Abstract
Plants resist infection and herbivory with innate immune responses that are often associated with reduced growth. Despite the importance of growth-defense tradeoffs in shaping plant productivity in natural and agricultural ecosystems, the molecular mechanisms that link growth and immunity are poorly understood. Here, we demonstrate that growth-defense tradeoffs mediated by the hormone jasmonate are uncoupled in an Arabidopsis mutant (jazQ phyB) lacking a quintet of Jasmonate ZIM-domain transcriptional repressors and the photoreceptor phyB. Analysis of epistatic interactions between jazQ and phyB reveal that growth inhibition associated with enhanced anti-insect resistance is likely not caused by diversion of photoassimilates from growth to defense but rather by a conserved transcriptional network that is hardwired to attenuate growth upon activation of jasmonate signalling. The ability to unlock growth-defense tradeoffs through relief of transcription repression provides an approach to assemble functional plant traits in new and potentially useful ways.
Similar content being viewed by others
Introduction
Plants continuously monitor external cues to tailor their growth, development and defensive capabilities in ways that optimize reproductive success, especially in environments where nutrients and light may be scarce. Theories to explain the costs and patterns of plant defense often invoke the existence of physiological tradeoffs that arise from allocation of limited resources to protective compounds and associated morphological structures1,2,3. Thus, a common interpretation of the ‘dilemma of plants to grow or defend’3 is that elevated expression of defense traits consumes metabolic resources at the expense of growth, whereas rapid growth, such as that triggered by shade light from competitors, diverts resources that could otherwise be invested in the defense arsenal. Although the concept of growth-defense tradeoffs is a major paradigm in ecological studies of plant resistance to herbivores and pathogens1,2,3,4, recent studies question the simplistic view that allocation of limited metabolic resources to one process necessarily reduces energetic expenditures in the other process5,6,7,8,9,10. These collective studies highlight the need to develop a more accurate conceptual framework for understanding how tradeoffs between growth and immunity optimize plant fitness in dynamic environments.
Here, we sought to exploit genetic tools in Arabidopsis thaliana to better understand how growth and defense are mechanistically integrated and to explicitly test whether the underlying signalling networks can be reconfigured to allow for simultaneous growth and defense. Our approach was based on the fact that defense responses mediated by the lipid-derived hormone jasmonate (JA) are associated with potent growth inhibition of leaves and roots11,12,13,14. We show that growth-defense tradeoffs are uncoupled in leaves of an Arabidopsis mutant (jazQ phyB) lacking five Jasmonate ZIM-domain (JAZ) transcriptional repressors and the photoreceptor phytochrome B (phyB). The concomitant robust growth and heightened anti-insect defense of jazQ phyB plants can be attributed, at least in part, to parallel activation of MYC and PIF transcription factors that in wild-type (WT) plants are repressed by JAZ and phyB, respectively. We further show that the capacity of jazQ phyB plants to grow and defend well simultaneously is associated with changes in leaf architecture and increased partitioning of carbon to leaf-area growth, but does not depend on increased leaf area-based photosynthesis. Thus, growth-defense antagonism in this system does not appear to be caused by constraints on the availability of metabolic resources that fuel growth and defensive processes but rather by a hormone-linked transcriptional network that is hardwired to restrict growth and upon activation of JA signalling. These collective findings highlight the importance of transcriptional repressor proteins in optimizing growth-defense balance, and further suggest that genetic modification of pathways that integrate defense and light signalling is a potential strategy to combine growth and defense traits in new ways.
Results
A jaz quintuple mutant exhibits constitutive JA responses
We devised a genetic screen to identify mutants of Arabidopsis that display enhanced JA-regulated defense against insect herbivory without an associated reduction in leaf growth. This screen was based on a signalling model15 predicting that removal of JAZ repressor proteins would constitutively activate anti-insect defense and inhibit growth of vegetative tissues (Fig. 1a). Such a phenotype has not been previously observed for jaz loss-of-function mutants of Arabidopsis, most likely because of functional redundancy between the 13 members of the JAZ gene family16. We thus developed an Arabidopsis line (jaz quintuple or jazQ) with T-DNA insertion mutations in five JAZ genes (JAZ1/3/4/9/10) (Supplementary Fig. 1). These JAZs were selected on the basis of their phylogenetic relationship, their demonstrated role in inhibiting various transcription factors (for example, MYCs) that execute defense responses, and their capacity to interact with DELLA proteins that antagonistically link JA signalling to gibberellic acid (GA)-mediated growth responses (Fig. 1a)17,18,19,20,21,22. Root growth assays showed that jazQ seedlings have both an increased sensitivity to exogenous JA and a constitutive short-root phenotype in comparison to WT seedlings (Fig. 1b and Supplementary Fig. 1). We also compared the hormone sensitivity of jazQ seedlings to the jaz10-1 mutant, which is one of the few jaz single mutants to exhibit enhanced sensitivity to JA11,23,24. Consistent with the notion that JAZ family members serve partially redundant roles in JA signalling, jazQ roots were significantly more sensitive to JA than the jaz10-1 mutant (Supplementary Fig. 1). These findings support a key role for multiple JAZs in the control of root growth. Our data are also consistent with recent genetic analysis of the co-repressor Novel Interactor of JAZ (NINJA), which directly interacts with JAZs to negatively regulate JA signalling in roots14,25.
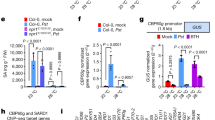
(a) Simple model of the JA-GA signalling network that governs growth and defense. (b) Photograph of WT and jazQ (jQ) seedlings grown in the absence or presence of 25 μM MeJA (MJ). (c) Accumulation of glucosinolates in WT (open bar) and jazQ (blue) seedlings. Compound abbreviations are listed in Methods section. (d) Anthocyanin accumulation in petioles of 4-week-old plants. (e) Trichoplusia ni weight after feeding on WT (33 larvae) and jazQ (38 larvae) plants for 10 days. (f) Photograph of 4-week-old soil-grown WT and jazQ plants. Data in all graphs represent the mean±s.e.m. of at least 10 biological replicates. Asterisks in c–e denote significant differences between WT and jazQ at P<0.05 (Student’s t-test).
Analysis of defense-related phenotypes showed that glucosinolates and anthocyanins, whose biosynthesis in Arabidopsis is positively regulated by JA22,26, accumulate to higher levels in jazQ seedlings than WT (Fig. 1c,d). We also found that soil-grown jazQ plants had remarkably heightened resistance to attack by the generalist herbivore Trichoplusia ni (Fig. 1e). In contrast to these elevated defense traits, leaf area, petiole length and rosette dry weight were all reduced in jazQ relative to WT (Fig. 1f and Supplementary Fig. 2). jazQ also delayed the time to bolting but did not affect the number of leaves at the time of bolting (Supplementary Fig. 2). These results demonstrate that jazQ plants exhibit constitutive growth-defense antagonism (that is, reduced growth with enhanced defense) and thus provide a unique genetic model with which to interrogate how the JA branch of immunity is linked to growth.
Loss of PHYB uncouples growth-defense tradeoffs in jazQ
We visually screened an ethyl methanesulfonate (EMS) -mutagenized population of jazQ for mutants with increased rosette size and persistence of elevated leaf anthocyanin content. Among several suppressor of jazQ (sjq) mutants identified, one line (sjq11) showed a particularly striking leaf growth pattern that was heritable in subsequent generations (Fig. 2a). Importantly, bioassays performed with T. ni larvae showed that sjq11 plants also maintained heightened resistance to herbivory (Fig. 2b). Initial characterization of sjq11 revealed phenotypes similar to those described for phytochrome B (phyB) photoreceptor mutants, including early flowering time and elongated hypocotyls and petioles under continuous white light27. Genetic allelism tests and DNA sequencing confirmed that sjq11 harbours a null mutation in the PHYB gene (Supplementary Fig. 3). To eliminate the possibility that additional EMS mutations contribute to the sjq11 phenotype, further studies were performed with a jazQ phyB sextuple mutant obtained by crossing the reference phyB-9 null allele into the jazQ background.
