Abstract
Cambodia harbours a variety of aboriginal (and presumably ancient) populations that have largely been ignored in studies of genetic diversity. Here we investigate the matrilineal gene pool of 1,054 Cambodians from 14 geographic populations. Using mitochondrial whole-genome sequencing, we identify eight new mitochondrial DNA haplogroups, all of which are either newly defined basal haplogroups or basal sub-branches. Most of the new basal haplogroups have very old coalescence ages, ranging from ~55,000 to ~68,000 years, suggesting that present-day Cambodian aborigines still carry ancient genetic polymorphisms in their maternal lineages, and most of the common Cambodian haplogroups probably originated locally before expanding to the surrounding areas during prehistory. Moreover, we observe a relatively close relationship between Cambodians and populations from the Indian subcontinent, supporting the earliest costal route of migration of modern humans from Africa into mainland Southeast Asia by way of the Indian subcontinent some 60,000 years ago.
Similar content being viewed by others
Introduction
Bordered by Thailand, Laos and Vietnam on the southern coast of the Indochina Peninsula, the Kingdom of Cambodia has a population of about 13.4 million (Cambodia National Census, 2008), 96% of which belongs to Khmer, a result of the historic expansion of the Khmer Empire in the 12th century1. Alongside the Khmer, there are 20 minority ethnic groups that reside primarily in the northeastern provinces, colloquially known in Cambodia as aborigines. Linguistically, Austro-Asiatic languages are the most common in Cambodia, being spoken by the Khmer and nearly all the aborigines. Being one of the most ancient language families in eastern Asia, Austro-Asiatic is also spoken in India, Bangladesh and southwestern China, implying that the Austro-Asiatic speaking populations may represent the descendants of the earliest settlers of modern humans who migrated from Africa and entered into eastern Asia about 60,000 years ago (ya)2,3,4.
Previous genetic studies of the mitochondrial DNA (mtDNA) and the Y-chromosome diversity of Asian populations suggested that modern humans of ultimate African origin initially entered into the southern part of eastern Asia around 60,000 ya and then migrated northward to mainland East Asia about 40,000 ya2,3,5,6,7. This southern origin and northward migration were confirmed by a recent analysis of genome-wide sequence variation, in which the Austro-Asiatic speaking populations were found at a basal position in the phylogenetic tree illustrating the genetic relationships among Asian populations.8 Collectively, currently available genetic data places mainland Southeast Asia (MSEA) and southern China as the potential cradle of modern humans for the initial peopling of eastern Asia. Consequently, revealing the pattern of ancient genetic diversity among early modern human populations is not only informative simply for tracing the prehistoric migrations but also important for understanding the underlying molecular mechanisms of adaptation to the varied environments early human settlers faced along their migratory paths.
Though many genetic studies have been conducted in populations from Southeast Asia (for example, populations from Thailand, Laos, Vietnam, Malaysia, Indonesia and other Island of Southeast Asian (ISEA) countries)9,10,11,12,13,14,15,16,17,18, there have been only a few studies with limited samples from Cambodia. In previous study, we found a relatively higher genetic diversity among Cambodians (in a relatively small sample comprising 26 male individuals) when compared with the surrounding populations2,3,4,5,19, an implication of inherited ancient genetic diversity among the Cambodian aborigines. In another study, however, no novel lineages were observed in the scans of mtDNA (31 samples) and Y-chromosome markers (125 samples)20, though this was most probably due to a limited sample size and insufficient coverage of aboriginal populations from Cambodia.
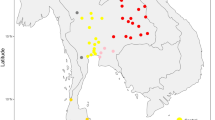
In this study, to elucidate the genetic background of Cambodian aborigines, we collected a total of 1,054 unrelated samples, representing 13 aboriginal ethnic populations and one Khmer population. These populations are from three provinces in northeastern Cambodia, where nearly 95% of all the aborigines live (fig. 1). We analyse sequence variation within the mtDNA control region (HVS-I and HVS-II) as well as parts of the coding region in all the samples. In addition, we sequence the entire mitochondrial genomes of 98 selected samples. These Cambodian aboriginal mitogenomes reveal four novel basal lineages and a further four novel sub-branches, indicating that present-day Cambodian aborigines still retain ancient genetic polymorphisms in their maternal lineages.
Results
Classification of the Cambodian mtDNA haplogroups
Through control region and partial coding region as well as whole-genome sequencing, a total of 1,000 out of 1,054 (94.88%) Cambodian individuals (Fig. 2) were classified into 69 known mtDNA haplogroups/subhaplogroups (Supplementary Data 1), previously identified in Southeast Asian and East Asian populations ( http://www.phylotree.org, mtDNA tree build 15 (30 September 2012))21. The remaining 54 individuals belong to eight novel mtDNA haplogroups newly identified in this study. Overall, the dominant haplogroups observed in the Cambodian populations were B5 (28.08%), F1 (18.22%), M12b (8.25%), R22 (5.79%) and B4 (5.60%), which altogether account for 65.94% (695/1,054) of all samples (Fig. 2).
