, E. et al. Evidence for the slow reaction of hypoxia-inducible factor prolyl hydroxylase 2 with oxygen. FEBS J. 277, 4089–4099 (2010).
Myllylä, R., Majamaa, K., Günzler, V., Hanauske-Abel, H. M. & Kivirikko, K. I. Ascorbate is consumed stoichiometrically in the uncoupled reactions catalyzed by prolyl-4-hydroxylase and lysyl hydroxylase. J. Biol. Chem. 259, 5403–5405 (1984).
Hausinger, R. P. Fe(II)/α-ketoglutarate-dependent hydroxylases and related enzymes. Crit. Rev. Biochem. Mol. Biol. 39, 21–68 (2004).
Flashman, E., Davies, S. L., Yeoh, K. K. & Schofield, C. J. Investigating the dependence of the hypoxia-inducible factor hydroxylases (factor inhibiting HIF and prolyl hydroxylase domain 2) on ascorbate and other reducing agents. Biochem. J. 427, 135–142 (2010).
Koivunen, P., Hirsilä, M., Kivirikko, K. I. & Myllyharju, J. The length of peptide substrates has a marked effect on hydroxylation by the hypoxia-inducible factor prolyl 4-hydroxylases. J. Biol. Chem. 281, 28712–28720 (2006).
Du, J., Cullen, J. J. & Buettner, G. R. Ascorbic acid: chemistry, biology and the treatment of cancer. Biochim. Biophys. Acta Rev. Cancer 1826, 443–457 (2012).
Townsend, D. M., Tew, K. D. & Tapiero, H. The importance of glutathione in human disease. Biomed. Pharmacother. 57, 145–155 (2003).
Leung, I. K. H. et al. Reporter ligand NMR screening method for 2-oxoglutarate oxygenase inhibitors. J. Med. Chem. 56, 547–555 (2013).
Leung, I. K. H., Flashman, E., Yeoh, K. K., Schofield, C. J. & Claridge, T. D. W. Using NMR solvent water relaxation to investigate metalloenzyme−ligand binding interactions. J. Med. Chem. 53, 867–875 (2010).
McNeill, L. A. et al. Hypoxia-inducible factor prolyl hydroxylase 2 has a high affinity for ferrous iron and 2-oxoglutarate. Mol. Biosyst. 1, 321–324 (2005).
Yeoh, K. K. et al. Dual-action inhibitors of HIF prolyl hydroxylases that induce binding of a second iron ion. Org. Biomol. Chem. 11, 732–745 (2013).
Hider, R. C. & Kong, X. Iron speciation in the cytosol: an overview. Dalton. Trans. 42, 3220–3229 (2013).
Walling, C. & Amarnath, K. Oxidation of mandelic acid by Fenton's reagent. J. Am. Chem. Soc. 104, 1185–1189 (1982).
Knight, R. J. & Sylva, R. N. Spectrophotometric investigation of iron(III) hydrolysis in light and heavy water at 25 °C. J. Inorg. Nucl. Chem. 37, 779–783 (1975).
Masson, N. et al. The FIH hydroxylase is a cellular peroxide sensor that modulates HIF transcriptional activity. EMBO Rep. 13, 251–257 (2012).
Buettner, G. R. & Jurkiewicz, B. A. Catalytic metals, ascorbate and free radicals: combinations to avoid. Radiat. Res. 145, 532–541 (1996).
Stadtman, E. R. Oxidation of free amino-acids and amino-acid residues in proteins by radiolysis and by metal-catalyzed reactions. Annu. Rev. Biochem. 62, 797–821 (1993).
Mecinović, J., Hamed, R. B. & Schofield, C. J. Iron-mediated cleavage of C-C bonds in vicinal tricarbonyl compounds in water. Angew Chem. Int. Ed. 48, 2796–2800 (2009).
Wall, S. B., Oh, J.-Y., Diers, A. R. & Landar, A. Oxidative modification of proteins: An emerging mechanism of cell signaling. Front. Physiol. 3, 369 (2012).
Liu, L., Wise, D. R., Diehl, J. A. & Simon, M. C. Hypoxic reactive oxygen species regulate the integrated stress response and cell survival. J. Biol. Chem. 283, 31153–31162 (2008).
Chandel, N. S. et al. Mitochondrial reactive oxygen species trigger hypoxia-induced transcription. Proc. Natl Acad. Sci. USA 95, 11715–11720 (1998).
Guzy, R. D. et al. Mitochondrial complex III is required for hypoxia-induced ROS production and cellular oxygen sensing. Cell Metab. 1, 401–408 (2005).