(a) Photograph of five-week-old WT, jazQ and sjq11 plants, the latter of which is a suppressor mutant of jazQ harbouring a null mutation in PHYB. (b) Trichoplusia ni weight after feeding for 10 day on WT (31 larvae), jazQ (31 larvae) and sjq11 (37 larvae) plants. Data show the mean±s.e.m. of at least 12 independent replicates. (c) Photograph of four-week-old plants grown in soil. (d,e) Rosette dry weight and anthocyanin accumulation in petioles, respectively. Data show the mean±s.e.m. of 10 plants per genotype. (f) T. ni weight after feeding for 10 days on WT (23 larvae), jazQ (29 larvae) and jazQ phyB (27 larvae) plants. Data show the mean larval weight±s.e.m. of insects reared on 12 plants per host genotype. Capital letters denote statistical differences according to Tukey HSD-test (P<0.05). Scale bars, 1 cm.
Analysis of growth and defense traits in jazQ phyB plants showed that the jazQ and phyB ‘single’ mutant phenotypes were largely additive and often organ specific. For example, jazQ phyB roots retained the JA-hypersensitive phenotype of jazQ, whereas jazQ phyB hypocotyls showed the red-light insensitive phenotype of phyB (Supplementary Fig. 4). Adult jazQ phyB plants grown in soil resembled phyB in having elongated petioles, flattened rosette leaves28 and early flowering time (Fig. 2c and Supplementary Fig. 5). The phyB mutation is thus epistatic to jazQ for these growth traits. Interestingly, the rosette diameter, projected leaf area and dry mass of jazQ phyB rosette leaves under our growth conditions exceeded that of the jazQ and phyB parents, suggesting that the combination of jazQ and phyB has transgressive effects on leaf growth (Fig. 2d and Supplementary Fig. 5). Despite its robust vegetative growth, jazQ phyB plants maintained the heightened anti-insect defense and anthocyanin content of jazQ (Fig. 2e,f). The effect of combining jazQ and phyB on resistance to T. ni feeding was particularly striking because phyB alone causes high susceptibility to this herbivore (Supplementary Fig. 6). jazQ is thus epistatic to phyB with respect to leaf defense traits. These collective data demonstrate that phyB fully suppresses the diminutive growth stature of jazQ aerial organs but, remarkably, does not compromise the heightened anti-insect resistance that is imparted by jazQ.
Co-expression of MYC and PIF regulons in jazQ phyB
A potential mechanistic explanation for the enhanced growth and defense attributes of jazQ phyB plants comes from recent studies that implicate crosstalk between the JA, phyB and GA signalling pathways in the modulation of growth-defense balance. Within this signalling network, GA stimulates cell extension growth by promoting the degradation of DELLA proteins that repress PIF transcription factors29 (Fig. 1a). Reciprocal antagonism between the JA and GA pathways involves JAZ-DELLA interactions that prevent these repressors from inhibiting their cognate transcription factors19,20,30. JA-GA crosstalk is integrated with the shade avoidance response through phyB-mediated perception of changes in the ratio of red to far red (R:FR) light. Low R:FR ratios indicative of leaf shading reduce phyB activity to relieve repression on PIFs, thereby promoting rapid growth through the concerted action of growth hormones such as auxin and brassinosteroid (Fig. 1a)31,32,33. Concurrent with this growth response to plant competitors, inactivation of phyB by low R:FR (or phyB mutation) is associated with depletion of DELLA proteins, increased JAZ stability, accelerated turnover of MYC transcription factors, and suppression of JA-triggered immune responses5,9,34.
These considerations led us to test the hypothesis that the combination of jazQ and phyB causes concomitant derepression of the MYC and PIF transcriptional programs to promote growth and defense simultaneously. In support of this idea, we found that overexpression of PIF4 in the jazQ background partially rescued the small rosette size and short petiole length of jazQ without affecting anthocyanin accumulation and resistance to T. ni feeding (Supplementary Fig. 7). This finding indicates that increased PIF4-mediated growth does not attenuate the defense stature of jazQ plants. The inability of PIF4 overexpression to fully recapitulate the jazQ phyB phenotype indicates that additional regulatory factors contribute to the growth vigour of jazQ phyB.
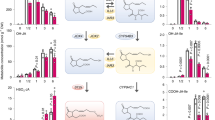
To further test the hypothesis that MYC and PIF transcriptional modules are simultaneously activated in jazQ phyB plants, we used mRNA sequencing to compare the transcript profile of WT, jazQ, phyB, and jazQ phyB seedlings. ‘Secondary metabolism’ and ‘response to stress’ and were among the biological processes most significantly overrepresented in ontologies of 257 genes expressed to higher levels in jazQ than WT (Fig. 3 and Supplementary Data 1). This gene set included glucosinolate biosynthesis genes that are direct targets of MYC2 (ref. 22) (Supplementary Fig. 8), as well as genes involved in the synthesis of triterpenoids, jasmonates, and various defensive proteins (Supplementary Fig. 9). In agreement with their enhanced defense stature, jazQ phyB plants maintained increased expression of the majority (68%) of genes that are upregulated in jazQ (Fig. 3, Supplementary Figs 8 and 9). Analysis of growth-related genes revealed that the set of 235 genes upregulated in both phyB and jazQ phyB is enriched for functional classes involved in responses to auxin, shade avoidance, cell wall organization and light stimulus (Fig. 3). Several genes within this group were previously shown to be direct targets for PIF transcription factor binding33,35,36 (Supplementary Fig. 9). These data provide evidence that the combination of jazQ and phyB promotes concomitant expression of defense and growth-related genes, at least in part, via the concurrent activation of MYC and PIF transcriptional modules.
WT, jazQ, phyB and jazQ phyB seedlings were grown for 8 days in continuous white light before RNA extraction and analysis of gene expression by mRNA sequencing. The Venn diagram shows the number of genes upregulated in comparisons between WT and each of the three mutants. GO analysis of functional categories was performed with gene sets that are shared between jazQ and jazQ phyB (blue intersect), shared between phyB and jazQ phyB (yellow intersect), or unique to jazQ phyB (green shade). See Supplementary Data 1 for detailed expression data and GO analysis.
Among the 576 transcripts whose abundance was significantly increased in jazQ phyB but not jazQ or phyB, there was a strong over-representation of GO terms related to secondary metabolism, cell wall organization, growth and auxin transport (Fig. 3). These data suggest that the combination of jazQ and phyB has synergistic effects on the expression of certain growth and defense responses in jazQ phyB. In support of this idea, quantitative PCR analysis showed that wound-induced expression of select JA-response genes was significantly higher in jazQ phyB than WT leaves, which may contribute to the heightened defense of jazQ phyB plants relative to WT (Supplementary Fig. 10). It is possible that these synergistic effects of jazQ and phyB on gene expression result from functional interaction between MYCs and PIFs at the level of protein–protein interaction or altered binding to common cis-regulatory elements in target genes21,33,34,37. Supporting this hypothesis, PIF4 targets previously identified by ChIP-seq33 include an over-representation of genes associated with the terms ‘jasmonate stimulus’ and ‘response to wounding’ (Supplementary Table 1). Moreover, several PIF4 targets are also targets of MYC2 (Supplementary Table 2). We cannot exclude the possibility that concurrent loss of phyB and JAZ1/3/4/9/10 affects the activity of transcriptional regulators other than PIFs and MYCs to contribute to the gene expression profile and other general phenotypes of jazQ phyB plants.