The branches in red colour are the newly defined haplogroups in this study. The branches in green colour are the re-defined haplogroups due to the adding of the Cambodian sequences. The mutations used to define the mtDNA lineages are given along the branches. For transversions, the derived alleles are shown in lowercase. Back-mutations are indicated with a ‘!’ suffix. Population abbreviations: Br, Brao; Ja, Jarai; Ka, Kachac; KH, Khmer; Kr, Kravet; Kre, Kreung; Ku, Kuy; La, Lao; Lu, Lun; Me, Mel; PH, Phnong; St, Stieng; To, Tompuon; and Kra, Kraol.
We compared the haplogroup profiles between the two non-Austro-Asiatic (Lao and Jarai) and the other Austro-Asiatic populations (Table 1), and no clustering by language families was observed (Fig. 3), implying that there has been extensive genetic exchange among regional populations in Cambodia despite the fact that they belong to different language families. Therefore, these two non-Austro-Asiatic populations were not treated separately in the following analyses. The Kraol population was omitted in the following analyses considering its small size (n=2).
Novel mtDNA basal lineages and sub-branches in Cambodians
Using mitochondrial whole-genome sequencing, we identified a total of eight novel lineages that have not been reported or classified previously. Phylogenetic analysis indicated that four of these are novel basal lineages within macrohaplogroup M (designated as M59, M69 and M78) and macrohaplogroup N (designated as N7). Three of the four newly defined lineages (M59, M69 and M78) also included previously reported but unclassified sequences from other Asian populations (Fig. 4). The other four lineages are novel sub-branches of the known M/N haplogroups (designated as M68 (which shares a root with M62), M68a, M3d and M3d1) with all of them also sitting at basal positions within the respective haplogroups. Figure 4 illustrates the phylogenetic positions of the eight newly identified haplogroups in Cambodians. Although they only account for a small portion (5.12%) of the Cambodian mtDNA pool, these presumably ancient haplogroups suggest a high genetic diversity due to antiquity of the Cambodian maternal lineages.
The tree includes 98 newly sequenced Cambodians genomes and 35 previously reported genomes from other Asian populations (the sample IDs are shown in grey colour). The branches in red colour are the novel basal haplogroups and sub-branches, and the branches in green colour are the re-defined haplogroups due to the adding of the Cambodian sequences. The mutations used to define the mtDNA lineages are given along the branches. For transversions, the derived alleles are shown in lowercase. Back-mutations are indicated with a ‘!’ suffix. Synonymous mutations in the coding region are underlined. The numbers in the brackets are sample size of the haplotypes.
Among the four newly identified basal lineages, haplogroup M78 is defined by a mutation motif of A93G-A4164G-A15652G-C16287T–C16327a, which includes three previously unclassified sequences, two from Myanmar (JX289097 and JX289130, NCBI GenBank nucleotide database) and one from Tibet (HM030537) (ref. 22), which belong to a M78 sub-branch deeply divergent from the four Cambodian samples of the Stieng ethnic group. The haplogroup M69 is defined by a short mutation motif C4392T-T11365C, which is shared by a sequence (HM596653) from Sumatra (Indonesia)23, but with a deep divergence with the twelve sequences from four different Cambodian ethnic groups. The haplogroup M59 is defined by a long motif of A249G-G9380A-T10256C-C11140T-G14040A-T16140C-C16278T, shared by a sequence (JQ702247) from Singapore24 but with a deep divergence from a Cambodian sample of the Tompoun ethnic group. The haplogroup N7 is a completely new basal lineage within macrohaplogoup N, defined by a long motif of A723G-G6570t-T11617C-A13542G-C14668T-C15945T- G16129A, and, interestingly, it is only observed in Cambodians.
A previously identified basal lineage M62 (refs 25, 26, 27) was joined with the novel lineage M68 (defined by G16255A-T16311C) identified in Cambodians, designated as M62'68, and defined by the mutation motif C150T-T4561C-G7664A. Eleven Lao sequences in Cambodians turned out to belong to a sub-branch of haplogroup M3, designated M3d (defined by T11827C-C16344T), to which also four previously published sequences, JF742206 (Nepal)28, FJ770946 (India)29, DQ112779 (Brahui, Pakistan)30 and JF742212 (Nepal)29 belong. The sequences JF742206 and FJ770946, together with the eleven Cambodian Lao samples, form a M3d sub-branch, designated M3d1 (defined by T10238C-T13820C). In addition, another Cambodian sequence (Lao09) refined the motif of haplogroup M46 (now defined by T146C-C3588T) and showed a deep divergence with a sequence (FJ442939) from Thailand31. Two Cambodian sequences (Khmer02 and Jarai06) refined the mutation motif of haplogroup M24 (now defined by T146C-T195C-G5773A-G13359A-T15601C), to which the two previously reported sequences DQ112783 (Cambodia)30 and JF739543 (Philippines)32 belong. Collectively, the novel basal mtDNA lineages found in Cambodians suggest that current Cambodian aborigines still carry ancient genetic diversity.
