Abstract
Unravelling the basis of variation in inflorescence architecture is important to understanding how the huge diversity in plant form has been generated. Inflorescences are divided between simple, as in Arabidopsis, with flowers directly formed at the main primary inflorescence axis, and compound, as in legumes, where they are formed at secondary or even higher order axes. The formation of secondary inflorescences predicts a novel genetic function in the development of the compound inflorescences. Here we show that in pea this function is controlled by VEGETATIVE1 (VEG1), whose mutation replaces secondary inflorescences by vegetative branches. We identify VEG1 as an AGL79-like MADS-box gene that specifies secondary inflorescence meristem identity. VEG1 misexpression in meristem identity mutants causes ectopic secondary inflorescence formation, suggesting a model for compound inflorescence development based on antagonistic interactions between VEG1 and genes conferring primary inflorescence and floral identity. Our study defines a novel mechanism to generate inflorescence complexity.
Similar content being viewed by others
Introduction
Understanding the basis of diversity in form is a major challenge in developmental biology. An important feature contributing to form diversity in angiosperms is the variation in the architecture of inflorescences, the structures that bear the flowers1,2. Inflorescence architecture is also important because it conditions flower and fruit production and, therefore, crop yield3. A process central to generation of inflorescence architecture diversity is inflorescence branching, with a major distinction between simple inflorescences, as in Arabidopsis, where flowers derive from the primary inflorescence axis (Fig. 1a,b,c), and compound inflorescences, where flowers form at secondary (or higher order) branches (Fig. 1d,e,f)2.
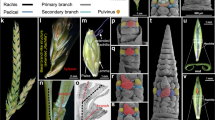
(a) Simple raceme of Arabidopsis thaliana. (b) Diagram of the architecture of the inflorescence showed in (a). (c) Schematic representation of meristem identity in the inflorescence showed in (a). (d) Compound raceme of the legume species Medicago sativa. (e) Diagram of the architecture of the inflorescence showed in (d). (f) Schematic representation of meristem identity in the inflorescence showed in (d). In Arabidopsis, flowers appear in the primary inflorescence stem (I1) whereas in M. sativa they appear in secondary inflorescence branches (I2). V, vegetative meristem; I1, primary inflorescence meristem; I2, secondary inflorescence meristem; F, floral meristem.
Genetic control of compound inflorescence development has been best studied in grasses, particularly in rice and maize, where flowers are formed from the spikelet meristem, an inflorescence meristem that frequently derives from lateral secondary or tertiary order inflorescence branches4,5. This higher complexity suggests that novel genetic functions must exist for the formation of the high-order inflorescence meristems, that often are not simple reiterations of the primary inflorescence meristem. In fact, a number of genes have been characterized that control specific aspects of compound inflorescence development in grasses, such as the formation of lateral inflorescences or the determinacy of spikelet meristems4,5,6,7. Compound inflorescence development has also been studied in Solanaceae, where two genes that control inflorescence complexity have been isolated. However, the formation of Solanaceae compound inflorescences differs from grasses, not involving high-order inflorescence meristems8.
Legumes (Fabaceae), the third largest angiosperm family, also have compound inflorescences9,10, where flowers are produced on lateral secondary inflorescence branches (Fig. 1d,e,f). In this respect, legume inflorescence architecture is similar to that of grasses and different to Solanaceae. Given that legumes are only distantly related to grasses, a relevant question is whether legumes have generated secondary inflorescence meristems through a distinct mechanism involving novel specific functions. To understand the development of the compound inflorescence in legumes, we analysed the classical vegetative1 (veg1) mutant from pea (Pisum sativum), which displays a phenotype that suggests severe defects in the formation of secondary inflorescence meristems11,12, and, therefore, we hypothesized that it might be defective in such novel function.
Results
VEG1 is required to make secondary inflorescences
The pea inflorescence is a compound raceme, typical of many legumes1,10,13. During the vegetative phase, each of the nodes produced by the vegetative shoot apical meristem (SAM) consists of a leaf with a shoot axillary meristem that generally remains dormant until the floral transition has occurred (Fig. 2a–c). At floral transition, the SAM becomes a primary inflorescence (I1) meristem, with indeterminate growth, that produces nodes with axillary meristems that grow out immediately (Fig. 2a–c). These secondary inflorescence (I2) meristems each produce 1–3 nodes bearing floral meristems before terminating in a stub1,13 (Fig. 2d). Therefore, the I2 meristem, interposed between the I1 and the floral meristems, represents an additional level of complexity compared with the simple raceme of Arabidopsis (Fig. 1c).
(a) Pea wild-type (WT) and veg1 plants grown for 11 weeks. Whereas the upper nodes of the wild type contain secondary inflorescences (I2) with flowers (pods, arrowheads), veg1 has not produced any flower. (b) The same plants as in (a), where leaves have been removed. As in the wild type, the axillary buds of the lower nodes of veg1 remain dormant. However, at the nodes where I2s (arrowheads) have grown in the wild type, branches (arrowheads) have grown out in veg1. (c) Diagrams (top) and schematic representation of meristem identity (bottom) of the wild-type and veg1 mutant plants. The vegetative meristem (V) becomes a primary inflorescence meristem (I1) that produces secondary inflorescence meristems (I2) that form flowers (F). In the veg1 plants the I2s are replaced by vegetative branches similar to I1s (I1*). Arrowheads, indeterminate shoots; open circles, flowers; orange triangles, stubs. (d) Structures formed in equivalent apical nodes of the inflorescence stems of wild-type and veg1 plants. Whereas the wild type has produced an I2 with two flowers and a stub (arrowhead in inset), veg1 has produced a vegetative branch. (e) Scanning electron micrographs of the main shoot apex of wild type and veg1. During the vegetative phase (top), in both the wild type and veg1, the vegetative SAM (V, highlighted in green) generates leaves with dormant axillary meristems. After transition to flowering (bottom), the SAM in the wild type becomes a primary inflorescence meristem (I1) that produces I2 meristems (orange) with floral meristems (F, pink). At an equivalent developmental stage, the SAM of veg1 produces leaves with axillary meristems (I1*, green) that grow out following the same pattern as the I1 meristem. Scale bars, 200 μm.