Genetic modulation of photosynthesis and leaf architecture
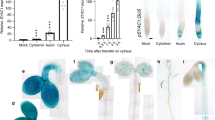
The robust growth of well-defended jazQ phyB leaves led us to investigate whether jazQ and phyB interact to modulate leaf photosynthetic efficiency. Indeed, PIF activity is known to repress chloroplast development and photosynthetic competency37, and we observed that ‘photosynthesis’ was the term most significantly overrepresented among genes that are repressed in both phyB and jazQ phyB seedlings (Supplementary Fig. 11 and Supplementary Data 1). We employed non-invasive, whole-plant chlorophyll fluorescence imaging13,38 to determine how genetic perturbations within the phyB-GA-JA signalling network affect photosystem II efficiency (ΦII) under various light regimes, including those designed to simulate natural environments (Fig. 4a and Supplementary Fig. 12). phyB plants had reduced ΦII under continuous low light intensity and this effect was exacerbated under the sinusoidal and fluctuating light regimes, consistent with previous studies39. A decrease in ΦII was also observed in a Col-0 transgenic line (35S:PIF4) that overexpresses PIF4. Interestingly, the negative effect of phyB and 35S:PIF4 on photosynthetic efficiency was rescued by jazQ, which alone had little (or very weak positive) effect on ΦII (Fig. 4a and Supplementary Fig. 12). Consistent with the role of PIFs in repressing photosynthesis, a pif1/3/4/5 quadruple mutant (pifq) showed increased ΦII under fluctuating light conditions, whereas loss of DELLAs in the della quintuple mutant (dellaQ) reduced ΦII under these conditions. That photosystem II efficiency was lower in phyB than in dellaQ leaves suggests that phyB exerts stronger repression on PIF activity than DELLA proteins under these growth conditions.
(a) Heat map of photosystem II quantum efficiency (ΦII) in response to varying light regimes. Chlorophyll fluorescence values for the indicated mutants were normalized to Col-0. Plants were exposed to three consecutive 16 h day−1 light regimes: constant light (day 1, left); sinusoidal increase and decrease in light intensity (day 2, middle); and sinusoidal light with higher intensity pulses (day 3, right). Statistical analysis of data is shown in Supplementary Fig. 12. (b) Photosynthetic rate in response to increasing light measured by gas exchange in 6–9 plants per genotype. Inset shows nonlinear curve-fitting to model the maximum velocity of Rubisco determined from foliage photosynthetic rates in response to increasing CO2. (c) Rubisco and (d) total chlorophyll concentration in leaves from 54-days-old plants (n=4). (e) Thickness of 22-day-old rosette leaves (n=4). In b–e, data show the mean±s.e.m., and capital letters indicate statistical difference at P<0.05 (Tukey HSD-test). In d, WT and jazQ phyB means are different at P<0.1.
To obtain additional insight into physiological processes that underlie growth-defense vigour of jazQ phyB, we investigated the relationship between photosynthesis and leaf growth to obtain an estimate of leaf construction costs40. Gas exchange experiments showed that phyB leaves have significantly lower photosynthetic rate per unit leaf area, whereas photosynthetic capacity of jazQ on a leaf area or dry weight basis was comparable to WT (Fig. 4b), consistent with our chlorophyll fluorescence measurements. phyB leaves also contained less area-based chlorophyll and Rubisco (D-ribulose-1,5-bisphosphate carboxylase/oxygenase) than WT. Modelling of photosynthetic parameters showed that the reduced photosynthetic capacity of phyB at high light results in part from a limitation in Rubisco activity. jazQ partially rescued the low photosynthetic capacity of phyB leaves, as well as the low area-based Rubisco and chlorophyll content of phyB (Fig. 4c,d). The mechanism by which jazQ partially restores the reduced photosynthetic performance of phyB leaves in unknown but deserves further attention.
We also found that phyB leaves were thinner than those of WT and jazQ and that this trait was retained in jazQ phyB (Fig. 4e). Because of the greater projected leaf area available to intercept light (due to longer petioles and flatter, thinner leaves), the whole-plant photosynthetic rate in jazQ phyB plants was similar to WT. Thus, costs associated with construction of jazQ phyB leaves may be lowered through increased partitioning of carbon to leaf area at the expense of leaf thickness40. These data suggest that changes in leaf architecture rather than increased efficiency of the photosynthetic apparatus may contribute to the growth-defense vigour of jazQ phyB relative to WT plants.
Discussion
A key innovation of our study was the development of a JA signalling mutant (jazQ), in which the removal of multiple JAZ repressors causes hyperactivation of JA responses. As a consequence, jazQ plants exhibit both enhanced resistance to insect herbivory and diminished growth of leaves and roots. The heightened defense of jazQ plants most likely involves increased activity of MYC transcription factors, which are well characterized for their role in promoting secondary metabolism and defense21,22. Our results also provide evidence that transcriptional changes account for the growth phenotypes of jazQ, but the molecular basis of growth restraint remains to be determined. DELLA proteins have previously been implicated in JA-mediated growth inhibition of Arabidopsis roots19 and hypocotyls20. Although these studies raise the possibility that removal of JAZs in jazQ increases the growth-inhibiting activity of DELLAs, previous studies showed that DELLA proteins are not required for wound- and JA-induced growth stunting of leaves12. Given the high connectivity of JAZs within protein interaction networks and their emerging role as integrators of various signalling pathways41,42, it is possible that JAZ-interacting transcriptional regulators other than MYCs and DELLAs also contribute to general phenotypes of jazQ. Further analysis of jazQ should provide new avenues to study how JA controls plant growth and development.
By exploiting jazQ phenotypes in a genetic suppressor screen, we identified jazQ phyB as a novel genotypic combination that uncouples growth-defense tradeoffs mediated by the JA signalling pathway. This major conclusion is based on the finding that phyB suppresses the growth restriction but not enhanced defense of jazQ leaves. Previous studies of phyB-JA crosstalk showed that decreased levels of the active form (Pfr) of phyB, as a consequence of either phyB mutation or exposure to shade light, compromises JA-mediated resistance to a broad spectrum of biotic attackers5,31,32. Recent molecular evidence further indicates that shade-triggered repression of JA-mediated defense in Arabidopsis involves stabilization of JAZs and reduced MYC activity9,34. Given these findings, a plausible mechanistic explanation for why jazQ-mediated leaf defense remains elevated in the presence of phyB is that one or more of the mutated JAZ genes in jazQ (JAZ1/3/4/9/10) are critical for suppressing JA-mediated defenses when levels of active phyB are reduced (for example, in the phyB genetic background). Consistent with this, analysis of the jaz10-1 single mutant has shown that JAZ10 is required for attenuation of JA-mediated defense by shade light9,43,44. Future studies are needed to ascribe specific functions to individual JAZ family members and, more generally, to better understand how light and defense perception pathways are integrated to achieve growth-balance in specific tissues.