Estimated coalescent times of Cambodian mtDNA haplogroups
The estimated coalescence ages of the mtDNA haplogroups are listed in Table 2. Most of the haplogroups are very ancient (>25,000 years), especially the newly identified haplogroups in Cambodians. For example, M78, M69 and M68, three of the newly identified haplogroups, have extremely old coalescence ages (55,188 years, 68,137 years and 82,782 years, respectively), falling into the suggested period of initial peopling of modern humans in eastern Asia2,3. Two other novel basal lineages in Cambodians are younger but still of considerable age (36,388 years for N7 and 27,594 years for M59). In addition, besides the newly identified mtDNA lineages in Cambodians, the other lineages (previously reported in other Asian populations) also have relatively old coalescence ages, most of them being older than 25,000 years, whereas only two of the observed lineages fell into the Neolithic period (<10,000 years) (Table 2). The old coalescence ages of the mtDNA lineages are consistent with the multiple novel basal lineages found in Cambodians, again suggesting that the Cambodian aborigines are ancient populations. We also used the Bayesian method in BEAST33 and the maximum likelihood method in PAML34 for age estimation, and the estimated ages by different methods are consistent (Table 3).
Phylogeographic patterns of the dominant haplogroups
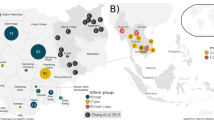
To delve into the phylogeographic and migratory patterns of the major haplogroups in Cambodians, we constructed networks (Fig. 5) and frequency contour maps (Fig. 6) of the haplogroups by combining all available data in Asian populations. B5a was the most dominant haplogroup (28.08%) in Cambodians (Fig. 2 and Table 2) and was also prevalent in southern China (Yunnan, Guangxi, Guangdong and Hainan provinces), northern Laos and southern Vietnam, but rare in northern Asian populations (Supplementary Data 2). The reduced median network of haplogroup B5a (Fig. 5a) indicates that the core haplotype of B5a is mostly shared among the southern populations in eastern Asia, suggesting a southern origin of this mtDNA lineage. The contour map of B5a indicates that the origin of B5a probably lies in MSEA (probably Cambodia), as reflected by the diversified haplotypes (Fig. 5a) and the high level of haplotype diversity (0.8364) of the haplogroup B5a HVS-I sequences in Cambodians (Table 2), which is comparable to the diversity levels of B5a in Vietnam (0.8824) and Thailand (0.8444). The estimated coalescence age of B5a in Cambodians is 34,220 years, which is much older than its coalescence age in other populations (for example, 16,200 years in ISEA11 and 26,000 years in Japan31), and close to the proposed time of the initial northward migration of modern humans in eastern Asia2,3,5. In addition, the star-like network of B5a suggests a relatively recent expansion of this lineage.
F1 is another prevalent haplogroup in Cambodians (18.22%), of which the most common sub-haplogroup is F1a (Fig. 2). The reduced median network of F1a showed that there are two major clusters (F1a* and F1a1a) (Fig. 5d), both of which have star-like shapes, suggesting recent expansions, consistent with the pattern seen in B5a (Fig. 5a). A previous study reported that the sister branches of F1a, that is, F1b and F1c, are largely restricted to southern China, and F1 and F1a were suggested to have a possible origin in this region11. Also, due to the high diversity of root types in Indochina, F1a1a was suggested to have expanded from MSEA to ISEA during the Holocene11. F1a1a is a prevalent haplogroup in Southeast Asia; besides Cambodia, it is also common in Thailand and in aboriginal Senoi groups of the Malay Peninsula12, and the coalescence age of F1a1a was estimated to be 9,000 years in ISEA11. F1a (including F1a1a) has a peak frequency (48%) in the Stieng ethnic group of Cambodia. Both F1a* and F1a1a in Cambodians have ancient coalescence ages (around 60,000 and 48,000 years, respectively) (Table 2), which are much older than their ages in other MSEA regions. Hence, F1a probably originated in MSEA (with Cambodia as a candidate region) and expanded to ISEA, which is also reflected by the F1a contour map (Fig. 6d).
M12b is a Cambodian-specific haplogroup with moderate prevalence (8.25%) in Cambodians, which was firstly defined by Fucharoen et al.35, with sporadic presence in northern India25, southwestern China (Yunnan province)36,37,38, northern Thailand35,39, Laos9 and ISEA12,40. The reduced median network of M12b (Fig. 5b) suggests that the M12b haplotypes in ISEA (mainly Indonesia) were derived from those in Cambodians, and an eastward migration can be inferred from the contour map of M12b (Fig. 6b). In addition, the phylogenetic structure of M12b derived from sequencing the mitogenomes of 19 representative Cambodian samples clearly supports a Cambodian origin of this mtDNA lineage (Fig. 4).
Haplogroup B4c2, which was first defined by Tanaka et al.31, accounts for 5.03% in Cambodians. It is also prevalent (10.12%) in southern Vietnam and Thailand14 and widely distributed in southern China and other Southeast Asian countries20 (Fig. 5c). The reduced median network of B4c2 indicated that the haplotypes from ISEA are sitting at the root position of the network, and there are two distinctive subclusters separated by a transversion (C16184a) (Fig. 5c), which seems to suggest an ISEA origin of this haplogroup. However, the reported coalescence age of B4c2 in ISEA is around 21,000 years11, which is much younger than the B4c2 age in Cambodians (~45,000 years) (Table 2), thus favouring an MSEA origin.