The vegetative1 (veg1) mutant derives from X-ray mutagenesis11, and, among numerous flowering-related mutants in pea, it is distinctive in that it never produces flowers under any conditions (Fig. 2a), and may persist in a vegetative state for over 6 months under conditions where wild-type plants flower after several weeks12. During the vegetative phase, veg1 and wild-type plants were indistinguishable, with dormant axillary meristems (Fig. 2a–c,e). However, after the floral transition, nodes equivalent to those occupied by I2s in wild type were also released from dormancy in veg1, but produced only vegetative shoots (Fig. 2a,b,d,e; Supplementary Fig. S1). This suggests that veg1 mutants undergo a phase transition but are impaired in the subsequent specification of secondary inflorescence identity.
To test this possibility, we compared the expression of inflorescence markers in wild type and veg1. In wild-type plants, the floral transition is associated with transcriptional induction of DETERMINATE (DET) and PROLIFERATING INFLORESCENCE MERISTEM (PIM)14,15,16,17 (Fig. 3a), which, like their Arabidopsis orthologues TERMINAL FLOWER 1 (TFL1)18 and APETALA 1 (AP1)19, control the identity of the inflorescence and the floral meristems14,15,17,18,19,20. In 5-week-old plants, where PIM is clearly expressed in wild-type plants, expression was not observed in veg1 (Fig. 3a), consistent with the absence of flowers or floral organs. However, the inflorescence marker DET was upregulated in veg1 similarly to wild type. Taken together, these results support the idea that the apex of veg1 plants go through floral transition at approximately the same time as the wild type. This implies that the extreme non-flowering phenotype of the veg1 mutant plants does not represent a block in floral induction or a defect in timing of the floral transition, but instead reflects a failure of the lateral meristems produced by the primary 'inflorescence' apex of veg1 to develop as secondary inflorescences. Therefore, VEG1 is required for the pea inflorescence apex to make the I2 meristems.
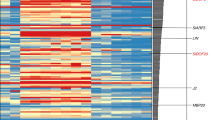
(a) mRNA levels of PIM (left) and DET (right) in the shoot apices of the wild type and veg1. Relative mRNA levels were determined by RT–qPCR. Samples were from main-shoot apices of 2-week old plants (veget. apices), before wild-type plants had gone through the floral transition, and of 5-week-old plants ('inflo' apices), when the wild type had started producing flowers. Values represent the means of two biological replicates ±s.e. (b–e) In situ hybridization of DET mRNA in the shoot apices of wild type and veg1. Samples were from the main apices of 4-week-old plants, when the wild type had gone through the floral transition. In the wild-type apex (b) DET expression was detected below the dome of the primary inflorescence meristem (I1), in the vasculature, and in the boundary between the secondary inflorescence meristem (I2) and the incipient leaf primordium (LP), but not in the I2 or in the floral meristems (F), as represented in the diagram (c), for clarity. Similarly, in veg1, expression was also detected below the primary shoot meristem (I1 SAM, arrowhead), in a section through the centre of the apex (d). A deeper section from the same apex (e) shows that in veg1 DET is also expressed in a lateral shoot meristem (I1*, arrowhead).
To elucidate the nature of the vegetative branches that replace the I2s in the veg1 mutant, we further analysed the expression of DET, in more detail, by in situ hybridization. In wild-type apices, DET expression was restricted to the primary inflorescence meristem (Fig. 3b,c). However, in veg1 apices DET was expressed not only in the I1 meristem but also in the lateral meristems that are formed in place of the I2 meristems (Fig. 3d,e). This indicates that the lateral branches produced by veg1 plants after the transition possess I1 identity, and further supports a role for VEG1 in the specification of I2 identity (Fig. 2c).
Although VEG1, thus, seems required for the specification of I2 meristems, a further question concerns whether it is also required for the formation of flowers. In contrast to the constitutive non-flowering phenotype of the veg1 mutant, the veg1 det double mutant does produce flowers13 (Supplementary Fig. S2a,b). However, in contrast to the det single mutant, where the primary inflorescence ends in a terminal I2, the veg1 det primary inflorescence does not produce I2s, and ends with the production of a terminal flower, often after producing a flower directly from the axil of a leaf at the node below13 (Supplementary Fig. S2c-f). Consistent with that, we saw that expression of PIM is induced in the apex of veg1 det plants (Supplementary Fig. S2g,h). This again indicates that VEG1 is required for the specification of the I2 meristems, and shows that it is not directly required for the expression of the floral meristem identity genes or the formation of floral meristems.
