Abstract
Chlamydiae are important pathogens and symbionts with unique cell biological features. They lack the cell-division protein FtsZ, and the existence of peptidoglycan (PG) in their cell wall has been highly controversial. FtsZ and PG together function in orchestrating cell division and maintaining cell shape in almost all other bacteria. Using electron cryotomography, mass spectrometry and fluorescent labelling dyes, here we show that some environmental chlamydiae have cell wall sacculi consisting of a novel PG type. Treatment with fosfomycin (a PG synthesis inhibitor) leads to lower infection rates and aberrant cell shapes, suggesting that PG synthesis is crucial for the chlamydial life cycle. Our findings demonstrate for the first time the presence of PG in a member of the Chlamydiae. They also present a unique example of a bacterium with a PG sacculus but without FtsZ, challenging the current hypothesis that it is the absence of a cell wall that renders FtsZ non-essential.
Similar content being viewed by others
Introduction
Chlamydiae are members of the Planctomycetes–Verrucomicrobia–Chlamydiae (PVC) bacterial superphylum1. Like most other bacteria, some PVC bacteria are already known to possess peptidoglycan (PG), that is, chains of alternating N-acetylglucosamine and N-acetylmuramic acid sugars crosslinked by short peptides. PVC bacteria also display striking eukaryote-like and archaea-like cell biological features, which have suggested intriguing hypotheses about their role in cellular evolution2,3.
In Verrucomicrobia and almost all other bacteria, septal PG synthesis is orchestrated by the FtsZ cytoskeleton4,5. In contrast, Planctomycetes lack both PG and FtsZ5. This fits to the notion that PG-loss renders FtsZ dispensable, like in mycoplasmas6 or L-form bacilli7. Although from genome sequences it is clear that chlamydiae do not possess FtsZ, the presence or absence of chlamydial PG has been highly controversial8,9,10,11,12,13,14,15,16. One early study reported the colorimetric detection of muramic acid in chlamydiae14, but more reliable chromatographic methods subsequently failed to confirm this result11,15. All attempts to purify chlamydial sacculi have failed10,12 and no periplasmic density layers have been detected between the inner and outer membranes of chlamydiae by electron microscopy (EM) (including for instance17,18,19). The apparent absence of PG in chlamydiae is surprising, however, as despite their highly reduced genomes, a nearly complete pathway for the synthesis of PG is present in the genomes of all chlamydiae13. In addition, several of the chlamydial PG biosynthetic enzymes have been characterized and shown to be functional in vitro and in complementation assays20,21,22,23,24,25.
Here we look for the evidence of PG cell walls in two diverse and deeply rooting chlamydiae26, Protochlamydia amoebophila and Simkania negevensis. Through electron cryotomography (ECT), biochemical purification, enzymatic digestion, mass spectrometry (MS), fluorescence microscopy and antibiotic treatment, we show that P. amoebophila are indeed surrounded by sacculi containing a new type of PG. In contrast, no evidence of PG is found in S. negevensis. These results prove that some chlamydiae do in fact synthesize PG sacculi, explaining the presence of PG-synthetic genes, but raising new questions about the identity and purpose of the modifications and the mechanisms of cell division in the absence of FtsZ.
Results
ECT of the chlamydial cell envelope
Two diverse and deeply rooting members of the chlamydial phylum, S. negevensis and P. amoebophila, were imaged by ECT in a near-native state. Bacteria were purified from amoeba cultures, plunge-frozen and 25 and 20 tomograms were collected of intact cells (Fig. 1). Density profiles through the cell envelopes of the two species were quite different. Although four layers were resolved in Simkania envelopes (Fig. 1b,c), five layers were resolved in Protochlamydia (Fig. 1e,f). Because the individual leaflets of lipid bilayers can be resolved in some cryotomograms, especially when the images are taken close to focus, in the case of Simkania, it is unclear whether the four layers represent the two leaflets of the outer and inner membranes (‘O’ and ‘A’ being the two leaflets of the outer membrane, and layers ‘B’ and ‘I’ being the two leaflets of the inner membrane), or whether one or more of these layers are non-membranous. The facts that layers O and A have fairly similar contrast and are consistently spaced even through the undulations are consistent with them being two leaflets of a single (outer) bilayer membrane. Their separation (~5 nm), however, is much larger than typical phospholipid bilayer membranes, whose two density peaks (from the phospholipid head groups) are only 3.7–4 nm apart27. Similarly, Simkania layers B and I may be the two leaflets of a single (inner) membrane, as they have similar contrast and a consistent spacing, but again they appear too far apart. In contrast to the Simkania envelope, the profile of Protochlamydia surprisingly resembled those of other Gram-negative bacteria with two membranes and a PG cell wall28,29,30. Between the Protochlamydia outer and inner membranes (labelled ‘O’ and ‘I’, respectively) there appeared to be three additional layers (labelled ‘C’–‘E’). The similar-looking three layers in Treponema pallidum (from the outside in) were identified as proteinaceous (lipoproteins), PG and again proteinaceous29. By analogy this suggests that layer C is composed of lipoproteins (perhaps connecting the outer membrane to the cell wall) and other outer-membrane-associated proteins, layer D is a cell wall and layer E is composed of lipoproteins and inner-membrane-associated proteins (perhaps including the penicillin-binding proteins responsible for cell wall synthesis, the lipoprotein OmcA and the cysteine-rich protein OmcB). Although other interpretations remain possible (cysteine-rich disulphide-crosslinked envelope proteins have been suggested to be the functional equivalent of PG in chlamydiae31), the most important and clear observation was that Protochlamydia exhibit a distinct periplasmic layer (D).