The haplogroup R22 accounts for 5.79% in Cambodians, which is a relatively young lineage (~19,000 years in Cambodians) (Table 2). The reduced median network of R22 (Fig. 4e) suggests an ISEA origin of this lineage with the root haplotypes mostly observed in ISEA and the Andaman Islands, consistent with a previous dating of ~29,000 years in ISEA11. The contour map of R22 indicates a westward migration from ISEA to MSEA (Fig. 6e).
The haplogroup M74 (4.93%) represents another lineage showing a migration from other regions to Cambodia. It was first reported by Kong et al.22 based on mitochondrial whole-genome sequencing, and was suggested to have a southern China origin, dated to 43,000 years ago. The reduced median network of M74 (Fig. 5f) agrees with this view with most of the core haplotypes being distributed in southern China. The contour map of M74 (Fig. 6f) suggests two possible expansion centres, one in southern China and another one in Cambodia. The coalescence time of M74 in Cambodians was estimated at ~39,000 years, younger than that in southern China (Table 2). Hence, a southern China origin of this haplogroup seems more congruent with the currently available data from Asian populations. There are also other minor haplogroups in Cambodians (R9b, N9a and M71) showing an into-Cambodia migratory pattern12,14,17.
Genetic relationship of Cambodians with nearby populations
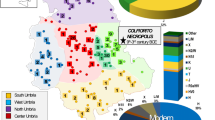
Finally, to examine the genetic relationship of Cambodians with surrounding populations, we conducted a principal component analysis (PCA) based on the mtDNA haplogroup frequencies of Cambodians and 225 different Asian populations, and the result is shown in Fig. 7. The first component (PC1), which explains 18.40% of the genetic variance, separates the northern and southern Asian populations. The second component (PC2, 10.86%) indicates divergence among the southern populations. Intriguingly, the Cambodian populations are clustered with populations from India (Dravidian, Indo-European and Austro-Asiatic speaking populations), Andaman Islands (one Austro-Asiatic speaking population), Australia (aborigines) and Madagascar, but relatively diverged from the other southern populations (Austronesian, Daic and Hmong-Mien speaking populations in MSEA and southern China). In addition, within the Austro-Asiatic language family, Cambodians are closer to the Austro-Asiatic speaking populations from India and the Andaman Islands than to those from MSEA and southern China.
Non-Cambodian populations were grouped by language families. The four Austro-Asiatic populations (red squares) located in the lower left corner are from India (three populations) and the Andaman Islands (one population). The first and the second components explain 18.40 and 10.86% of the genetic variance, respectively.
Inference of demographic changes
To infer demographic changes through time for Cambodian aborigines, we carried out Bayesian skyline plot (BSP)41 using all Cambodian mtDNA HVS-I sequences (Fig. 8). The BSP showed a relatively constant female effective population size in the past 50,000 years, suggesting that the Neolithic agricultural diffusion had a minor impact on the Cambodian aborigines.
The plot was generated using 1,054 Cambodian HVS-I (16,038–16,462) sequences. The MCMC chain was run for 1 × 108 steps, with sampling of parameters every 2,500 steps, and the initial 1 × 107 steps were discarded as burn-in. The purple area reflects the s.d. values of the estimated effective populations sizes.
Discussion
Through an extensive sampling in Cambodian aborigines and high-resolution mtDNA diversity analyses, we showed that Cambodian aboriginal populations still carry ancient sequence polymorphisms in their maternal lineages, suggesting that Cambodia was probably located in the region where the earliest modern human settlers initially populated eastern Asia. We identified eight novel mtDNA lineages in Cambodians, including four basal haplogroups and four sub-branches. This was rather unexpected given the extensive surveys of mtDNA diversity that have already been conducted in many MSEA regions (Thailand, Vietnam, Myanmar and Laos) and southern China6,7,9,11,14,22,35,36,37,38,42,43,44,45.
The dating of the mtDNA haplogroups supports the antiquity of Cambodian populations, with most of the estimated haplogroup ages exceeding 25,000 years. In particular, the ages of the newly defined basal haplogroups M59, M69, M78 and N7 were estimated between ~36,000 and ~68,000 years ago (Table 2), falling within the suggested time period of the earliest settlement of modern humans in eastern Asia3,4. Notably, although most of the newly defined basal haplogroups in Cambodians also exist in other Asian populations, the Cambodian sequences are highly diverged from the non-Cambodian sequences for M59, M69 and M78 (Fig. 4), therefore, it is unlikely that these ancient mtDNA lineages were brought into Cambodia only recently via migration. Until now, no ancient human fossils or cultural relics beyond the Neolithic time have yet been discovered in Cambodia46,47, probably due to the limited excavations conducted by archaeologists and/or the tropical environments in Cambodia, which are not ideal for fossil and relic preservation.