A central premise underlying current views of growth-defense balance is that the production of defensive compounds and associated morphological structures is energetically costly. The ability of jazQ phyB leaves to grow and defend well at the same time provides evidence that JA-mediated growth-defense antagonism is not simply a consequence of ‘metabolic competition’ that shunts resources to defense at the expense of growth6,7,45. Our data support an alternative (but not mutually exclusive) model, in which tradeoffs between growth and defense are orchestrated by a hormone-based transcriptional network that is hardwired to restrict growth upon activation of the JA signalling pathway. This interpretation is bolstered by an increasing number of molecular studies that have addressed to role of hormones in the intersection of growth and immunity4,20,46,47. In light of current theories regarding the costs of induced defense45, it remains to be determined whether growth inhibition resulting from genetic removal (for example, jazQ) or natural JA-induced degradation of JAZs provides a fitness benefit to the plant. It is conceivable, for example, that JA-mediated growth restriction enhances the plant’s capacity to anticipate future threats or to cope with multiple-stress conditions48. This hypothesis predicts that simultaneous loss of phyB and JAZ, as major drivers of phenotypic plasticity, may reduce the fitness of jazQ phyB plants in natural environments because these plants lack the regulatory potential to adapt to complex and rapidly changing environments. Support for this idea comes from our RNA-seq data showing that the expression of genes involved in abiotic stress response is repressed in jazQ phyB (Supplementary Fig. 11). An important future direction will thus be to assess the reproductive fitness of these genotypes in the presence of various predators, competitors and abiotic stressors.
The ability to unlock growth-defense tradeoffs through relief of transcription repression may provide an approach to assemble plant traits in ways that have practical application in biotechnology and agronomy. One example concerns the use of exogenous JA to elicit the biosynthesis of commercially valuable plant secondary metabolites49,50. A significant bottleneck in JA-elicited production of these compounds, including the anti-cancer drug paclitaxel (taxol), is the loss of plant biomass resulting from JA-mediated growth inhibition50,51. The design of plants with increased flux through JA-responsive metabolic pathways without concomitant growth restriction could provide a solution to this problem. Similarly, unlinking growth-defense antagonism may be useful in cropping systems that increasingly depend upon high planting densities to maximize yield. Because the ratio of R:FR light perceived by canopy leaves decreases as planting densities increase, densely planted crops may become more susceptible insect pests and pathogens that are sensitive to JA-mediated defenses31,32. In demonstrating that jazQ and phyB can be combined to produce what is essentially a well-defended shade leaf, our results suggest a strategy to cultivate densely planted crops with less dependence on pesticides. The evolutionary conservation of light and defense signalling pathways suggest that the findings described here can readily be applied to most plant species using modern genetic approaches.
Methods
Plant material and growth conditions
A. thaliana Columbia ecotype (Col-0) was used as a WT parent for all experiments. Soil-grown plants were maintained at 20 °C (±1 °C) under 16 h at a light intensity of 120 μE m−2 s−1 and 8 h dark unless otherwise noted. For the first 10 day after seed sowing, trays containing potted plants were covered with a transparent plastic dome to increase humidity. For experiments involving growth of seedlings on agar plates, seeds were surface sterilized for 15 min in a solution containing 50% (v/v) bleach and 0.1% (v/v) Triton X-100, washed 10 times with sterile water and then stratified in dark at 4 °C for 2 days. Seeds were then sown on 0.7% (w/v) agar media containing half-strength Murashige and Skoog (MS; Caisson Labs) salts supplemented with 0.8% (w/v) sucrose. Transfer DNA (T-DNA) insertion mutants used for construction of jazQ were obtained from the Arabidopsis Biological Research Center (ABRC; The Ohio State University) and named as follows: jaz1-2 (JIC-SM.22668), jaz3-4 (GK-097F09), jaz4-1 (SALK_141628), jaz9-4 (GK-265H05) and jaz10-1 (SAIL_92_D08). jaz3-4 and jaz9-4 lines were backcrossed to Col-0 to remove unlinked T-DNA insertions. jaz10-1 was backcrossed to Col-0 to remove a qrt1-2 mutation present in the SAIL lines52. jaz4-1 and jaz10-1 mutants have been described23,53. The jazQ phyB sextuple mutant was obtained from a genetic cross between jazQ and the phyB reference allele phyB-9 (ref. 27). The higher-order pifq (pif1-1/pif3-3/pif4-2/pif5-3) and dellaQ (gai-t6/rgat2/rgl1-1/rgl2-1/rgl3-1) mutants have been described54,55.
PCR analysis
PCR-based genotyping of jazQ and lower-order mutants relied on primer sets flanking T-DNA insertion sites, together with a third primer recognizing the border of the inserted T-DNA. The forward, reverse, and border primers used were the following: JAZ1 (At1g19180), 5′-ACCGAGACACATTCCCGATT-3′, 5′-CATCAGGCTTGCATGCCATT-3′, and 5′-ACGAATAAGAGCGTCCATTTTAGAG-3′; JAZ3 (At3g17860), 5′-ACGGTTCCTCTATGCCTCAAGTC-3′, 5′-GTGGAGTGGTCTAAAGCAACCTTC-3′, and 5′-ATAACGCTGCGGACATCTACATT-3′; JAZ4 (At1g48500), 5′-TCAGGAAGACAGAGTGTTCCC-3′, 5′-TGCGTTTCTCTAAGAACCGAG-3′, and 5′-TTGGGTGATGGTTCACGTAG-3′; JAZ9 (At1g70700), 5′-TACCGCATAATCATGGTCGTC-3′, 5′-TCATGCTCATTGCATTAGTCG-3′, and 5′-CTTTGAAGACGTGGTTGGAACG-3′; JAZ10 (At5g13220), 5′-ATTTCTCGATCGCCGTCGTAGT-3′, 5′-GCCAAAGAGCTTTGGTCTTAGAGTG-3′, and 5′-GTCTAAGCGTCAATTTGTTTACACC-3′. PCR with reverse transcription (RT–PCR) was used to confirm the presence or absence of JAZ transcripts in WT and jazQ plants. For this purpose, RNA was extracted from 8-day-old seedlings grown on MS plates containing 20 μM MeJA. Frozen tissue was homogenized with a mortar and pestle and RNA was extracted using an RNeasy kit (Qiagen) with on-column DNase (Qiagen) treatment. cDNA was reverse transcribed from 1 μg total RNA with a High-Capacity cDNA Reverse Transcription kit (Applied Biosystems, ABI). RT–PCR was performed using primer sets designed to amplify the five JAZ genes and the internal control ACTIN1 (At2g37620). The forward and reverse primer sets used were as follows: JAZ1, 5′-ATGTCGAGTTCTATGGAATGTTCTG-3′ and 5′-TCATATTTCAGCTGCTAAACCGAGCC-3′; JAZ3, 5′-ATGGAGAGAGATTTTCTCGGG-3′ and 5′-TTAGGTTGCAGAGCTGAGAGAAG-3′; JAZ4, 5′-ATGGAGAGAGATTTTCTCGG-3′ and 5′-CAGATGATGAGCTGGAGGAC-3′; JAZ9, 5′-ATGGAAAGAGATTTTCTGGGTTTG-3′ and 5′-TTATGTAGGAGAAGTAGAAGAGTAATTCA-3′; JAZ10, 5′-ATGTCGAAAGCTACCATAGAAC-3′ and 5′-GATAGTAAGGAGATGTTGATACTAATCTCT-3′; and ACTIN1, 5′-ATGGCTGATGGTGAAGACATTCAA-3′ and 5′-TCAGAAGCACTTCCTGTGAACAAT-3′. RT–PCR reactions were performed with the following conditions: 94 °C for 5 min, followed by 30 cycles of denaturation (45 s at 94 °C), annealing (30 s at 52 °C) and elongation (1.5 min at 72 °C). Final elongation step was performed at 72 °C for 10 min and completed reactions were maintained at 12 °C. Forty elongation cycles were used to detect the JAZ4 transcripts, which accumulate at low levels in WT plants56.