Simkania (a–c) and Protochlamydia (d–f) cells were purified from asynchronously infected amoeba cultures, plunge-frozen and imaged by ECT. Shown are tomographic slices through reticulate bodies (a,d and b,e enlarged) and corresponding density profiles (c,f) of the cell envelopes. Profiles are enlarged, aligned and cropped relative to the outer membrane. Distances between peaks (in nm) are indicated. In contrast to the Simkania profile, the Protochlamydia profile resembles those of other bacteria with PG cell walls (see text for a full discussion of each profile and layer). Scale bar, 100 nm in a,d and 20 nm in b,c,e,f.
Purification and imaging of sacculi
In order to explore whether any of the observed periplasmic layers consisted of PG, we attempted to purify sacculi by boiling chlamydial cells (obtained from asynchronously infected amoeba cultures) in 4% sodium dodecyl sulphate. Strikingly, in three independent experiments, we observed sacculus-like structures in preparations from Protochlamydia (Fig. 2a–d), but not from Simkania (two experiments). Protochlamydia sacculi diameters (679 nm ±34 s.d. n=10) and morphologies matched the size and shape of intact cells. Protochlamydia sacculi had one or two 5–7 nm thick layers (arrowheads in Fig. 2e), plus mesh-like (up to 30 nm long) high-density aggregates attached to the outside (arrows in Fig. 2d).
Cryoprojections (a) and tomographic slices (b–e) through sacculi-like structures purified from Protochlamydia cells. We were unable to obtain similar structures from Simkania. Sacculi had one or two layers (arrowheads) plus short high-density filaments (arrows) on the outside (c enlarged in d,e). Sacculi were digested, reduced and separated by HPLC (f). MS analysis of peaks 1–3 (neutral masses indicated in Da) indicated the presence of modified PG (see also Table 1 and Supplementary Fig. S1). Scale bars, 100 nm.
Digestion and biochemical analyses of sacculi
To check for the presence of PG in the purified sacculi, we digested the samples with cellosyl, a glycan strand-cleaving PG muramidase. Cellosyl released soluble material from insoluble sacculi, which was reduced with sodium borohydride and analysed by high-pressure liquid chromatography (HPLC) using conditions for separating muropeptides32. The chromatogram (Fig. 2f) showed three main peaks in the monomeric region (20–50 min) and many peaks after 60 min that are poorly separated at higher retention time (>75 min) forming a ‘hump’, which is typical for highly crosslinked and/or incompletely digested PG material33. The retention times and overall pattern of cellosyl digestion products were different, however, from those of muropeptide mixtures obtained from other Gram-positive and Gram-negative bacteria.
To characterize this material, the three main cellosyl products in the monomeric region and one well-separated main product at the beginning of the ‘hump’ region were analysed by MS. The determined neutral masses of the earlier three products were higher than what would be expected for monomeric muropeptides, but the masses of products 1 and 2 and of products 2 and 3 both differed by 71 Da, a typical feature of monomeric muropeptides with a tri-, tetra- and pentapeptide, respectively, due to the presence of none, one or two D-alanine residues (Fig. 2f). In MS/MS analysis, the three products were fragmented in a similar way, showing that they are related (Supplementary Fig. S1, Table 1). For all three peaks, we observed mass differences to the parent ion corresponding to the loss of GlcNAc, GlcNAcMurNAc(r) (r, indicates reduction to N-acetylmuramitol), GlcNAcMurNAc(r)-L-alanine and GlcNAcMurNAc(r)-L-alanine-D-glutamate, confirming that products 1–3 are all muropeptides. The neutral masses of the Protochlamydia products 1, 2 and 3 were all 314.12 Da larger than the masses of the reduced monomeric muropeptides (with tri-, tetra- or pentapeptide) from Gram-negative bacteria34, however, suggesting the presence of a common modification in the Protochlamydia muropeptides. The neutral mass of product 4 was consistent with a peptide crosslinked dimer of product 2. Additional mass differences that occurred in all fragmentation spectra indicated the presence of the same and as yet unknown modifications with 129 and 203 Da, respectively, explaining the higher mass of Protochlamydia muropeptides 1–3 compared to the monomeric muropeptides of Escherichia coli.