Besides the newly identified mtDNA lineages, many dominant haplogroups in Cambodia, which are shared with populations in MSEA and southern China, probably also originated locally in Cambodia. As reflected by the contour maps of the major haplogroups (Fig. 6), most of the dominant haplogroups in Cambodians (B5a, F1a, M12b and B4c2) have high frequencies and high haplotype diversities in Cambodia as compared with the surrounding regions, an implication of a Cambodian dispersal centre of these haplogroups, northward to mainland East Asia and southward to ISEA. Notably, there are also less-prevalent haplogroups, which seem to have been brought into Cambodia from other regions, including R22, M74 and several other minor haplogoups (R9b, N9a and M71).
The genetic relationships inferred from the PCA map among extended Asian populations are consistent with the proposed antiquity of the Cambodian aborigines. In the PCA map (Fig. 7), Cambodians are clustered with populations from the Indian subcontinent, the Andaman Islands, Australia (aborigines) and Madagascar, congruent with the widely accepted costal migratory route of modern humans, which starting in Africa advanced through the Indian subcontinent and into MSEA around 60,000 years ago3,48. This clustering pattern tends to support a single early migration wave of modern humans from Africa to eastern Asia8,48 though a recent analysis of the genome of an aboriginal Australian suggested multiple dispersals49. In addition, the findings of ancient maternal lineages in Cambodians supports the idea of a MSEA dispersal centre of modern humans in eastern Asia, consistent with the proposed southern origin and early northward migrations of modern humans in mainland East Asia inferred from Y-chromosome data2. Hence, further studies of the aboriginal populations from other Southeast Asian countries, such as Myanmar, may reveal more unidentified ancient mtDNA lineages.
Alternatively, Cambodia also experienced recent influence from India. Cambodia historically had a highly Indianized state known as Funan (first to sixth century) and its successor Chenla (late sixth to the early ninth century), harbouring a culture similar to that in India. Substantial Indian immigration during the first to thirteenth century was suggested to have made a hefty contribution to the modern gene pool of Cambodians50. However, as we argued above, most of the dominant haplogoups in Cambodians showed an out-of-Cambodia expansion pattern, and the influence of the suggested relatively recent gene flows from India does not seem to have had a large impact on the Cambodian aborigines, at least regarding their matrilineal ancestry.
In conclusion, Cambodia harbours ancient and indigenous mtDNA haplogroups, which have accumulated abundant mutations that show patterns of long-term in situ evolution. Many of the prevalent mtDNA haplogroups in ISEA and mainland East Asia probably originated in MSEA (presumably Cambodia). Hence, the Cambodian aborigines are important ethnic populations for reconstructing the genetic makeup of early modern human settlers in Asia.
Methods
Sample collection
We collected 5 ml of blood samples from a total of 1,054 unrelated individuals (693 females and 361 males) from Cambodia. These samples were from 14 geographic populations, including 13 aboriginal populations and 1 Khmer population from three provinces in northeastern Cambodia (Fig. 1). Written informed consents were obtained from all sampled Cambodian individuals, and the study protocol was approved by the Internal Review Board of Kunming Institute of Zoology, Chinese Academy of Sciences. As shown in Table 1, except for Jarai and Lao, which belong to Austronesian and Daic language families, respectively, all the other 12 populations belong to the Mon-Khmer branch of the Austro-Asiatic family.
MtDNA sequencing and genotyping
Following the previously described method45, for all the 1,054 samples, we first sequenced the mtDNA HVS-I (range: 16,038–16,462) and HVS-II (range: 65–417) regions, as well as a coding region (range: 10,220–10,610) containing two sites, 10,398 and 10,400, diagnostic for the macrohaplogroups N and M, respectively. We next sequenced several diagnostic-site-containing coding regions to define the mtDNA haplogroup of each individual (Supplementary Data 1). Based on this strategy, 860 individuals could be assigned to a known mtDNA haplogroup. From the remaining 194 samples of unknown haplogroups, we selected 75 representative samples, together with 4 samples of known haplogroups, which were subjected to whole-mitochondrial genome sequencing. Based on the phylogenetic information gained from the newly sequenced mtDNA genomes, we next sequenced at least one specific coding-region position in the remaining 119 samples to define their haplogroups (Supplementary Data 1). Moreover, to clarify the phylogeny of haplogroup M12b, which is specific to Cambodians, we additionally conducted whole-genome sequencing of 19 samples belonging to this haplogroup. In total, we acquired the mitochondrial whole-genome sequences of 98 Cambodian individuals. The protocol for mitochondrial whole-genome sequencing was adopted from a published study51 involving amplification of two overlapping fragments each of 8.5 kb in length. For the newly discovered mtDNA haplogroups, the nomenclature follows the system suggested by van Oven26.
We also collected previously published mtDNA data from extensive array of Asian populations (21,100 samples from 225 populations), mostly containing HVS and partial coding-region sequences (Supplementary Data 2).