VEG1 is a MADS-box gene from the AGL79 clade
To identify a candidate gene for VEG1, we adopted a comparative mapping strategy using the related model legume Medicago truncatula, which has inflorescence architecture identical to pea21. The VEG1 locus was initially observed to map to the bottom of pea linkage group V (top of Medicago chromosome 7), near the MADS-box gene PsSEPALLATA1 (PsSEP1)22, and we found that this gene was deleted in veg1. However, consistent with the well-documented role for SEP genes in floral organ identity23,24, we found that PsSEP1 was expressed in floral but not in I2 meristems, thus arguing against PsSEP1 as a candidate for VEG1. In M. truncatula, a second MADS-box gene, MtFULc, is located on the same BAC clone as MtSEP1 (ref. 22) (Fig. 4a). We isolated the pea orthologue of this gene (PsFULc), confirmed its map position very close to PsSEP1, and observed that it is also deleted in the veg1 mutant (Fig. 4b). PsFULc and MtFULc belong to the AGL79 clade of the AP1/SQUA/FUL genes25,26 (Supplementary Fig. S3). Genes from the AP1/SQUA/FUL lineage are involved in the specification of meristem identity20, which suggested that PsFULc was a good candidate for VEG1.
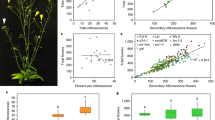
(a) In M. truncatula, MtFULc is located 17 Kb from MtSEP1, whose pea homologue maps to the VEG1 locus and is deleted in the veg1 mutant. (b) Lesions in the psfulc mutant alleles described in this work. Left, Southern blot on DNA of plants from a F2 population segregating for veg1, hybridized with a PsFULc probe. The same DNA samples were digested with BamHI and with EcoRI. Hybridization was observed with DNA from plants with a wild-type phenotype (2, 4, 6 and 7) but not from homozygous veg1 plants (3 and 5), showing that PsFULc is deleted in veg1. Right, mutations in the psfulc-2 and psfulc-3 alleles. The nucleotide changes in the mutant sequences are indicated in red. The G-to-A mutation in fulc-2 destroyed the splice donor site of exon 3. The C-to-T mutation in psfulc-3 caused a Q102STOP change. (c,d) Phenotype of the psfulc-2 and psfulc-3 mutant plants. Both plants show a non-flowering veg1 phenotype, where secondary inflorescences (I2) are replaced by vegetative branches. (e) veg1-like phenotype of a PsFULc-VIGS plant. In the PsFULc-silenced plant, the I2s appear in later nodes than in the control plant, and branches (arrowheads) develop in the nodes where I2s (arrowheads) appear in the control plant.
To assess whether the deletion of PsFULc was the cause of the veg1 phenotype, we characterized additional mutant alleles of PsFULc. Forward screening of an ethyl methanesulfonate (EMS)-mutagenized population27 identified a single M2 plant in which secondary inflorescences were replaced by vegetative branches, as in veg1 (Fig. 4c). Sequencing of PsSEP1- and PsFULc-coding regions in this mutant (psfulc-2) revealed a wild-type PsSEP1 sequence, but identified a G-to-A mutation typical of EMS mutagenesis at the 5′-splice junction of the fourth intron in PsFULc (Fig. 4b). A third PsFULc mutant allele (psfulc-3), carrying a Q102STOP mutation, was identified in reverse genetic screening of an EMS-mutagenized Targeting-Induced Local Lesions in Genomes (TILLING) population28. Like psfulc-2, this mutant also showed a clear veg1 phenotype (Fig. 4d) and carried a PsSEP1-coding region identical to wild type. As a third line of evidence, we used virus-induced gene silencing (VIGS)29 to specifically suppress expression of PsFULc, and found that PsFULc-VIGS plants partly phenocopied the veg1 phenotype. The node at which the first I2 appeared was significantly higher in PsFULc-VIGS plants (14.5±1.6) than in control plants (10.7±0.9) and the intervening nodes were occupied by vegetative branches (Fig. 4e; Supplementary Fig. S1), as in veg1 plants. In summary, the phenotype of the PsFULc-VIGS plants, and the defects of psfulc-2 and psfulc-3, identical to those of the original veg1 X-ray mutant, show that the veg1 phenotype is specifically caused by the loss of PsFULc and that the deletion of other sequences in the X-ray veg1 mutant does not significantly contribute to it. Therefore, we subsequently refer to PsFULc as VEG1.
VEG1 expression marks secondary inflorescence meristems
To assess whether the expression of VEG1 is consistent with its proposed role in the specification of I2 meristem identity, we examined its expression pattern in wild-type plants and in mutants either completely lacking or producing ectopic secondary inflorescences. Analysis by RT–qPCR in wild-type plants showed that VEG1 is expressed in inflorescence apices and mature flowers (Supplementary Fig. S4a). Consistent with that, a time-course expression analysis showed that VEG1 upregulation occurs during the floral transition, after FTb2, a leaf marker of floral induction16, and before the floral meristem identity gene PIM (Supplementary Fig. S4b). Further analysis by in situ hybridization on wild-type inflorescence apices (Fig. 5a,b) showed that VEG1 is expressed in the I2 meristems but is not expressed in the apical meristem of the primary inflorescence (I1) or in the young floral meristems, which show strong expression of PIM (Fig. 5c,d). These results further support the idea that VEG1 specifies the identity of I2 meristems.