To confirm that PG was a major component of Protochlamydia sacculi and check for the presence of disulphide-crosslinked protein components, purified sacculi were subjected to lysozyme and dithiothreitol treatment, respectively, and imaged with negative stain EM. Only the incubation with lysozyme degraded the sacculi (Supplementary Fig. S2).
Fluorescence imaging of D-alanine incorporation in vivo
To further confirm the presence of PG in Protochlamydia sacculi, we tested whether fluorescently labelled amino acids (fluorescent D-amino acids (FDAA); labelled D-alanine in our experiments)35 would be incorporated into chlamydial cells in vivo. Incubation of amoeba cultures continuously infected with Protochlamydia (including reticulate bodies, elementary bodies and transitional stages) with FDAA (HADA (hydroxy coumarin-carbonyl-amino-D-alanine) and BADA (4,4-difluoro-5,7-dimethyl-4-bora-3a,4a-diaza-s-indacene-3-propionic acid-3-amino-D-alanine)) resulted in multiple strong and chlamydial cell-sized signals inside amoeba cells (Fig. 3a–d, Supplementary Fig. S3). In many cases, the FDAA labelling in infected amoebae overlapped with staining of the chlamydial cells by 4′,6-diamidino-2-phenylindole (DAPI) or chlamydiae-specific fluorescence in situ hybridization (FISH). Not all cells stained by FISH/DAPI showed a corresponding FDAA signal, at least in part because chlamydial cells were in different developmental stages, including non-replicating elementary bodies. No signals were detected when uninfected amoebae were incubated with FDAA (Supplementary Fig. S3D, E), or when infected amoebae were incubated with dimethylsulphoxide only. Interestingly, purified Protochlamydia elementary bodies (which cannot undergo cell division and are therefore probably not actively synthesizing new PG) also did not show labelling upon incubation with FDAA (Supplementary Fig. S3B,C), indicating that PG synthesis takes place during Protochlamydia replication inside the host. Amoeba cultures infected with Simkania, on the other hand, showed either no signals (using BADA) or signals similar to the background level (using HADA) upon labelling with FDAA (Supplementary Fig. S4A,B), consistent with the absence of purifiable sacculi. Purified Simkania cells were not labelled by either dye (Supplementary Fig. S4C,D).
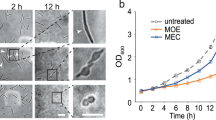
Protochlamydia-infected amoebae (a–d), but not purified Protochlamydia cells (Supplementary Fig. S2) or uninfected amoeba cells (Supplementary Fig. S2), stained positively for new PG synthesis with fluorescently labelled D-alanine dyes HADA (a) and BADA (Supplementary Fig. S2), confirming the synthesis of PG in actively growing Protochlamydia. Specific FISH staining of chlamydial cells (b, overlay with a shown in c) and the eukaryotic host (d, shows overlay with c) are shown. The treatment of Protochlamydia-infected amoeba cultures (amoeba cell outlines in white) with cell-wall-synthesis-targeting fosfomycin (h, control shown in g) resulted in a dramatic decrease in infection rate and aberrant or larger chlamydial cell shapes (shown are immunofluorescent stainings of chlamydial outer membrane proteins), suggesting a crucial role of PG in the Protochlamydia life cycle. Fosfomycin did not affect Simkania (f, control shown in e). Scale bars, 5 μm in d, 10 μm in e–h.