Mansfield, K. D. et al. Mitochondrial dysfunction resulting from loss of cytochrome c impairs cellular oxygen sensing and hypoxic HIF-α activation. Cell Metab. 1, 393–399 (2005).
Figueroa, M. E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18, 553–567 (2010).
Kim, K. J. et al. Succinic semialdehyde dehydrogenase: biochemical-molecular-clinical disease mechanisms, redox regulation, and functional significance. Antioxid. Redox Signaling 15, 691–718 (2011).
Pearl, P. L., Novotny, E. J., Acosta, M. T., Jakobs, C. & Gibson, K. M. Succinic semialdehyde dehydrogenase deficiency in children and adults. Ann. Neurol. 54, S73–S80 (2003).
Hopkinson, R. J., Hamed, R. B., Rose, N. R., Claridge, T. D. W. & Schofield, C. J. Monitoring the activity of 2-oxoglutarate dependent histone demethylases by NMR spectroscopy: direct observation of formaldehyde. Chembiochem. 11, 506–510 (2010).
Chowdhury, R. et al. Structural basis for binding of hypoxia-inducible factor to the oxygen-sensing prolyl hydroxylases. Structure 17, 981–989 (2009).
Elkins, J. M. et al. Structure of factor-inhibiting hypoxia-inducible factor (HIF) reveals mechanism of oxidative modification of HIF-1α. J. Biol. Chem. 278, 1802–1806 (2003).
Ng, S. S. et al. Crystal structures of histone demethylase JMJD2A reveal basis for substrate specificity. Nature 448, 87–91 (2007).
Leung, I. K. H. et al. Structural and Mechanistic Studies on γ-Butyrobetaine Hydroxylase. Chem. Biol. 17, 1316–1324 (2010).
Aguilar, J. A., Nilsson, M., Bodenhausen, G. & Morris, G. A. Spin echo NMR spectra without J modulation. Chem. Commun. 48, 811–813 (2012).
Mecinović, J., Chowdhury, R., Flashman, E. & Schofield, C. J. Use of mass spectrometry to probe the nucleophilicity of cysteinyl residues of prolyl hydroxylase domain 2. Anal. Biochem. 393, 215–221 (2009).
Yang, M. et al. Substrate selectivity analyses of factor inhibiting hypoxia-inducible factor. Angew Chem. Int. Ed. 52, 1700–1704 (2013).
Acknowledgements
We thank the Wellcome Trust, European Research Council, Biotechnology and Biological Sciences Research Council, the British Heart Foundation, Clarendon-St Hugh’s College-W. Louey Scholarship (H.T.), Dulverton Trust (A.M.R.) and a Royal Society Dorothy Hodgkin Fellowship (E.F.) for financial support. We also thank Dr E. Sánchez-Fernández for the synthesis of H3K9me3 peptide, Dr G. Kochan for the purification of BBOX1, and Mr P. Holdship (Department of Earth Sciences, University of Oxford) for conducting ICP-MS analysis.
Author information
Authors and Affiliations
Contributions
H.T. performed enzymatic assays, UV–vis experiments, non-enzymatic LC-MS studies and wrote the manuscript. A.M.R. carried out BBOX1 assays, non-enzymatic NMR studies, organic synthesis and wrote the manuscript. I.K.H.L. performed NMR experiments, including binding studies, under the supervision of T.D.W.C. N.D.L. carried out amino-acid analysis and LC-MS method development under the supervision of J.S.O.M. M.C.C. performed AlphaScreen assays. A.K. carried out initial experiments and provided KDM4A enzyme. E.F. and C.J.S. designed and supervised the study and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-21, Supplementary Tables 1-3 and Supplementary References (PDF 1333 kb)
Rights and permissions
About this article
Cite this article
Tarhonskaya, H., Rydzik, A., Leung, I. et al. Non-enzymatic chemistry enables 2-hydroxyglutarate-mediated activation of 2-oxoglutarate oxygenases. Nat Commun 5, 3423 (2014). https://doi.org/10.1038/ncomms4423
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms4423
This article is cited by
-
The immunometabolite S-2-hydroxyglutarate exacerbates perioperative ischemic brain injury and cognitive dysfunction by enhancing CD8+ T lymphocyte-mediated neurotoxicity
Journal of Neuroinflammation (2022)
-
The return of quiescence metabolites
Nature Cell Biology (2021)
-
Heart failure—emerging roles for the mitochondrial pyruvate carrier
Cell Death & Differentiation (2021)
-
Oncometabolites in renal cancer
Nature Reviews Nephrology (2020)
-
Coupling Krebs cycle metabolites to signalling in immunity and cancer
Nature Metabolism (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.