Root growth assays
The effect of exogenous JA on seedling root growth inhibition was determined by growing seedlings on square Petri plates (Fisher) containing MS medium supplemented with the indicated concentration of methyl-JA (MeJA; Sigma-Aldrich)57. Plates were incubated vertically in a growth chamber maintained at 21 °C under continuous light for 8 days. Primary root length was measured using the ImageJ software (http://imagej.nih.gov/ij/). WT and mutant lines were grown on the same plate to control for plate-to-plate variation.
Quantification of secondary metabolites
Anthocyanins were quantified using established procedures58, with the following modifications. Petioles were excised from 4-week-old plants and extracted in 1 ml MeOH containing 1% (v/v) HCl. Samples were incubated overnight at 4 °C with constant agitation. Anthocyanin pigments in the resulting extract were measured spectrophotometrically and calculated as A530−0.25(A657) g−1 fresh weight. Glucosinolates were quantified using 8-day-old seedlings grown on solid MS medium59. Seedlings were collected into 2 ml tubes (∼50 seedlings per tube) and immediately frozen in liquid nitrogen. WT and mutant lines were grown on the same plate to avoid plate-to-plate variation. Frozen tissue was lyophilized, ground to a fine powder and extracted with 1 ml 80% MeOH containing an internal standard (25 nmol sinigrin, Sigma-Aldrich). Samples were briefly vortexed, incubated at 75 °C for 15 min, and then centrifuged at 23 °C at 10,000g for 10 min. Resulting supernatants were applied to Sephadex A-25 columns (Amersham). Glucosinolates were released from the columns as desulfoglucosinolates with a solution containing 30 μl of aryl sulfatase (3.0 mg ml−1; Sigma) and 70 μl water (high-performance liquid chromatography grade). Following an overnight incubation in the dark at 23 °C, samples were eluted from the columns with 200 μl 80% MeOH and 200 μl water. Samples were then lyophilized to complete dryness and re-dissolved in 100 μl water. Desulfoglucosinoaltes were detected by high-performance liquid chromatography59. Compound abbreviations in Fig. 1c correspond to the following: 3MSP, 3-methylsulfinylpropylglucosinolate; 4MSB, 4-methylsulfinylbutylglucosinolate; 7MSH, 7-methylsulfinylheptylglucosinolate; 4MTB, 4-methylthiobutylglucosinolate; 8MSO, 8-methylsulfinyloctylglucosinolate; I3M, indol-3-ylmethylglucosinolate; 4MI3M, 4-methoxyindol-3-ylmethylglucosinolate; 1MI3M, 1-methoxyindol-3-ylmethylglucosinolate and 8MTO, 8-methylthiooctylglucosinolate.
Insect-feeding assays
Insect-feeding assays were performed with soil-grown plants maintained in a growth chamber at 19 °C and a photoperiod of 8 h light (120 μE m−2 s−1) and 16 h dark. Neonate Trichoplusia ni larvae (Benzon Research) were transferred to the centre of fully expanded rosette leaves of 6-week-old plants60. Four larvae were reared on each of 12 plants per genotype. Plants were then covered with a transparent dome and returned to the chamber for 10 days, after which larval weights were measured.
Growth and flowering time measurements
Three-to-four week-old soil-grown plants were used for all measurements (typically 10 plants per measurement), unless indicated otherwise. Petiole length of the third true leaf was measured with a caliper after leaf excision. Bolting time was measured in a separate set of plants by counting the number of true leaves on the main stem and the number of days from sowing until bolting (that is, floral buds visible in the centre of the rosette). The same set of plants was subsequently used to assess the length of time to opening of the first flower. Rosette diameter and leaf area were determined by photographing rosettes from the top with a Nikon D80 camera. The resulting images were used to calculate Feret diameter using ImageJ analysis. Total leaf area was determined with GIMP software (http://www.gimp.org). Leaf dry weight was determined by weighing excised rosettes (without roots) after freeze drying for 2 days in a lyophilizer.
jazQ suppressor screen and identification of sjq11
Approximately 50,000 jazQ seeds were mutagenized by immersion in a solution of 0.1% or 0.2% (v/v) EMS (Sigma-Aldrich) for 16 h at room temperature, with constant agitation. Seeds (M1 generation) were thoroughly washed with H2O, stratified in the dark at 4 °C for 2 days and then immediately sown on soil. M2 seed was collected from 16 pools of self-pollinated M1 plants (approximately 1,000 M1 plants/pool). Soil-grown M2 plants (∼2,000 plants/pool) were visually screened for individuals having a larger rosette size than jazQ. Putative sjq (suppressors of the jazQ) mutants were rescreened in the M3 generation to confirm heritability of phenotypes. Insight into the causal mutation in sjq11 came from the observation that sjq11 seedlings grown on MS medium in continuous white light for 3 days have elongated hypocotyls. Subsequent hypocotyl growth assays in monochromatic red light61 confirmed a defect in red-light signalling. Briefly, sjq11 (M3 generation) and control seeds were plated on MS medium lacking sucrose and stratified at 4 °C in dark for 2 days. Mutant and control lines were grown on the same plate to control for plate-to-plate variation. A 3 h pulse of white light was then administered to improve synchronous seed germination. Plates were then returned to darkness for one d at 21 °C and then transferred to a monochromatic light-emitting diode chamber outfitted to emit red light (670±20 nm; 25 μE μE m−2 s−1). As a control, a set of plates containing each genotype was maintained in darkness. Following 3 days of growth, seedling hypocotyls were measured by ImageJ software analysis of scanned images. Allelism tests performed with F1 seedlings (obtained from the cross between sjq11 and phyB-9) revealed a lack of genetic complementation. Finally, sequencing of the PHYB gene (AT2G18790) in sjq11 revealed a C→T transition that introduces a stop codon in a region of the gene that encodes the chromophore-binding domain of phyB.
Gene expression profiling
Global gene expression profiling in 8-day-old whole seedlings (Col-0 WT, jazQ, phyB-9, jazQ phyB-9) was assessed by mRNA sequencing (RNA-seq) performed on the Ilumina HiSeq 2000 platform. Seedlings were grown in continuous light on solid MS medium supplemented with sucrose. For each replicate sample, ∼200 seedlings were harvested for RNA extraction. WT and mutant seedlings were grown on the same plate to minimize plate-to-plate variation. Three independent RNA samples (biological replicates) were sequenced per genotype. Total RNA was isolated as described above and RNA integrity was assessed with a 2100 Bioanalyzer (Agilent Technologies). All samples utilized had an integrity score of at least 7.0. Single-end (50 bp) sequencing was performed at the Michigan State University Research Technologies Service Facility (https://rtsf.natsci.msu.edu). Barcoded sequencing libraries were constructed using the Illumina RNAseq kit according to the manufacturer’s instructions and were multiplexed in six libraries per lane. The average number of sequencing reads was 18.4±4.3 million per sample. Raw sequencing reads were assessed with Illumina quality control tools filters and FASTX toolkit (http://hannonlab.cshl.edu/fastx_toolkit/). Reads were mapped to gene models in TAIR10 with the program RSEM (version 1.2.11) set for default parameters62. Data was expressed as transcripts per million. The average transcripts per million±s.e.m. for all Arabidopsis genes is provided in Supplementary Data 1. DESeq (version 1.18.0) (ref. 63) was used to normalize expected counts from RSEM and to assess differential gene expression by comparing normalized counts in WT to those in a particular mutant. Gene ontology (GO) analysis of enriched functional categories was performed using BiNGO (version 2.44) (ref. 64). The hypergeometric test with Benjamini & Hochberg’s FDR correction was used to calculate over- and underrepresented GO categories among differentially expressed genes, using a P value<0.05.