Protochlamydia sensitivity to cell-wall-targeting antibiotics
Owing to the high conservation of PG throughout the bacterial domain of life, many antibacterial drugs target PG synthesis. The so-called chlamydial anomaly16 is that despite the fact that PG has not been detected in pathogenic chlamydiae, these organisms are sensitive to cell-wall-targeting β-lactam antibiotics. Penicillin, for instance, leads to the formation of enlarged aberrant cells36,37 and blocks the conversion between developmental stages38. Environmental chlamydiae, in contrast, are resistant to β-lactams39,40 –possibly due to putative β-lactamases encoded in their genomes. To explore the role of the PG sacculus in the Protochlamydia life cycle, we used an alternative PG synthesis-targeting antibiotic (fosfomycin) to treat infected amoeba cells. The addition of 500 μg ml−1 fosfomycin to Protochlamydia-infected amoeba cultures led to a significant decrease in infection rate (20.2%±8 infected amoebae for fosfomycin-treated cultures versus 95.8%±2.2 infected amoebae for untreated cultures; P<0.0001, unpaired t-test). Protochlamydia cells within treated cultures were also up to eight-times larger than normal (diameters of up to 6 μm) (Fig. 3g,h). Lower fosfomycin concentrations (25 μg ml−1 and 100 μg ml−1) induced the formation of fewer aberrant forms and did not affect the infection rate (not shown). Fosfomycin treatment of Simkania-infected amoeba cultures led to only a slight decrease in infection rate (59.9%±5.6 infected amoebae for fosfomycin-treated cultures versus 67%±1.7 infected amoebae for untreated cultures) and no differences in cell size were detected (Fig. 3e,f).
Discussion
We conclude that Protochlamydia synthesize sacculi containing PG that can be hydrolysed by cellosyl, contains monomeric and crosslinked muropeptides, and carries yet unknown modifications at virtually every subunit. No evidence of PG in Simkania was found. Fluorescence imaging of D-Ala incorporation in vivo and monitoring of cell wall antibiotic sensitivity further suggested that the Protochlamydia PG sacculus has an important role in cell cycle and shape. This challenges previous speculations that chlamydiae synthesize a small ring of PG only during cell division41. Because this might still be true for Simkania and pathogenic chlamydiae, however, our data prompts a reconsideration of whether these organisms lack PG entirely (and the effects of β-lactams are pleiotropic) or if they synthesize novel PG structures that are not purified by standard sacculus preparation protocols.
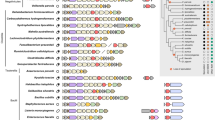
The presence of sacculi in Protochlamydia but not in Simkania matches the less complete set of synthetic genes in the latter: Simkania, as well as pathogenic chlamydiae, lack an undecaprenyl-diphosphate phosphatase (UppP) and alanine/glutamine racemases (Alr, MurI) (Supplementary Table S1)13,42,43. Interestingly, transglycosylases have not been found in any chlamydial genomes (Supplementary Table S1)13 or in the genomes of a few other PG-possessing bacteria44,45, so some other enzyme(s) must be capable of synthesizing glycan strands.
The presence of PG sacculi in Protochlamydia and in the Chlamydiae’s sister phylum Verrucomicrobia46, together with the fact that the more basal chlamydial lineages have more complete PG synthesis pathways make it likely that the last common chlamydial ancestor synthesized a PG sacculus. The detection of a PG-containing sacculus in Protochlamydia challenges the view that FtsZ is essential in PG-possessing bacteria4,6,7, however, because to our knowledge, Protochlamydia is the first example of a bacterium with a PG cell wall, but without FtsZ. Studying cell division and septal development in this organism could help clarify the role of FtsZ and the evolutionary transition to PG- and FtsZ-independency.
Methods
Cultivation of organisms
Acanthamoeba castellanii Neff infected with P. amoebophila UWE25, or A. castellanii UWC1 infected with S. negevensis, were cultivated in TSY medium (30 g l−1 trypticase soy broth, 10 g l−1 yeast extract, pH 7.3) at 20 °C. Amoebal growth was monitored by light microscopy and medium was exchanged every 3–6 days. The presence and identity of the chlamydial symbionts was verified by isolation of DNA from cultures followed by amplification and sequencing of the 16S rRNA genes. In addition, FISH using specific probes combined with DAPI staining of infected cultures was performed using specific probes for the respective symbiont47. Amoebae infected with chlamydiae were allowed to attach on slides and were fixed with 4% formaldehyde at 20 °C. Cells were hybridized for 1.5 h at 46 °C at a formamide concentration of 25% with the Protochlamydia-specific probe E25-454 (5′-GGA TGT TAG CCA GCT CAT-3′) or the Simkania-specific probe Simneg-183 (5′-CAG GCT ACC CCA GCT CTC-3′) and the probe EUB338 (ref. 48). Subsequently, cells were stained with DAPI (0.5 μg ml−1 in PBS) for 5 min, and slides were analysed using an epifluorescence microscope.
Purification of chlamydiae
Infected A. castellanii cultures were harvested by centrifugation (7,197 × g, 10 min), washed in Page’s Amoebic Saline (PAS)49, centrifuged and resuspended in PAS. Amoeba cells were disrupted by vortexing with an equal volume of glass beads for 3 min. Glass beads and cell debris were removed by centrifugation (5 min, 300 × g). The supernatant was filtered through a 1.2 μm filter and centrifuged at maximum speed for 10 min, 18,000 × g. The obtained pellet was resuspended in PAS.