Phylogenetic analysis and haplogroup age estimation
To determine the phylogenetic positions of the newly discovered Cambodian haplogroups, we employed 133 complete mtDNA genome sequences (including the 98 newly sequenced mitogenomes in the present study and the 35 previously reported mitogenomes from Asia14,17,22,23,24,25,28,29,30,32,42,43,48,52) to construct the phylogenetic tree. From the NCBI nucleotide database ( http://www.ncbi.nlm.nih.gov), we selected 35 mitogenomes, including 10 reported but unclassified mitogenomes related with the newly defined haplogroups in Cambodians. We also randomly selected 25 reported mitogenomes that share known haplogroups with Cambodians. With these 35 reference mitogenomes, the phylogenetic positions of the Cambodian sequences can be indicated (Fig. 4). To reveal the detailed structure of the major haplogroups among Cambodians, based on the HVS-I sequence data, reduced median networks were constructed using the programme NETWORK version 4.6.1.0 (Fluxus Engineering)53. The unbiased HVS-I haplotype diversity was calculated with DnaSP (verion 5.10)54 following the method described by Nei55.
The coalescence time to the most recent common ancestor of each haplogroup was estimated using the ρ±σ statistics. We also used the Bayesian method and the maximum likelihood method embedded in BEAST33 and PAML34, respectively, for age estimation. The mutation rates of 16,677 years, 7,884 years and 3,624 years per mutation were used for HVS-I (16,051–16,400), coding region (577–16,023) synonymous mutations and entire mtDNA genome, respectively56. For Bayesian and maximum likelihood, the haplogroups with whole mtDNA genome data and a sample size of n>5 were analysed, and the estimated ages are consistent with those by the rho method (Table 3). To establish the genetic relationships between Cambodians and other Asian populations, we performed PCA based on the frequencies of mtDNA haplogroups according to the method developed by Richards et al.57 in the MVSP3.13 software. Contour maps of the major haplogroups in Cambodians were constructed using Golden Software Surfer 10.0 (Golden Software Inc., USA) with the Kriging algorithm.
Bayesian skyline plot
To reconstruct the demographic changes through time for Cambodian aborigines, we reconstructed BSP41 in BEAST (version 1.7.5)33 with MCMC algorithms58. The BSP was generated using 1,054 Cambodian HVS-I (16038–16462) sequences. A strict molecular clock with a fixed rate of 1.784 × 10−7 substitutions per site per year56 was applied. The MCMC chain was run for 1 × 108 steps, with sampling of parameters every 2,500 steps, and the initial 1 × 107 steps were discarded as burn-in. In all runs, the effective sample size for eight parameters of interest was over 200. The BSP was visualized with Tracer 1.5 ( http://tree.bio.ed.ac.uk/ software/tracer), and the female effective population size was plotted on a log scale by assuming a female generation time of 25 years. Population growth rate was calculated from the skyline plot using method described elsewhere59.
Additional information
Accession Numbers: Sequence data for the 98 mtDNA whole-genome sequences and the 1,054 HVS-I sequences have been deposited in GenBank/EMBL/DDBJ nucleotide core database under accession numbers KC505067 to KC505122, KC887456 to KC887497 and KC504013 to KC505066.
How to cite this article: Zhang, X. et al. Analysis of mitochondrial genome diversity identifies new and ancient maternal lineages in Cambodian aborigine. Nat. Commun. 4:2599 doi: 10.1038/ncomms3599 (2013).
References
Evans, D. et al. A comprehensive archaeological map of the world’s largest preindustrial settlement complex at Angkor, Cambodia. Proc. Natl Acad. Sci. USA 104, 14277–14282 (2007).
Su, B. et al. Y-Chromosome evidence for a northward migration of modern humans into Eastern Asia during the last Ice Age. Am. J. Hum. Genet. 65, 1718–1724 (1999).
Shi, H. et al. Y-chromosome evidence of southern origin of the East Asian-specific haplogroup O3-M122. Am. J. Hum. Genet. 77, 408–419 (2005).
Shi, H. et al. Y chromosome evidence of earliest modern human settlement in East Asia and multiple origins of Tibetan and Japanese populations. BMC Biol. 6, 45 (2008).
Jin, L. & Su, B. Natives or immigrants: modern human origin in east Asia. Nat. Rev. Genet. 1, 126–133 (2000).
Kivisild, T. et al. The emerging limbs and twigs of the East Asian mtDNA tree. Mol. Biol. Evol. 19, 1737–1751 (2002).
Wen, B. et al. Genetic evidence supports demic diffusion of Han culture. Nature 431, 302–305 (2004).
Consortium, H. P.-A. S et al. Mapping human genetic diversity in Asia. Science 326, 1541–1545 (2009).
Bodner, M. et al. Southeast Asian diversity: first insights into the complex mtDNA structure of Laos. BMC Evol. Biol. 11, 49 (2011).
Endicott, P. et al. The genetic origins of the Andaman Islanders. Am. J. Hum. Genet. 72, 178–184 (2003).
Hill, C. et al. A mitochondrial stratigraphy for island southeast Asia. Am. J. Hum. Genet. 80, 29–43 (2007).
Hill, C. et al. Phylogeography and ethnogenesis of aboriginal Southeast Asians. Mol. Biol. Evol. 23, 2480–2491 (2006).
Mona, S. et al. Genetic admixture history of Eastern Indonesia as revealed by Y-chromosome and mitochondrial DNA analysis. Mol. Biol. Evol. 26, 1865–1877 (2009).