(a) Inflorescence of a pea wild-type plant. The inflorescence apex (arrowhead) exhibits indeterminate growth. (b) SEM image showing the different meristem types in a wild-type inflorescence. The SAM is a primary inflorescence (I1) meristem that has produced secondary inflorescence (I2) meristems, one of which has produced a floral meristem (F). (c) In situ hybridization of VEG1 mRNA in the inflorescence apex of the wild type. VEG1 is expressed in the lateral I2 meristems, not in the I1 neither in the floral meristem (F). (d) In situ hybridization of PIM mRNA in the wild-type inflorescence, in a contiguous section to that in (c). The PIM signal is observed only in the floral meristem, which does not express VEG1. (e) Inflorescence of a det mutant, where the apex has converted into an I2 (arrowhead). The terminal I2 stem subtends a flower and ends into a stub (arrowhead in inset). (f) SEM image of a det inflorescence apex. The SAM has the characteristic shape of an I2 meristem (I2*). (g) Schematic representation of meristem identity in the det inflorescence. V, vegetative meristem; orange triangles, stubs. (h) Inflorescence of a pim mutant, showing proliferative I2 s (arrowhead) with abnormal flowers. (i) SEM image of a pim inflorescence apex, showing I2s that, rather than flowers, produce new I2s (I2*). One of the proliferative I2s is producing a floral primordium (F*). (j) Schematic representation of meristem identity in the pim inflorescence. (k) In situ hybridization of VEG1 mRNA in the inflorescence apex of det. In the det mutant VEG1 is also expressed in the SAM (I2*), which is converted into an I2. (l) In situ hybridization of VEG1 mRNA in the inflorescence apex of pim. Expression of VEG1 is observed not only in the lateral I2s but also in the meristem being formed by the I2 at the right (I2*), which is converted from a floral meristem to an I2 meristem. Scale bars, 100 μm.
Mutations in DET or PIM, homologues of TFL1 and AP1, respectively, cause the conversion of other meristems of the pea inflorescence into I2 meristems. Thus, while the primary inflorescence meristem (the inflorescence SAM) of wild-type pea plants shows indeterminate growth, producing I2s only in lateral positions (Figs 2c and 5a), in det mutants, the inflorescence SAM shows determinate growth, and terminates in a typical I2 (refs. 15,30) (Fig. 5e,f,g). On the other hand, the lateral meristems in the I2 of pim mutant plants, rather than acquiring floral identity, as in the wild type (Figs 2c and 5b), retain I2 meristem identity and themselves generate lateral meristems with I2 identity in a reiterative manner, with some of these supernumerary I2 meristems eventually producing flowers17,30 (Fig. 5h,i,j). If VEG1 specifies I2 identity, it would be expected that the formation of the ectopic I2 meristems in these mutants were accompanied by changes in VEG1 expression. In fact, in situ hybridization showed that in the det mutant inflorescence VEG1 is expressed not only in the lateral I2 meristems but also in the apical meristem (Fig. 5k), and that the supernumerary I2 meristems in the pim mutant exhibit VEG1 expression (Fig. 5l). This indicates that DET and PIM repress VEG1 expression and restrict it to the I2, and suggests that the ectopic expression of VEG1 in det and pim mutants causes the conversion of I1 and floral meristems, respectively, into I2 meristems.
Severe mutations in the VEGETATIVE2 (VEG2) or GIGAS genes cause a non-flowering phenotype similar to the veg1 mutant (gigas only under long-day photoperiods), with no formation of secondary inflorescences16,31. As in veg1, the plants of both mutants show outgrowth of vegetative lateral branches at nodes occupied in wild-type plants by I2s (ref. 16) (Sussmilch et al., unpublished). This similarity suggests that these genes participate in the same genetic network as VEG1 and raises the question of what is their relative position in that network. Analysis by RT–qPCR showed that although VEG1 was upregulated in shoot apices of 4-week-old wild-type plants, its expression was not detectable in either mutant (Supplementary Fig. S4c) after 6 weeks. This indicates that VEG1 acts downstream of VEG2 and GIGAS and suggests that they participate in the activation of VEG1 expression. This seems particularly likely for GIGAS16, in view of the fact that its Arabidopsis homologue FLOWERING LOCUS T (FT) is a direct activator of the floral identity genes AP1 and FUL32,33,34,35, which are MADS-box genes from the same lineage as VEG1 (Supplementary Fig. S3).
Discussion
Our data suggest a genetic model that explains the specification of the identity of the different meristem types formed in the pea compound inflorescence (Fig. 6a). In this model, expression of VEG1, which is required to specify I2 identity, is restricted to the I2 meristem by DET, which represses it in the I1 meristem, and by PIM, which represses it in the floral meristem. Conversely, expression of DET itself is restricted to the I1 and excluded from the I2 through repression by VEG1. The simplest interpretation of the veg1 phenotype is, therefore, that it results from ectopic DET expression in the lateral I2 meristems, converting them into I1 meristems (Fig. 6b). In addition, the formation of flowers in the veg1 det double mutant suggests that DET represses not only VEG1 but also PIM expression. Finally, the fact that the flowers in veg1 det are directly formed from the I1, as in simple inflorescences, suggests that the default state of the meristems in the pea inflorescence is floral identity, which is normally restricted to the lateral meristems of the I2 by the concerted action of DET and VEG1. In this sense, VEG1 would be required to maintain 'vegetativeness', as defined by Prusinkiewicz et al.36, in the lateral meristems of the I1.