For wounding experiments, 3-week old soil-grown seedlings were wounded twice across the midvein of four leaves (leaves 3–6, counted from first rosette leaf). After 1 h, the wounded leaves of two plants were pooled and immediately frozen in liquid nitrogen. Equivalent leaves of two unwounded plants were pooled and collected as controls. The experiment was independently replicated twice, with each experiment consisting of 3–4 biological replicates. Frozen tissue was homogenized with a TissueLyser II (Qiagen) and RNA was extracted using an RNeasy kit (Qiagen) with on-column DNase (Qiagen) treatment, as per the manufacturer's instructions. RNA quality was assessed by A260/A280 ratios using a ND-1000 UV Nanodrop spectrophotometer (Thermo Scientific). cDNA was reverse transcribed using a High-Capacity cDNA Reverse Transcription kit (Applied Biosystems, ABI), as per the manufacturer's instructions, and cDNA was diluted to 0.5 ng μl−1 with nuclease-free water. Quantitative real-time PCR reactions consisted of 5 μl of 2 × Power SYBR Green (ABI) master mix, 2 ul diluted cDNA template (1 ng total), 1 μl 5 uM forward and reverse primers, and nuclease-free water for 10 μl total reaction volume. The forward and reverse primers used were the following: PP2A, 5′-AAGCAGCGTAATCGGTAGG-3′ and 5′-GCACAGCAATCGGGTATAAAG-3′; AOS, 5′-GGAGAACTCACGATGGGAGCGATT-3′ and 5′-GCGTCGTGGCTTTCGATAACCAGA-3′; LOX3, 5′-GCTGGCGGTTCGACATG-3′ and 5′-GCCATTCCTCTGCGAATTAGA-3′; and MYC2, 5′-AGAAACTCCAAATCAAGAACCAGCTC-3′ and 5′-CCGGTTTAATCGAAGAACACGAAGAC-3′. Reactions were run on an ABI 7500 Fast qPCR instrument with the following conditions: 95 °C for 10 min, then 40 cycles of 15 s at 95 °C for denaturation and 60 s at 60 °C for annealing and polymerization. A dissociation curve was performed at the end of each reaction to confirm primer specificity using default parameters (15 s at 95 °C, 60 s at 60 °C–95 °C in 1 °C increments, and 15 s at 95 °C). Target gene expression was normalized to the expression of PP2a, which is stable under JA-inducing conditions13. The normalization incorporated primer efficiencies determined for each primer pair using LinRegPCR v2012.0 (ref. 65) from the log-linear phase of each amplification plot.
Overexpression of PIF4 in the jazQ background
The 35S::PIF4-TAP overexpression construct66 was kindly provided by Dr Michael Thomashow (Michigan State University). Transformation of jazQ plants with Agrobacterium tumefaciens (strain C58C1) was performed using the flower dip method67. Multiple independent transformed lines (T1 generation) were selected on MS plates containing gentamycin and transferred to soil for subsequent analysis. Homozygous lines were selected by testing the T3 progeny for gentamycin resistance.
Photosynthesis measurements
Gas exchange measurements were performed with plants grown in plastic containers (‘Cone-tainers’, Steuwe and Sons, Tangent, OR, USA) on an 8 h light (19 °C)/ 16 h dark (16 °C) photoperiod and 120 μmol m−2 s−1 photosynthetic photon flux density (PPFD)40,68. Single mature rosette leaves (attached) from 8- to 10-week-old plants were used to obtain CO2 response curves on a LI-6400XT system (LI-COR Biosciences, Lincoln, NE, USA) outfitted with a standard leaf chamber (chamber area=6 cm2). Leaves were supplied with an artificial air mixture consisting of 20% O2, 80% N2, and 400 ppm CO2 at intensity of light 500 μmol m−2 s−1. Leaf temperature was maintained at ∼20 °C (block temperature set to 18 °C). Leaves were acclimated under this condition for at least 30 min before the start of each experiment. Assimilation rates were normalized to projected leaf area as measured by image analysis with the GIMP software. Area- and whole-plant-based photosynthesis and respiration was determined at four time points of the Arabidopsis growth cycle using plants grown under short-day conditions40.
In situ chlorophyll fluorescence measurements were performed in a Percival AR41L2 (Geneva Scientific, http://www.geneva-scientific.com) refitted as a Dynamic Environment Photosynthesis Imager (DEPI) system13,38. Images were processed using visual phenomics software69. The quantum yield of PSII (ΦII) was calculated as (F’M−FS)/F’M, where FS is the steady-state fluorescence and F’M is the fluorescence maximum at steady state. Statistical analysis of the ΦII data is provided in Supplementary Fig. 12.
Leaf thickness measurements
Leaf cross sections obtained from the 5th leaf of 22-day old rosette leaves were examined under an Olympus FluoView FV1000, Confocal Laser Scanning Microscope (Olympus, NJ, USA) in the Center for Advanced Microscopy, Michigan State University. Leaf thickness was measured as the distance between the abaxial and adaxial surfaces of the leaf40.
Measurement of total chlorophyll and Rubisco
Chlorophyll was extracted from 54-d old Arabidopsis rosette leaves with 96% ethanol. Absorbance of the extracted chlorophyll was measured spectrophotometrically at 665 nm and 649 nm and the total chlorophyll was calculated using the following equation: Chla+Chlb=(13.95A665–6.88A649)+(24.96A649–7.32A665)70. Total protein was extracted from 54-d old Arabidopsis rosette leaves using a Plant Total Protein Extraction Kit (Sigma-Aldrich, MO, USA). A modified Lowry Assay was performed to measure the total protein concentration in the extract and the purity and quality of the extracted protein were determined by denaturing polyacrylamide gel electrophoresis. Equal amounts of total protein were loaded onto an automated capillary-based size western blotting system (ProteinSimple Wes System, San Jose CA, USA). All procedures were performed with manufacturer’s reagents according to their user manual. Protein separation and immunodetection were performed automatically on the individual capillaries using the default settings. Antibodies raised against the large subunit of Rubisco (rabbit, AS03 037; Agrisera, Sweden; dilution used 1:650) were used to detect Rubisco in each protein sample. For quantification, all subsequent data generated was analysed with Compass Software provided by manufacturer (ProteinSimple, San Jose CA). Peak heights of the fluorescence signals were used to calculate relative differences of Rubisco concentration between samples. Rubisco concentration per unit leaf area was calculated based on the total protein concentration and measurements of leaf area per unit mass.
Data Availability
RNA sequencing data is deposited at the National Center for Biotechnology Information Gene Expression Omnibus (GEO) as series record GSE79012. The authors declare that all other data supporting the findings of this study are available within the article and its Supplementary Information files or are available from the corresponding author upon request.
Additional information
How to cite this article: Campos, M. L. et al. Rewiring of jasmonate and phytochrome B signalling uncouples plant growth-defense tradeoffs. Nat. Commun. 7:12570 doi: 10.1038/ncomms12570 (2016).
References
Stamp, N. Out of the quagmire of plant defense hypotheses. Q. Rev. Biol. 78, 23–55 (2003).
Zust, T., Rasmann, S. & Agrawal, A. A. Growth-defense tradeoffs for two major anti-herbivore traits of the common milkweed Asclepias syriaca. Oikos 124, 1404–1415 (2015).
Herms, D. A. & Mattson, W. J. The dilemma of plants - to grow or defend. Q. Rev. Biol. 67, 283–335 (1992).
Huot, B., Yao, J., Montgomery, B. L. & He, S. Y. Growth-defense tradeoffs in plants: a balancing act to optimize fitness. Mol. Plant 7, 1267–1287 (2014).
Moreno, J. E., Tao, Y., Chory, J. & Ballare, C. L. Ecological modulation of plant defense via phytochrome control of jasmonate sensitivity. Proc. Natl Acad. Sci. USA 106, 4935–4940 (2009).