Plunge-freezing
For plunge-freezing, copper/rhodium EM grids (R2/2 or R2/1, Quantifoil) were glow-discharged for 1 min. A 20 × -concentrated bovine serum albumin-treated solution of 10 nm colloidal gold (Sigma) was added to purified chlamydiae or sacculi (1:4 v/v) immediately before plunge freezing. A 4-μl droplet of the mixture was applied to the EM grid, then automatically blotted and plunge-frozen into a liquid ethane–propane mixture50 using a Vitrobot (FEI Company)51.
Electron cryotomography
Images were collected using a Polara 300 kV FEG transmission electron microscope (FEI Company) equipped with an energy filter (slit width 20 eV; Gatan) on a lens-coupled 4 k × 4 k UltraCam CCD (Gatan). Pixels on the CCD represented 0.95 nm (22,500 × ) or 0.63 nm (34,000 × ) at the specimen level. Typically, tilt series were recorded from −60° to +60° with an increment of 1° at 10 μm under-focus. The cumulative dose of a tilt-series was 180–220 e−/Å2. UCSF Tomo52 was used for automatic acquisition of tilt-series and 2D projection images. Three-dimensional reconstructions were calculated using the IMOD software package53 or Raptor54. Tomograms were visualized using 3dMOD53. Density profiles were generated using ImageJ.
Fluorescent labelling of PG
Newly synthesized PG was labelled using FDAA35. A. castellanii cells continuously infected with Protochlamydia or Simkania were harvested and resuspended in a mixture of TSY and PAS (1:1). Cells were incubated with 1.5 mM HADA35 or BADA for 6 h with gentle shaking. Cells were pelleted, washed three times and fixed with 4% formaldehyde followed by FISH using the chlamydia-specific probe Chls-0523 or DAPI-staining. As a control, uninfected amoebae and purified chlamydiae were treated in the same way.
Antibiotic treatment of infected amoeba cultures
A. castellanii were seeded into the wells of a multi-well dish and infected with purified Simkania and Protochlamydia elementary bodies55. After centrifugation at 600 × g for 10 min, the medium was exchanged for TSY supplemented with fosfomycin (0, 25, 100 or 500 μg ml−1, respectively). Medium was exchanged once at 48 h post infection. Cells were fixed with methanol at 96 h post infection followed by immunofluorescence analysis using either anti-PomS antibodies55 or anti-Simkania antibodies raised against purified chlamydiae. The number of infected amoebae was counted for each treatment.
Preparation and composition analysis of sacculi
Chlamydial cells were purified from 1.8 l (Protochlamydia) and 3 l (Simkania) of infected amoeba culture and, depending on the amount of harvested cells, resuspended in 2.6–5.2 ml 4% SDS (w/v). After shipping (overnight, room temperature), the suspensions were dripped into 4% SDS (preheated to 90 °C, 6 ml final volume) and stirred for 2.5 h at 90 °C. Sacculi were pelleted (30 min, 135,000 × g) in a TLA-100.3 rotor (Beckman Coulter), washed 4 × in 3 ml water, resuspended in 150 μl water and supplemented with 0.02% (w/v) sodium azide. Muropeptides were released from the PG by an overnight incubation with the muramidase Cellosyl (Hoechst, Frankfurt am Main, Germany) on a thermal shaker at 37 °C and 800 rpm. The sample was boiled for 10 min and centrifuged for 10 min at 13,500 × g. The muropeptides present in the supernatant were reduced with sodium borohydride and separated on a 250 × 4.6 mm 3-μm Prontosil 120-3-C18 AQ reversed phase column (Bischoff, Germany) at 55 °C using a 135 min gradient from 50 mM sodium phosphate pH 4.31 to 75 mM sodium phosphate pH 4.95, 15% methanol and a flow rate of 0.5 ml min−1 (ref. 32). Muropeptides were detected at 205 nm. Fractions were collected, concentrated in a SpeedVac, acidified by 1% trifluoroacetic acid and analysed by offline electrospray MS on a Finnigan LTQ-FT mass spectrometer (ThermoElectron, Bremen, Germany) in positive ion mode using mass scans over the mass range from m/z=300 to m/z=1,900 at a typical spray voltage of 1.1–1.5 kV34. Parent ion scans were acquired with an FT MS resolution setting of 100,000 (at m/z=400) with a typical mass accuracy of 3 ppm. MS/MS scans were performed in the ion trap, which has a typical mass accuracy for the fragment ion of±0.3 Da. MS spectra were deconvoluted to generate uncharged masses using the QualBrowser program (ThermoElectron, Bremen, Germany)34.