(a) In the pea compound inflorescence, expression of DET (orthologue of TFL1) in the I1, VEG1 in the I2 and PIM (orthologue of AP1) in the floral meristems are required for those meristems to acquire their identity. Expression of these genes in their correct domains is maintained by a network of mutual repressive interactions. (b) The genetic model explains the phenotypes of the pea meristem identity mutants. The absence of DET in the det mutant allows expression of VEG1 in the SAM, which gets converted into an I2. The absence of PIM allows the expression of VEG1 in the floral meristem, which gets converted into a proliferative I2. In absence of VEG1, DET is expressed in all the meristems in the apex, and they get converted into I1s. (c) Meristem identity in the simple inflorescence of Arabidopsis is maintained by a similar genetic network than in pea, but VEG1 function is absent and I2 meristem is not formed.
The network of mutually repressive interactions between DET, VEG1 and PIM resembles the simpler genetic network that controls meristem identity in the inflorescence apex of Arabidopsis (Fig. 6c), where the separation of the inflorescence and floral meristematic domains is achieved by mutual repression between TFL1 and the floral meristem genes AP1 and LEAFY (LFY)20. This similarity suggests a mechanism in which evolutionary modification of a simple raceme (for example, Arabidopsis) into a compound form may have occurred through the appearance of a new function, VEG1, acting between the inflorescence identity function of DET (TFL1) and the floral identity function of PIM (AP1), which leads to the formation of a new intermediate-step meristem, the I2, and therefore to the development of a compound inflorescence.
The VEG1 gene belongs to the AP1/SQUA/FUL lineage, represented in core eudicots by three distinct clades, euAP1, euFUL and AGL79, that likely arose from a common eudicot ancestor through duplication25,26. While no functional information from mutant phenotypes is available for any AGL79-like gene, analysis of several euAP1 genes and of the Arabidopsis euFUL gene FRUITFULL (FUL) indicates that genes in these other sub-clades control the identity of reproductive meristems20,37. This suggests that this basic function was already present in the ancestor of the core-eudicot AP1/SQUA/FUL genes, and we speculate that AGL79 genes, such as VEG1, may have sub-functionalized to specify the identity of I2 meristems.
Is the mechanism of I2 identity specification, through VEG1-like genes, also central for compound inflorescence development in species other than pea? Compound inflorescences are widespread in the Fabaceae family, suggesting a common evolutionary origin. As VEG1 orthologues are found in several papillionoid legumes (Supplementary Fig. S3), and more widely in eudicots, it seems likely that VEG1 function was present early in the evolution of Fabaceae and may have arisen before the origin of this group. However, no monocot orthologue exists for VEG1/AGL79, a core-eudicot-specific gene25,26, and consistent with this, the genetic network controlling compound inflorescence formation in grasses is different to that in legumes and does not seem to involve a VEG1-related function4,5,6,7,38. Within the eudicots, compound inflorescences have also been studied in Solanaceae8, but, in this group, the ontogeny of the compound inflorescence does not involve high-order inflorescence meristems8 and is thus quite distinct from that in either legumes or grasses. Accordingly, the genes known to control complexity in this family are distinct from those in legumes and grasses and also do not include a VEG1 orthologue8. Thus, these groups apparently use distinct genetic networks for inflorescence complexity, suggesting that compound inflorescences have independently appeared several times during angiosperm evolution. This is consistent with the fact that compound inflorescences occur in phylogenetically distant plant families2.
In summary, our work provides the first insight into the genetic network controlling the legume compound inflorescence. We identify a novel mechanism for generation of inflorescence complexity, distinct from that in grasses and Solanaceae, which is based on the function of the VEG1 gene, which acts between the I1 and floral meristem identity genes to specify formation of the I2 meristem. The identification of VEG1 as an AGL79-like MADS-box gene suggests that VEG1 function derives from sub-functionalization of the AGL79 clade within the eudicot AP1/SQUA/FUL genes. More generally, our identification of VEG1 provides an illustration of how the expansion and functional divergence within key regulatory gene families can contribute to the evolution of morphological complexity.
Methods
Plant material and growth conditions
Plants were grown in a greenhouse at 22 °C (day) and 18 °C (night); long-day photoperiods (16 h light/8 h darkness) were maintained with supplementary lighting (400 W Phillips HDK/400 HPI (R) (N). Plants were irrigated with a Hoagland No. 1 solution supplemented with oligoelements39. The origins of the veg1/psfulc-1, det-2, pim-1 and gigas-2 mutants have been previously described11,16,17,40. The psfulc-2 mutant was generated from line NGB5839 by EMS mutagenesis27. The psfulc-3 mutant was isolated by reverse screening in an EMS-mutagenized TILLING population of cultivar Caméor28. The veg2-1 mutant was isolated from fast-neutron mutagenesis of cv. Kaliski41. VIGS experiments29 were performed on the cultivar Boneville. Each mutant was analysed in comparison with its corresponding parental wild-type line.
Gene isolation and phylogenetic analysis
The PsFULc complementary DNA was isolated from cDNA from inflorescence apices using PCR techniques; first, a 550-bp cDNA fragment was amplified by RT–PCR with primers derived from the M. truncatula FULc sequence and the remaining 5′- and 3′-fragments were amplified by PCR from a pea cDNA library14 with primers from the vector and from the PsFULc cDNA fragment. To analyse the sequences of the PsFULc and PsSEP1 genes in the psfulc-2 mutant, several overlapping genomic fragments from each gene were amplified from genomic DNA from the mutant and from the parental wild type. All PCR fragments were cloned in pGEM-T easy (Promega) and sequenced. For the phylogenetic tree of the AP1/SQUA/FUL genes, the deduced amino acid sequences were aligned using the CLUSTALW tool in MACVECTOR 12.0 software (MacVector http://www.macvector.com/) and further refined by hand. Pairwise Poisson genetic distances were estimated from the alignment and a neighbour joining best tree was estimated using systematic tie-breaking and rooted to AMtrAGL2, an Amborella trichopoda orthologue of SEPALLATA.