Havko, N. E. et al. Control of carbon assimilation and partitioning by jasmonate: an accounting of growth-defense balance. Plants 5, 7 (2016).
Kliebenstein, D. J. False idolatry of the mythical growth versus immunity tradeoff in molecular systems plant pathology. Physiol. Mol. Plant Pathol. 95, 55–59 (2016).
Hu, P. et al. JAV1 controls jasmonate-regulated plant defense. Mol. Cell 50, 504–515 (2013).
Leone, M., Keller, M. M., Cerrudo, I. & Ballare, C. L. To grow or defend? Low red:far-red ratios reduce jasmonate sensitivity in Arabidopsis seedlings by promoting DELLA degradation and increasing JAZ10 stability. New Phytol. 204, 355–367 (2014).
Schuman, M. C. & Baldwin, I. T. The layers of plant responses to insect herbivores. Annu. Rev. Entomol. 61, 373–394 (2016).
Yan, Y. et al. A downstream mediator in the growth repression limb of the jasmonate pathway. Plant Cell 19, 2470–2483 (2007).
Zhang, Y. & Turner, J. Wound-induced endogenous jasmonates stunt plant growth by inhibiting mitosis. PLoS ONE 3, e3699 (2008).
Attaran, E. et al. Temporal dynamics of growth and photosynthesis suppression in response to jasmonate signaling. Plant Physiol. 165, 1302–1314 (2014).
Gasperini, D. et al. Multilayered organization of jasmonate signalling in the regulation of root growth. PLoS Genet. 11, e1005300 (2015).
Howe, G. & Jander, G. Plant immunity to insect herbivores. Annu. Rev. Plant. Biol. 59, 41–66 (2008).
Thireault, C. et al. Repression of jasmonate signaling by a non-TIFY JAZ protein in Arabidopsis. Plant J. 82, 669–679 (2015).
Thines, B. et al. JAZ repressor proteins are targets of the SCFCOI1 complex during jasmonate signalling. Nature 448, 661–665 (2007).
Chini, A. et al. The JAZ family of repressors is the missing link in jasmonate signalling. Nature 448, 666–671 (2007).
Hou, X., Lee, L. Y., Xia, K., Yan, Y. & Yu, H. DELLAs modulate jasmonate signaling via competitive binding to JAZs. Dev. Cell 19, 884–894 (2010).
Yang, D. L. et al. Plant hormone jasmonate prioritizes defense over growth by interfering with gibberellin signaling cascade. Proc. Natl Acad. Sci. USA 109, 1192–1200 (2012).
Fernandez-Calvo, P. et al. The Arabidopsis bHLH Transcription Factors MYC3 and MYC4 are targets of JAZ repressors and act additively with MYC2 in the activation of jasmonate responses. Plant Cell 23, 701–715 (2011).
Schweizer, F. et al. Arabidopsis basic helix-loop-helix transcription factors MYC2, MYC3, and MYC4 regulate glucosinolate biosynthesis, insect performance, and feeding behavior. Plant Cell 25, 3117–3132 (2013).
Sehr, E. M. et al. Analysis of secondary growth in the Arabidopsis shoot reveals a positive role of jasmonate signalling in cambium formation. Plant J. 63, 811–822 (2010).
Demianski, A. J., Chung, K. M. & Kunkel, B. N. Analysis of Arabidopsis JAZ gene expression during Pseudomonas syringae pathogenesis. Mol. Plant Pathol. 13, 46–57 (2012).
Acosta, I. F. et al. Role of NINJA in root jasmonate signaling. Proc. Natl Acad. Sci. USA 110, 15473–15478 (2013).
Qi, T. et al. The Jasmonate-ZIM-domain proteins interact with the WD-Repeat/bHLH/MYB complexes to regulate jasmonate-mediated anthocyanin accumulation and trichome initiation in Arabidopsis thaliana. Plant Cell 23, 1795–1814 (2011).
Reed, J. W., Nagpal, P., Poole, D. S., Furuya, M. & Chory, J. Mutations in the gene for the red/far-red light receptor phytochrome B alter cell elongation and physiological responses throughout Arabidopsis development. Plant Cell 5, 147–157 (1993).
Kozuka, T., Suetsugu, N., Wada, M. & Nagatani, A. Antagonistic regulation of leaf flattening by phytochrome B and phototropin in Arabidopsis thaliana. Plant Cell Physiol. 54, 69–79 (2013).
de Lucas, M. et al. A molecular framework for light and gibberellin control of cell elongation. Nature 451, 480–484 (2008).
Wild, M. et al. The Arabidopsis DELLA RGA-LIKE3 is a direct target of MYC2 and modulates jasmonate signaling responses. Plant Cell 24, 3307–3319 (2012).
Casal, J. J. Photoreceptor signaling networks in plant responses to shade. Annu. Rev. Plant Biol. 64, 403–427 (2013).
Ballare, C. L. Light regulation of plant defense. Annu. Rev. Plant Biol. 65, 335–363 (2014).
Oh, E., Zhu, J. Y. & Wang, Z. Y. Interaction between BZR1 and PIF4 integrates brassinosteroid and environmental responses. Nat. Cell Biol. 14, 802–809 (2012).
Chico, J. M. et al. Repression of jasmonate-dependent defenses by shade involves differential regulation of protein stability of MYC transcription factors and their JAZ repressors in Arabidopsis. Plant Cell 26, 1967–1980 (2014).
Hornitschek, P. et al. Phytochrome interacting factors 4 and 5 control seedling growth in changing light conditions by directly controlling auxin signaling. Plant J. 71, 699–711 (2012).
Zhang, Y. et al. A quartet of PIF bHLH factors provides a transcriptionally centered signaling hub that regulates seedling morphogenesis through differential expression-patterning of shared target genes in Arabidopsis. PLoS Genet. 9, e1003244 (2013).
Leivar, P. & Monte, E. PIFs: systems integrators in plant development. Plant Cell 26, 56–78 (2014).
Cruz, J. A. et al. Dynamic environmental photosynthetic imaging reveals emergent phenotypes. Cell Syst. 2, 365–377 (2016).
Boccalandro, H. E. et al. Phytochrome B enhances photosynthesis at the expense of water-use efficiency in Arabidopsis. Plant Physiol. 150, 1083–1092 (2009).
Weraduwage, S. M. et al. The relationship between leaf area growth and biomass accumulation in Arabidopsis thaliana. Front. Plant Sci. 6, 167 (2015).
Wager, A. & Browse, J. Social network: JAZ protein interactions expand our knowledge of jasmonate signaling. Front. Plant Sci. 3,, doi: doi:10.3389/fpls.2012.00041 (2012).
Campos, M. L., Kang, J. H. & Howe, G. A. Jasmonate-triggered plant immunity. J. Chem. Ecol. 40, 657–675 (2014).
Cerrudo, I. et al. Low red/far-red ratios reduce Arabidopsis resistance to Botrytis cinerea and jasmonate responses via a COI1-JAZ10-dependent, salicylic acid-independent mechanism. Plant Physiol. 158, 2042–2052 (2012).
Cargnel, M. D., Demkura, P. V. & Ballare, C. L. Linking phytochrome to plant immunity: low red: far-red ratios increase Arabidopsis susceptibility to Botrytis cinerea by reducing the biosynthesis of indolic glucosinolates and camalexin. New Phytol. 204, 342–354 (2014).
Heil, M. & Baldwin, I. T. Fitness costs of induced resistance: emerging experimental support for a slippery concept. Trends Plant Sci. 7, 61–67 (2002).
Lozano-Duran, R. & Zipfel, C. Trade-off between growth and immunity: role of brassinosteroids. Trends Plant Sci. 20, 12–19 (2015).