Peng, M. S. et al. Tracing the Austronesian footprint in Mainland Southeast Asia: a perspective from mitochondrial DNA. Mol. Biol. Evol. 27, 2417–2430 (2010).
Soares, P. et al. Ancient voyaging and Polynesian origins. Am. J. Hum. Genet. 88, 239–247 (2011).
Soares, P. et al. Climate change and postglacial human dispersals in southeast Asia. Mol. Biol. Evol. 25, 1209–1218 (2008).
Tabbada, K. A. et al. Philippine mitochondrial DNA diversity: a populated viaduct between Taiwan and Indonesia? Mol. Biol. Evol. 27, 21–31 (2010).
Tumonggor, M. K. et al. The Indonesian archipelago: an ancient genetic highway linking Asia and the Pacific. J. Hum. Genet. 58, 165–173 (2013).
Su, B. et al. Polynesian origins: insights from the Y chromosome. Proc. Natl Acad. Sci. USA 97, 8225–8228 (2000).
Black, M. L., Dufall, K., Wise, C., Sullivan, S. & Bittles, A. H. Genetic ancestries in northwest Cambodia. Ann. Hum. Biol. 33, 620–627 (2006).
van Oven, M. & Kayser, M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum. Mutat. 30, E386–E394 (2009).
Kong, Q. P. et al. Large-scale mtDNA screening reveals a surprising matrilineal complexity in east Asia and its implications to the peopling of the region. Mol. Biol. Evol. 28, 513–522 (2011).
Gunnarsdottir, E. D. et al. Larger mitochondrial DNA than Y-chromosome differences between matrilocal and patrilocal groups from Sumatra. Nat. Commun. 2, 228 (2011).
Behar, D. M. et al. A ‘Copernican’ reassessment of the human mitochondrial DNA tree from its root. Am. J. Hum. Genet. 90, 675–684 (2012).
Chandrasekar, A. et al. Updating phylogeny of mitochondrial DNA macrohaplogroup M in India: dispersal of modern human in South Asian corridor. PLoS One 4, e7447 (2009).
van Oven, M. Revision of the mtDNA tree and corresponding haplogroup nomenclature. Proc. Natl Acad. Sci. USA 107, E38–E39 (2010).
Zhao, M. et al. Mitochondrial genome evidence reveals successful Late Paleolithic settlement on the Tibetan Plateau. Proc. Natl Acad. Sci. USA 106, 21230–21235 (2009).
Wang, H. W. et al. Revisiting the role of the Himalayas in peopling Nepal: insights from mitochondrial genomes. J. Hum. Genet. 57, 228–234 (2012).
Fornarino, S. et al. Mitochondrial and Y-chromosome diversity of the Tharus (Nepal): a reservoir of genetic variation. BMC Evol. Biol. 9, 154 (2009).
Kivisild, T. et al. The role of selection in the evolution of human mitochondrial genomes. Genetics 172, 373–387 (2006).
Tanaka, M. et al. Mitochondrial genome variation in eastern Asia and the peopling of Japan. Genome Res. 14, 1832–1850 (2004).
Scholes, C. et al. Genetic diversity and evidence for population admixture in Batak Negritos from Palawan. Am. J. Phys. Anthropol. 146, 62–72 (2011).
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Fucharoen, G., Fucharoen, S. & Horai, S. Mitochondrial DNA polymorphisms in Thailand. J. Hum. Genet. 46, 115–125 (2001).
Kong, Q. P. et al. Updating the East Asian mtDNA phylogeny: a prerequisite for the identification of pathogenic mutations. Hum. Mol. Genet. 15, 2076–2086 (2006).
Wen, B. et al. Genetic structure of Hmong-Mien speaking populations in East Asia as revealed by mtDNA lineages. Mol. Biol. Evol. 22, 725–734 (2005).
Li, H. et al. Mitochondrial DNA diversity and population differentiation in southern East Asia. Am. J. Phys. Anthropol. 134, 481–488 (2007).
Oota, H., Settheetham-Ishida, W., Tiwawech, D., Ishida, T. & Stoneking, M. Human mtDNA and Y-chromosome variation is correlated with matrilocal versus patrilocal residence. Nat. Genet. 29, 20–21 (2001).
Wong, H. Y. et al. Sequence polymorphism of the mitochondrial DNA hypervariable regions I and II in 205 Singapore Malays. Leg Med (Tokyo) 9, 33–37 (2007).
Drummond, A. J., Rambaut, A., Shapiro, B. & Pybus, O. G. Bayesian coalescent inference of past population dynamics from molecular sequences. Mol. Biol. Evol. 22, 1185–1192 (2005).
Dancause, K. N., Chan, C. W., Arunotai, N. H. & Lum, J. K. Origins of the Moken Sea Gypsies inferred from mitochondrial hypervariable region and whole genome sequences. J. Hum. Genet. 54, 86–93 (2009).
Peng, M. S., He, J. D., Liu, H. X. & Zhang, Y. P. Tracing the legacy of the early Hainan Islanders--a perspective from mitochondrial DNA. BMC Evol. Biol. 11, 46 (2011).