Mapping of PsFULc and PsSEP1
A population of 92 F2 individuals from a cross between NGB5839 and JI1794 (Sussmilch et al., unpublished) was used to map PsFULc and PsSEP1 in relation to other markers on the bottom of pea linkage group V. The description of the markers used is given in Supplementary Table S1. No recombination was found between PsFULc and PsSEP1, placing them 20 cM below COLa and 40 cM above FTb1 suggesting that they are less than 1 cM apart.
TILLING
To identify TILLING mutants in PsFULc, an EMS mutant population of 4,800 M2 families from Pisum sativum cultivar Cameor was screened. DNA isolation and pooling, PCR amplification and mutation detection were performed, as previously described28. For primer sequences in this and following sections in Methods see Supplementary Table S1.
PsFULc VIGS
Two PsFULc-VIGS plasmids, pCAPE2-PsFULc193 and pCAPE2-PsFULc416, were used for the VIGS experiments. The pCAPE2-PsFULc193 and pCAPE2-PsFULc416 constructs derived from two non-overlapping PsFULc cDNA fragments of 193 bp (positions 166–359 from ATG) and 416 bp (positions 490–906 from ATG), respectively, which were generated by PCR and separately cloned into the VIGS vector pCAPE2, using XbaI and BglII cloning sites present in the PCR primers. The plasmid pCAPE2-Con, containing 400 bp of the GUS gene, was included for comparison as the VIGS control42. Inoculation of plants was carried out as previously described29 with the following modifications. In each experiment, 12 plants were inoculated which each plasmid. At day 5 after inoculation, plants were decapitated and, for each plant, a single, basal, axial shoot was allowed to proliferate into a new primary shoot. At day 50 after decapitation, the newly outgrown primary shoot of each plant was analysed by scoring the number and nodal position of secondary inflorescences and of vegetative axial shoots exceeding 1 cm in length. Similar results were obtained in different experiments with each of the two PsFULc-VIGS plasmids; the data presented in the text corresponds to a representative experiment with the pCAPE2-PsFULc416 plasmid.
Genotyping
To discriminate between the wild-type and fulc-3 mutant alleles in the TILLING M3 family, we used a dCAPS marker with the primers FULc-dCF and FULc-dCR, which amplify a 228 bp fragment from the PsFULc gene (positions 2304–2532 from ATG at the PsFULc gene). FULc-dCF creates one mismatch generating a Sau3AI target site in the wild-type product (but not in psfulc-3), which after digestion produces two fragments of 26 bp and 202 bp. The veg1-1 single mutants and veg1-1 det-2 double mutants used for qPCR analysis were identified from segregating families. veg1-1 mutants were identified by absence of PCR product from primers FULc-2F and FULc-2R, which amplify a 950-bp fragment from the PsFULc gene, with primers TFL1a-1F and TFL1a-Rev03 used for a positive control PCR. det-2 mutants were determined using a CAPS marker with TFL1a-1F and TFL1a-Rev03 primers, which amplify a 764-bp fragment of the PsTFL1a/DET gene, which after digestion with EarI produces two fragments of 143 and 621 bp in the det-2 mutant because of the det-2 CAA-CGA substitution15.
Southern analysis
10 μg of genomic DNA were digested with restriction enzymes and separated on 0.6%. TAE 1× agarose gels run overnight at 1 V cm−1. Southern Blot analysis was performed by standard methods. The probe was a 550-bp fragment, amplified by PCR from the PsFULc cDNA (nucleotides 221–771 from the ATG) and cloned into the pGEM-T easy vector (Promega).
RT–qPCR
Total RNA was extracted using the SV Total RNA isolation system (Promega). RNA concentrations were determined by spectrophotometer analysis using a NanoDrop 8000 (Thermo Scientific). Reverse transcription was conducted in 20 μl with 1 μg of total RNA using the MMLV high-performance reverse transcriptase (Epicenter), according to the manufacturer's instructions. RT-negative (no enzyme) controls were performed to monitor for contamination with genomic DNA. First-strand cDNA was diluted 5 times, and 2 μl was used in each real-time PCR reaction. Real-time PCR reactions using SYBR green chemistry (Sensimix, Quantace, Bioline) were set up with a CAS-1200N robotic liquid handling system (Corbett Research) and run for 50 cycles in a Rotor-Gene RG3000 (Corbett). Two technical replicates and two-to-three biological replicates were performed for each sample. Relative transcript levels were evaluated using the reference gene ACTIN, as previously described43.
In situ hybridization
RNA in situ hybridization with digoxigenin-labelled probes was performed as described44. For PSFULc/VEG1 and DET, RNA antisense probes were generated using as substrate a 450-bp fragment of the PsFULc cDNA (236–686 from ATG) or a 460-bp fragment of the DET cDNA (358–818 from ATG), amplified by PCR and cloned into the pGEM-T Easy vector (Promega). For PIM, the probe was generated from a 767-bp of the 3′-region of the PIM cDNA, cloned into pGEM3Zf (Promega), as described14.
Scanning electron microscopy (SEM)
Samples for SEM were prepared and analysed as previously described45.