Wang, W. F. & Wang, Z. Y. At the intersection of plant growth and immunity. Cell Host Microbe 15, 401–403 (2014).
Coolen, S. et al. Transcriptome dynamics of Arabidopsis during sequential biotic and abiotic stresses. Plant J. 86, 249–267 (2016).
Geyter, D., Gholami, A. & Goossens, A. Transcriptional machineries in jasmonate-elicited plant secondary metabolism. Trend Plant Sci. 17, 349–359 (2012).
Ramirez-Estrada, K. et al. Elicitation, an Effective strategy for the biotechnological production of bioactive high-added value compounds in plant cell factories. Molecules 21, 182 (2016).
Yukimune, Y., Tabata, H., Higashi, Y. & Hara, Y. Methyl jasmonate-induced overproduction of paclitaxel and baccatin III in Taxus cell suspension cultures. Nat. Biotechnol. 14, 1129–1132 (1996).
McElver, J. et al. Insertional mutagenesis of genes required for seed development in Arabidopsis thaliana. Genetics 159, 1751–1763 (2001).
Jiang, Y., Liang, G., Yang, S. & Yu, D. Arabidopsis WRKY57 functions as a node of convergence for jasmonic acid- and auxin-mediated signaling in jasmonic acid-induced leaf senescence. Plant Cell 26, 230–245 (2014).
Feng, S. H. et al. Coordinated regulation of Arabidopsis thaliana development by light and gibberellins. Nature 451, 475–479 (2008).
Leivar, P. et al. Multiple phytochrome-interacting bHLH transcription factors repress premature seedling photomorphogenesis in darkness. Curr. Biol. 18, 1815–1823 (2008).
Chung, H. S. et al. Regulation and function of Arabidopsis JASMONATE ZIM-domain genes in response to wounding and herbivory. Plant Physiol. 146, 952–964 (2008).
Shyu, C. et al. JAZ8 lacks a canonical degron and has an EAR motif that mediates transcriptional repression of jasmonate responses in Arabidopsis. Plant Cell 24, 536–550 (2012).
Kang, J. H. et al. The flavonoid biosynthetic enzyme chalcone isomerase modulates terpenoid production in glandular trichomes of tomato. Plant Physiol. 164, 1161–1174 (2014).
Barth, C. & Jander, G. Arabidopsis myrosinases TGG1 and TGG2 have redundant function in glucosinolate breakdown and insect defense. Plant J. 46, 549–562 (2006).
Herde, M., Koo, A. J. & Howe, G. A. Elicitation of jasmonate-mediated defense responses by mechanical wounding and insect herbivory. Methods Mol. Biol. 1011, 51–61 (2013).
Warnasooriya, S. N. & Montgomery, B. L. Detection of spatial-specific phytochrome responses using targeted expression of biliverdin reductase in Arabidopsis. Plant Physiol. 149, 424–433 (2009).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Maere, S., Heymans, K. & Kuiper, M. BiNGO: a Cytoscape plugin to assess overrepresentation of Gene Ontology categories in biological networks. Bioinformatics 21, 3448–3449 (2005).
Ruijter, J. M. et al. Amplification efficiency: linking baseline and bias in the analysis of quantitative PCR data. Nucleic Acids Res. 37, e45 (2009).
Lee, C. M. & Thomashow, M. F. Photoperiodic regulation of the C-repeat binding factor (CBF) cold acclimation pathway and freezing tolerance in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 109, 15054–15059 (2012).
Clough, S. J. & Bent, A. F. Floral Dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Li, Z. R., Gao, J. P., Benning, C. & Sharkey, T. D. Characterization of photosynthesis in Arabidopsis ER-to-plastid lipid trafficking mutants. Photosynth. Res. 112, 49–61 (2012).
Tessmer, O. L., Jiao, Y., Cruz, J. A., Kramer, D. M. & Chen, J. Functional approach to high-throughput plant growth analysis. BMC Syst. Biol. 7, (Suppl 6): S17 (2013).
Lichtenthaler, H. K. & Wellburn, A. R. Determinations of total carotenoids and chlorophylls a and b of leaf extracts in different solvents. Biochem. Soc. Trans. 11, 591–592 (1983).
Acknowledgements
We thank Drs Carlos Ballare and Javier Moreno for critical comments on the manuscript. We also acknowledge Fransisca Anozie for assistance with growth data analysis, Sean Weise for assistance with photosynthesis measurements, Alicia Withrow and Melinda Frame in the MSU Center for Advanced Microscopy for help with leaf thickness measurements, Beronda Montgomery for use of monochromatic light chambers, and Linda Savage for technical assistance in the Michigan State University Center for Advanced Algal and Plant Phenotyping. This work was primarily funded by the Chemical Sciences, Geosciences and Biosciences Division, Office of Basic Energy Sciences, Office of Science, US Department of Energy through grant no. DE–FG02–91ER20021. Construction of mutant lines was supported by the National Institutes of Health award number GM57795 to G.A.H. Partial support for M.L.C. was provided by a MSU College of Natural Science Dissertation Completion Fellowship and the MSU Discretionary Funding Initiative. Y.Y. was supported in part by a postdoctoral fellowship from the Japan Society for the Promotion of Science. D.d.O.F. was supported by a fellowship from the Brazilian National Council for Scientific and Technological Development (Science Without Borders–CNPq). G.J. acknowledges support from NSF award IOS-1139329 and G.A.H. acknowledges support from the Michigan AgBioResearch Project MICL02278.
Author information
Authors and Affiliations
Contributions
M.L.C., Y.Y. and G.A.H. were responsible for the conception and planning of the project. M.L.C. performed all growth and defense phenotypic analysis of adult plants and was responsible for photography of whole plants. Y.Y. generated all Arabidopsis mutant and transgenic lines used in the work and performed phenotypic analysis of mutants at the seedling stage, including light response assays. M.L.C. and D.d.O.F. performed the genetic suppressor screen and characterized sjq11 with assistance from Y.Y. G.J. and M.L.C. designed and performed experiments for quantification of glucosinolates. M.L.C. harvested samples for RNA-seq analysis and analysed the resulting data. I.T.M. and T.D.S. designed and performed photosynthetic measurements, and S.M.W. and T.D.S. performed leaf-area growth modelling, leaf thickness measurements and chlorophyll quantification. D.M.K. designed chlorophyll fluorescence experiments. J.E.F., B.F.J., S.M.W. and T.D.S. quantified leaf Rubisco protein levels. The manuscript was primarily written by M.L.C. and G.A.H. with input from all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-12, Supplementary Tables 1-2 and Supplementary References (PDF 6486 kb)
Supplementary Data 1
Full dataset comprising RNAseq and gene onthology (GO) analysis performed on WT, jazQ, phyB and jazQ phyB seedlings (XLSX 4640 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Campos, M., Yoshida, Y., Major, I. et al. Rewiring of jasmonate and phytochrome B signalling uncouples plant growth-defense tradeoffs. Nat Commun 7, 12570 (2016). https://doi.org/10.1038/ncomms12570
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms12570
This article is cited by
-
Jasmonate signaling drives defense responses against Alternaria alternata in chrysanthemum
BMC Genomics (2023)
-
Subtype-selective agonists of plant hormone co-receptor COI1-JAZs identified from the stereoisomers of coronatine
Communications Biology (2023)
-
Plant susceptibility to a shared herbivore is reduced by belowground competition with neighbors
Oecologia (2023)
-
Developmental Phytohormones: Key Players in Host-Microbe Interactions
Journal of Plant Growth Regulation (2023)
-
Transcriptomics analysis reveals Xanthoceras sorbifolia Bunge leaves’ adaptation strategy to low nitrogen
Trees (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.