Negative-stain EM of treated sacculi preparations
Purified sacculi were incubated at 37 °C for 12 h in 20 μl total volume with either lysozyme (10 mg ml−1), dithiothreitol (5 mM) or phosphate buffered saline. Samples were applied to a Formvar-coated, carbon-coated, glow-discharged copper EM grid (Electron Microscopy Sciences). Samples were aspirated, stained with 1.5% uranylacetate and imaged on an FEI Tecnai T12 electron microscope.
Additional information
How to cite this article: Pilhofer, M. et al. Discovery of chlamydial peptidoglycan reveals bacteria with murein sacculi but without FtsZ. Nat. Commun. 4:2856 doi: 10.1038/ncomms3856 (2013).
Change history
03 December 2013
The HTML version of this Article previously published did not acknowledge Grant J. Jensen as a corresponding author. This has now been corrected in the HTML; the PDF version of the paper was correct from the time of publication.
References
Wagner, M. & Horn, M. The Planctomycetes, Verrucomicrobia, Chlamydiae and sister phyla comprise a superphylum with biotechnological and medical relevance. Curr. Opin. Biotechnol. 17, 241–249 (2006).
Devos, D. P. & Reynaud, E. G. Evolution. Intermediate steps. Science 330, 1187–1188 (2010).
Reynaud, E. G. & Devos, D. P. Transitional forms between the three domains of life and evolutionary implications. Proc. Biol. Sci. 278, 3321–3328 (2011).
Erickson, H. P., Anderson, D. E. & Osawa, M. FtsZ in bacterial cytokinesis: cytoskeleton and force generator all in one. Microbiol. Mol. Biol. Rev. 74, 504–528 (2010).
Pilhofer, M. et al. Characterization and evolution of cell division and cell wall synthesis genes in the bacterial phyla Verrucomicrobia, Lentisphaerae, Chlamydiae, and Planctomycetes and phylogenetic comparison with rRNA genes. J. Bacteriol. 190, 3192–3202 (2008).
Lluch-Senar, M., Querol, E. & Pinol, J. Cell division in a minimal bacterium in the absence of ftsZ. Mol. Microbiol. 78, 278–289 (2010).
Leaver, M., Dominguez-Cuevas, P., Coxhead, J. M., Daniel, R. A. & Errington, J. Life without a wall or division machine in Bacillus subtilis. Nature 457, 849–853 (2009).
Manire, G. P. & Tamura, A. Preparation and chemical composition of the cell walls of mature infectious dense forms of meningopneumonitis organisms. J. Bacteriol. 94, 1178–1183 (1967).
Tamura, A. & Manire, G. P. Preparation and chemical composition of the cell membranes of developmental reticulate forms of meningopneumonitis organisms. J. Bacteriol. 94, 1184–1188 (1967).
Caldwell, H. D., Kromhout, J. & Schachter, J. Purification and partial characterization of the major outer membrane protein of Chlamydia trachomatis. Infect. Immun. 31, 1161–1176 (1981).
Fox, A. et al. Muramic acid is not detectable in Chlamydia psittaci or Chlamydia trachomatis by gas chromatography-mass spectrometry. Infect. Immun. 58, 835–837 (1990).
Barbour, A. G., Amano, K., Hackstadt, T., Perry, L. & Caldwell, H. D. Chlamydia trachomatis has penicillin-binding proteins but not detectable muramic acid. J. Bacteriol. 151, 420–428 (1982).
McCoy, A. J. & Maurelli, A. T. Building the invisible wall: updating the chlamydial peptidoglycan anomaly. Trends. Microbiol. 14, 70–77 (2006).
Perkins, H. R. & Allison, A. C. Cell-wall constituents of Rickettsiae and Psittacosis-Lymphogranuloma organisms. J. Gen. Microbiol. 30, 469–480 (1963).
Garrett, A. J., Harrison, M. J. & Manire, G. P. A search for the bacterial mucopeptide component, muramic acid, in Chlamydia. J. Gen. Microbiol. 80, 315–318 (1974).
Moulder, J. W. Why is Chlamydia sensitive to penicillin in the absence of peptidoglycan? Infect. Agents. Dis. 2, 87–99 (1993).
Matsumoto, A. Recent progress of electron microscopy in microbiology and its development in future: from a study of the obligate intracellular parasites, chlamydia organisms. J. Electron. Microsc. 28, 57–64 (1979).
Tamura, A., Matsumoto, A., Manire, G. P. & Higashi, N. Electron microscopic observations on the structure of the envelopes of mature elementary bodies and developmental reticulate forms of Chlamydia psittaci. J. Bacteriol. 105, 355–360 (1971).