Wen, B. et al. Analyses of genetic structure of Tibeto-Burman populations reveals sex-biased admixture in southern Tibeto-Burmans. Am. J. Hum. Genet. 74, 856–865 (2004).
Yao, Y. G., Kong, Q. P., Man, X. Y., Bandelt, H. J. & Zhang, Y. P. Reconstructing the evolutionary history of China: a caveat about inferences drawn from ancient DNA. Mol. Biol. Evol. 20, 214–219 (2003).
Stark, M. T Pre-Angkorian and Angkorian Cambodia. Southeast Asia: From Prehistory to History 89–120 (2004).
Reinecke, A., Laychour, V., Sophady, H. & Sonetra, S. The First Golden Civilization of Cambodia: Unexpected Archaeological Discoveries, 1–39 (Memot Centre for Archaeology. Phnom Penh, Cambodia, (2009).
Macaulay, V. et al. Single, rapid coastal settlement of Asia revealed by analysis of complete mitochondrial genomes. Science 308, 1034–1036 (2005).
Rasmussen, M. et al. An Aboriginal Australian genome reveals separate human dispersals into Asia. Science 334, 94–98 (2011).
Chandler, D. A History of Cambodia In: Westview Press ((2000).
Fendt, L., Zimmermann, B., Daniaux, M. & Parson, W. Sequencing strategy for the whole mitochondrial genome resulting in high quality sequences. BMC Genomics 10, 139 (2009).
Gunnarsdottir, E. D., Li, M., Bauchet, M., Finstermeier, K. & Stoneking, M. High-throughput sequencing of complete human mtDNA genomes from the Philippines. Genome Res. 21, 1–11 (2011).
Bandelt, H. J., Forster, P. & Rohl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48 (1999).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Nei, M. Molecular Evolutionary Genetics[M] Columbia University Press (1987).
Soares, P. et al. Correcting for purifying selection: an improved human mitochondrial molecular clock. Am. J. Hum. Genet. 84, 740–759 (2009).
Richards, M., Macaulay, V., Torroni, A. & Bandelt, H. J. In search of geographical patterns in European mitochondrial DNA. Am. J. Hum. Genet. 71, 1168–1174 (2002).
Drummond, A. J., Nicholls, G. K., Rodrigo, A. G. & Solomon, W. Estimating mutation parameters, population history and genealogy simultaneously from temporally spaced sequence data. Genetics 161, 1307–1320 (2002).
Gignoux, C. R., Henn, B. M. & Mountain, J. L. Rapid, global demographic expansions after the origins of agriculture. Proc. Natl Acad. Sci. USA 108, 6044–6049 (2011).
Acknowledgements
We are grateful to all the volunteers for providing their blood samples. We would like to thank the Departments of Geography & Land Management and of Biology of the Royal University of Phnom Penh of Cambodia for their assistance during sample collection. We also wish to thank Dr Min-Sheng Peng for his assistance in data analyses. This study was supported by the National 973 Program of China (2012CB518202 to X.Q.), the National Natural Science Foundation of China (31130051 and 91231203 to B.S., 31371268 and 91131001 to H.S. and 31371269 to X.Q.) and the Natural Science Foundation of Yunnan Province (2010CI044 to H.S.). M.v.O. was supported in part by a grant from the Netherlands Genomic Initiative (NGI)/Netherlands Organization for Scientific Research (NWO) within the framework of the Forensic Genomics Consortium Netherlands (FGCN) and by the Erasmus MC University Medical Center Rotterdam.
Author information
Authors and Affiliations
Contributions
B.S. and H.S. designed the experiment; X.Z., X.Q., Z.Y., B.Se., T.S., L.B., H.S.A., H.Sa. and H.S. collected the samples; X.Z. and X.Q. collected the data; H.Z. and Q.L. provided technical assistance in the experiments; X.Z., X.Q., M.v.O., H.S. and B.S. conducted data analysis; X.Z., X.Q., H.S. and B.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Data 1
MtDNA Sequence Variations of 1054 Samples from Cambodia (XLS 229 kb)
Supplementary Data 2
MtDNA Haplogroup Frequencies (%) in Asian Populations (XLS 439 kb)
Rights and permissions
About this article
Cite this article
Zhang, X., Qi, X., Yang, Z. et al. Analysis of mitochondrial genome diversity identifies new and ancient maternal lineages in Cambodian aborigines. Nat Commun 4, 2599 (2013). https://doi.org/10.1038/ncomms3599
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms3599
This article is cited by
-
Unraveling the mitochondrial phylogenetic landscape of Thailand reveals complex admixture and demographic dynamics
Scientific Reports (2023)
-
The matrilineal ancestry of Nepali populations
Human Genetics (2023)
-
An in-depth analysis of the mitochondrial phylogenetic landscape of Cambodia
Scientific Reports (2021)
-
Massively parallel sequencing of human skeletal remains in Vietnam using the precision ID mtDNA control region panel on the Ion S5™ system
International Journal of Legal Medicine (2021)
-
Complete human mtDNA genome sequences from Vietnam and the phylogeography of Mainland Southeast Asia
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.