Additional information
Accession codes: The sequence data have been deposited in the NCBI GenBank database under accession codes JN974184 (PsFULC/VEG1 cDNA sequence) and JN974185 (PsFULC/VEG1 genomic sequence).
How to cite this article: Berbel, A. et al. VEGETATIVE1 is essential for development of the compound inflorescence in pea. Nat. Commun. 3:797 doi: 10.1038/ncomms1801 (2012).
References
Benlloch, R., Berbel, A., Serrano-Mislata, A. & Madueño, F. Floral initiation and inflorescence architecture: a comparative view. Ann. Bot. 100, 659–676 (2007).
Weberling, F. Morphology of Flowers and Inflorescences (Cambridge University Press, 1992).
Wang, Y. & Li, J. Molecular basis of plant architecture. Annu. Rev. Plant Biol. 59, 253–279 (2008).
Kellogg, E. A. Floral displays: genetic control of grass inflorescences. Curr. Opin. Plant Biol. 10, 26–31 (2007).
Thompson, B. E. & Hake, S. Translational biology: from Arabidopsis flowers to grass inflorescence architecture. Plant Physiol. 149, 38–45 (2009).
Bommert, P., Satoh-Nagasawa, N., Jackson, D. & Hirano, H.- Y. Genetics and evolution of inflorescence and flower development in grasses. Plant Cell Physiol. 46, 69–78 (2005).
Bortiri, E. & Hake, S. Flowering and determinacy in maize. J. Exp. Bot. 58, 909–916 (2007).
Lippman, Z. B. et al. The making of a compound inflorescence in tomato and related nightshades. PLoS Biol. 6, e288 (2008).
Doyle, J. J. & Luckow, M. A. The rest of the iceberg. Legume diversity and evolution in a phylogenetic context. Plant Physiol. 131, 900–910 (2003).
Weberling, F. In Advances in Legume Biology Vol. 29 (eds C.H. Stirton & J.L. Zarucchi) 35–58 (Missouri Botanoical Gardens, 1989).
Gottschalk, W. A Pisum gene preventing transition from the vegetative stage. Pisum Newslet. 11, 10 (1979).
Reid, J. B. & Murfet, I. C. Flowering in Pisum: a fifth locus, veg. Ann. Bot. 53, 369–382 (1984).
Singer, S. Inflorescence Architecture: a developmental genetics approach. Bot. Rev. 65, 1–26 (1999).
Berbel, A. et al. Analysis of PEAM4, the pea AP1 functional homologue, supports a model for AP1-like genes controlling both floral meristem and floral organ identity in different plant species. Plant J. 25, 441–451 (2001).
Foucher, F. et al. DETERMINATE and LATE FLOWERING are two TERMINAL FLOWER1/CENTRORADIALIS homologs that control two distinct phases of flowering initiation and development in pea. Plant Cell 15, 2742–2754 (2003).
Hecht, V. et al. The pea GIGAS gene is a FLOWERING LOCUS T homolog necessary for graft-transmissible specification of flowering but not for responsiveness to photoperiod. Plant Cell 23, 147–161 (2011).
Taylor, S. A. et al. PROLIFERATING INFLORESCENCE MERISTEM, a MADS-box gene that regulates floral meristem identity in pea. Plant Physiol. 129, 1150–1159 (2002).
Bradley, D., Ratcliffe, O., Vincent, C., Carpenter, R. & Coen, E. Inflorescence commitment and architecture in Arabidopsis. Science 275, 80 (1997).
Mandel, M. A., Gustafson-Brown, C., Savidge, B. & Yanofsky, M. F. Molecular characterization of the Arabidopsis floral homeotic gene APETALA1. Nature 360, 273–277 (1992).
Blázquez, M. A., Ferrándiz, C., Madueño, F. & Parcy, F. How floral meristems are built. Plant Mol. Biol. 60, 855–870 (2006).
Benlloch, R. et al. Isolation of mtpim proves Tnt1 a useful reverse genetics tool in Medicago truncatula and uncovers new aspects of AP1-like functions in legumes. Plant Physiol. 142, 972–983 (2006).
Hecht, V. et al. Conservation of Arabidopsis flowering genes in model legumes. Plant Physiol. 137, 1420–1434 (2005).
Ditta, G., Pinyopich, A., Robles, P., Pelaz, S. & Yanofsky, M. F. The SEP4 gene of Arabidopsis thaliana functions in floral organ and meristem identity. Curr. Biol. 14, 1935–1940 (2004).
Pelaz, S., Ditta, G. S., Baumann, E., Wisman, E. & Yanofsky, M. F. B and C floral organ identity functions require SEPALLATA MADS-box genes. Nature 405, 200–203 (2000).
Litt, A. & Irish, V. F. Duplication and diversification in the APETALA1/FRUITFULL floral homeotic gene lineage: implications for the evolution of floral development. Genetics 165, 821–833 (2003).
Shan, H. et al. Patterns of gene duplication and functional diversification during the evolution of the AP1/SQUA subfamily of plant MADS-box genes. Mol. Phylogenet. Evol. 44, 26–41 (2007).
Hecht, V. et al. Pea LATE BLOOMER1 is a GIGANTEA ortholog with roles in photoperiodic flowering, deetiolation, and transcriptional regulation of circadian clock gene homologs. Plant Physiol. 144, 648–661 (2007).
Dalmais, M. et al. UTILLdb, a Pisum sativum in silico forward and reverse genetics tool. Genome Biol. 9, R43 (2008).