Huang, Z. et al. Cryo-electron tomography of Chlamydia trachomatis gives a clue to the mechanism of outer membrane changes. J. Electron. Microsc. 59, 237–241 (2010).
McCoy, A. J. & Maurelli, A. T. Characterization of Chlamydia MurC-Ddl, a fusion protein exhibiting D-alanyl-D-alanine ligase activity involved in peptidoglycan synthesis and D-cycloserine sensitivity. Mol. Microbiol. 57, 41–52 (2005).
McCoy, A. J., Sandlin, R. C. & Maurelli, A. T. In vitro and in vivo functional activity of Chlamydia MurA, a UDP-N-acetylglucosamine enolpyruvyl transferase involved in peptidoglycan synthesis and fosfomycin resistance. J. Bacteriol. 185, 1218–1228 (2003).
Hesse, L. et al. Functional and biochemical analysis of Chlamydia trachomatis MurC, an enzyme displaying UDP-N-acetylmuramate:amino acid ligase activity. J. Bacteriol. 185, 6507–6512 (2003).
Patin, D., Bostock, J., Blanot, D., Mengin-Lecreulx, D. & Chopra, I. Functional and biochemical analysis of the Chlamydia trachomatis ligase MurE. J. Bacteriol. 191, 7430–7435 (2009).
Patin, D., Bostock, J., Chopra, I., Mengin-Lecreulx, D. & Blanot, D. Biochemical characterisation of the chlamydial MurF ligase, and possible sequence of the chlamydial peptidoglycan pentapeptide stem. Arch. Microbiol. 194, 505–512 (2012).
Henrichfreise, B. et al. Functional conservation of the lipid II biosynthesis pathway in the cell wall-less bacteria Chlamydia and Wolbachia: why is lipid II needed? Mol. Microbiol. 73, 913–923 (2009).
Horn, M. Chlamydiae as symbionts in eukaryotes. Annu. Rev. Microbiol. 62, 113–131 (2008).
Leforestier, A., Lemercier, N. & Livolant, F. Contribution of cryoelectron microscopy of vitreous sections to the understanding of biological membrane structure. Proc. Natl Acad. Sci. USA 109, 8959–8964 (2012).
Tocheva, E. I., Li, Z. & Jensen, G. J. Electron cryotomography. Cold Spring Harb. Perspect. Biol. 2, a003442 (2010).
Liu, J. et al. Cellular architecture of Treponema pallidum: novel flagellum, periplasmic cone, and cell envelope as revealed by cryo electron tomography. J. Mol. Biol. 403, 546–561 (2010).
Tocheva, E. I. et al. Peptidoglycan remodeling and conversion of an inner membrane into an outer membrane during sporulation. Cell 146, 799–812 (2011).
Hatch, T. Disulfide cross-linked envelope proteins: the functional equivalent of peptidoglycan in chlamydiae? J. Bacteriol. 178, 1–5 (1996).
Glauner, B. Separation and quantification of muropeptides with high-performance liquid chromatography. Anal. Biochem. 172, 451–464 (1988).
de Jonge, B. L. & Tomasz, A. Abnormal peptidoglycan produced in a methicillin-resistant strain of Staphylococcus aureus grown in the presence of methicillin: functional role for penicillin-binding protein 2A in cell wall synthesis. Antimicrob. Agents. Chemother. 37, 342–346 (1993).
Bui, N. K. et al. The peptidoglycan sacculus of Myxococcus xanthus has unusual structural features and is degraded during glycerol-induced myxospore development. J. Bacteriol. 191, 494–505 (2009).
Kuru, E. et al. In Situ probing of newly synthesized peptidoglycan in live bacteria with fluorescent D-amino acids. Angew. Chem. 51, 12519–12523 (2012).
Matsumoto, A. & Manire, G. P. Electron microscopic observations on the effects of penicillin on the morphology of Chlamydia psittaci. J. Bacteriol. 101, 278–285 (1970).
Skilton, R. J. et al. Penicillin induced persistence in Chlamydia trachomatis: high quality time lapse video analysis of the developmental cycle. PLoS One 4, e7723 (2009).
Beatty, W. L., Morrison, R. P. & Byrne, G. I. Persistent chlamydiae: from cell culture to a paradigm for chlamydial pathogenesis. Microbiol. Rev. 58, 686–699 (1994).
Maurin, M., Bryskier, A. & Raoult, D. Antibiotic susceptibilities of Parachlamydia acanthamoeba in amoebae. Antimicrob. Agents Chemother. 46, 3065–3067 (2002).
Kahane, S., Gonen, R., Sayada, C., Elion, J. & Friedman, M. G. Description and partial characterization of a new chlamydia-like microorganism. FEMS. Microbiol. Lett. 109, 329–334 (1993).
Brown, W. J. & Rockey, D. D. Identification of an antigen localized to an apparent septum within dividing chlamydiae. Infect. Immun. 68, 708–715 (2000).
Horn, M. et al. Illuminating the evolutionary history of chlamydiae. Science 304, 728–730 (2004).
Collingro, A. et al. Unity in variety—the Pan-genome of the Chlamydiae. Mol. Biol. Evol. 28, 3253–3270 (2011).
McPherson, D. C. & Popham, D. L. Peptidoglycan synthesis in the absence of Class A penicillin-binding proteins in Bacillus subtilis. J. Bacteriol. 185, 1423–1431 (2003).
Arbeloa, A. et al. Role of class A penicillin-binding proteins in PBP5-mediated β-lactam resistance in Enterococcus faecalis. J. Bacteriol. 186, 1221–1228 (2004).
Hedlund, B. P., Gosink, J. J. & Staley, J. T. Phylogeny of Prosthecobacter, the fusiform caulobacters: members of a recently discovered division of the Bacteria. Int. J. Syst. Bacteriol. 46, 960–966 (1996).
Schmitz-Esser, S. et al. Diversity of bacterial endosymbionts of environmental Acanthamoeba isolates. Appl. Environ. Microbiol. 74, 5822–5831 (2008).
Amann, R. I. et al. Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl. Environ. Microbiol. 56, 1919–1925 (1990).
Page, F. C. An Illustrated Key to Freshwater and Soil Amoebae Freshwater Biological Association (1976).
Tivol, W. F., Briegel, A. & Jensen, G. J. An improved cryogen for plunge freezing. Microsc. Microanal. 14, 375–379 (2008).
Iancu, C. V. et al. Electron cryotomography sample preparation using the Vitrobot. Nat. Protocols 1, 2813–2819 (2006).
Zheng, S. Q. et al. UCSF tomography: an integrated software suite for real-time electron microscopic tomographic data collection, alignment, and reconstruction. J. Struct. Biol. 157, 138–147 (2007).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 1996/01/01, 71–76 (1996).
Amat, F. et al. Markov random field based automatic image alignment for electron tomography. J. Struct. Biol. 161, 260–275 (2008).
Aistleitner, K. et al. Identification and characterization of a novel porin family highlights a major difference in the outer membrane of chlamydial symbionts and pathogens. PLoS One 8, e55010 (2013).
Frirdich, E. et al. Peptidoglycan-modifying enzyme Pgp1 is required for helical cell shape and pathogenicity traits in Campylobacter jejuni. PLoS. Pathog. 8, e1002602 (2012).
Acknowledgements
This work was funded by the Austrian Science Fund FWF (Y277-B03 to M.H.), the European Research Council (ERC StG ‘EvoChlamy’ to M.H.), NIH grant GM094800B (to G.J.J.), the Caltech Center for Environmental Microbiology Interactions (to G.J.J., M.P.), a gift from the Gordon and Betty Moore Foundation to Caltech, the BBSRC (BB/I020012/1 to W.V.), NIH grant AI059327 (to M.S.V.) and NIH grant GM051986 (to Y.V.B.). We thank Elitza Tocheva for discussions on the preparation of sacculi.
Author information
Authors and Affiliations
Contributions
M.P. initiated the study. M.P. and K.A. performed all experiments except HPLC/MS analyses of sacculi, which were done by J.B., J.G. and W.V. E.K., E.H., Y.V.B. and M.S.V. provided the FDAA dyes and advice on the FDAA labelling experiments. M.P., K.A., W.V., M.H. and G.J.J. designed the experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1-S4 and Supplementary Table S1 (PDF 3948 kb)
Rights and permissions
About this article
Cite this article
Pilhofer, M., Aistleitner, K., Biboy, J. et al. Discovery of chlamydial peptidoglycan reveals bacteria with murein sacculi but without FtsZ. Nat Commun 4, 2856 (2013). https://doi.org/10.1038/ncomms3856
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms3856
This article is cited by
-
Using click chemistry to study microbial ecology and evolution
ISME Communications (2023)
-
Effects of DNA preservation solution and DNA extraction methods on microbial community profiling of soil
Folia Microbiologica (2021)
-
Non-essentiality of canonical cell division genes in the planctomycete Planctopirus limnophila
Scientific Reports (2020)
-
Maioricimonas rarisocia gen. nov., sp. nov., a novel planctomycete isolated from marine sediments close to Mallorca Island
Antonie van Leeuwenhoek (2020)
-
Cell morphology and nucleoid dynamics in dividing Deinococcus radiodurans
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.