Constantin, G. D. et al. Virus-induced gene silencing as a tool for functional genomics in a legume species. Plant J. 40, 622–631 (2004).
Singer, S. R., Maki, S. L. & Mullen, H. J. Specification of meristem identity in Pisum sativum inflorescence development. Flowering Newslet. 18, 26–32 (1994).
Reid, J., Murfet, I., Singer, S., Weller, J. & Taylor, S. Physiological-genetics of flowering inPisum. Seminars in Cell and Developmental Biology 7, 455–463 (1996).
Abe, M. et al. FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science 309, 1052–1056 (2005).
Giakountis, A. & Coupland, G. Phloem transport of flowering signals. Curr. Opin. Plant Biol. 11, 687–694 (2008).
Teper-Bamnolker, P. & Samach, A. The flowering integrator FT regulates SEPALLATA3 and FRUITFULL accumulation in Arabidopsis leaves. Plant Cell 17, 2661–2675 (2005).
Wigge, P. A. et al. Integration of spatial and temporal information during floral induction in Arabidopsis. Science 309, 1056–1059 (2005).
Prusinkiewicz, P., Erasmus, Y., Lane, B., Harder, L. D. & Coen, E. Evolution and development of inflorescence architectures. Science 316, 1452–1456 (2007).
Litt, A. An evaluation of A-Function: evidence from the APETALA1 and APETALA2 gene lineages. Int. J. Plant Sci. 168, 73–91 (2007).
Gallavotti, A. et al. The control of axillary meristem fate in the maize ramosa pathway. Development 137, 2849–2856 (2010).
Hewitt, Y. Sand and Water Culture Methods used in the Study of Plant Nutrition 2nd edn, (Farnham: Commonwealth Agricultural Bureau, 1966).
Singer, S., Hsiung, L. & Huber, S. Determinate (det) mutant of Pisum sativum (Leguminosae: Papilionoideae) exhibits an indeterminate growth pattern. Am. J. Bot. 77, 130–135 (1990).
Murfet, I. C. & Reid, J B. In Peas - Genetics, Molecular Biology and Biotechnology (eds R Casey & D R Davies) 165–216 (CAB International, 1993).
Constantin, G. D., Grønlund, M., Johansen, I. E., Stougaard, J. & Lund, O. S. Virus-induced gene silencing (VIGS) as a reverse genetic tool to study development of symbiotic root nodules. Mol. Plant Microbe Interact. 21, 720–727 (2008).
Weller, J., Hecht, V., Vander Schoor, J., Davidson, S. & Ross, J. Light regulation of gibberellin biosynthesis in Pea is mediated through the COP1/HY5 Pathway. Plant Cell 3, 800–813 (2009).
Ferrándiz, C., Gu, Q., Martienssen, R. & Yanofsky, M. F. Redundant regulation of meristem identity and plant architecture by FRUITFULL, APETALA1 and CAULIFLOWER. Development 127, 725–734 (2000).
Berbel, A. et al. Functional conservation of PISTILLATA activity in a pea homolog lacking the PI motif. Plant Physiol. 139, 174–185 (2005).
Acknowledgements
We thank Christine Le Signor for sending the seed from the pea TILLING lines, and to Miguel Blázquez for critical reading of the manuscript. This work was supported by grants from Spanish Ministerio de Ciencia e Innovación (BIO2009-10876 to F.M. and BIO2009-09920 to C.F.), the Consellería d'Empresa Universitat i Ciencia from the Generalitat Valenciana (GV03-066 to F.M., C.F. and J.P.B.), the EU-FP6 (GL-IP, FP6-2002–FOOD–1–506223, to F.M., C.F., J.P.B., T.H.N.E and A.Bendahmane), the EU-FP4 (EUdicotMap, BIOCT972170, to T.H.N.E. and S.A.T.), the Australian Research Council (Discovery Porject DP6556508 to J.L.W., V.H. and F.C.S.) and the Danish Agricultural and Veterinary Research Council (grants nos. 9702802 and 53-00-0330 to O.S.L.).
Author information
Authors and Affiliations
Contributions
F.M. and C.F. conceived the project and designed the experiments together with J.L.W., O.S.L., A.Ben., T.H.N.E. and J.P.B. A.Ber. and C.F. performed the experiments together with V.H., O.S.L., M.D., F.C.S. and S.A.T., F.M. wrote the paper together with J.L.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1-S4 and Supplementary Table S1 (PDF 3626 kb)
Rights and permissions
About this article
Cite this article
Berbel, A., Ferrándiz, C., Hecht, V. et al. VEGETATIVE1 is essential for development of the compound inflorescence in pea. Nat Commun 3, 797 (2012). https://doi.org/10.1038/ncomms1801
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms1801
This article is cited by
-
Innovations in functional genomics and molecular breeding of pea: exploring advances and opportunities
aBIOTECH (2024)
-
Recombinant inbred lines derived from wide crosses in Pisum
Scientific Reports (2023)
-
Transcriptome landscape of early inflorescence developmental stages identifies key flowering time regulators in chickpea
Plant Molecular Biology (2022)
-
TILLING by sequencing (TbyS) reveals mutations in flowering control genes that are associated with altered plant architecture in Mungbean (Vigna radiata (L.) R. Wilczek)
Genetic Resources and Crop Evolution (2021)
-
GmFULa improves soybean yield by enhancing carbon assimilation without altering flowering time or maturity
Plant Cell Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.