Abstract
The breast cancer susceptibility gene BRCA1 is well known for its function in double-strand break (DSB) DNA repair. While BRCA1 is also implicated in transcriptional regulation, the physiological significance remains unclear. COBRA1 (also known as NELF-B) is a BRCA1-binding protein that regulates RNA polymerase II (RNAPII) pausing and transcription elongation. Here we interrogate functional interaction between BRCA1 and COBRA1 during mouse mammary gland development. Tissue-specific deletion of Cobra1 reduces mammary epithelial compartments and blocks ductal morphogenesis, alveologenesis and lactogenesis, demonstrating a pivotal role of COBRA1 in adult tissue development. Remarkably, these developmental deficiencies due to Cobra1 knockout are largely rescued by additional loss of full-length Brca1. Furthermore, Brca1/Cobra1 double knockout restores developmental transcription at puberty, alters luminal epithelial homoeostasis, yet remains deficient in homologous recombination-based DSB repair. Thus our genetic suppression analysis uncovers a previously unappreciated, DNA repair-independent function of BRCA1 in antagonizing COBRA1-dependent transcription programme during mammary gland development.
Similar content being viewed by others
Introduction
Germline mutations in BRCA1 predispose women to breast and ovarian cancers1. BRCA1 is best known for its role in promoting the homologous recombination (HR)-based pathway of DNA double-strand break (DSB) repair2. In addition to DSB repair, BRCA1 has also been implicated in other cellular processes, including transcription3,4. BRCA1 binds to RNA polymerase II (RNAPII)5 and various site-specific transcription factors, including oestrogen receptor α (ERα) and GATA3 (refs 6, 7), which are involved in mammary gland development and breast cancer. Our previous work indicates that BRCA1 is capable of regulating transcription through high-order chromatin reorganization and unfolding8,9,10,11,12. Consistent with a role for BRCA1 in transcriptional regulation, genome-wide analysis indicates that chromatin binding of BRCA1 is enriched at transcription start sites (TSS) across the human genome13,14,15. Notwithstanding these in vitro findings, there is a gap of knowledge concerning the physiological relevance of these transcription-related activities of BRCA1 in vivo.
We previously identified a BRCA1-binding protein, cofactor of BRCA1 or COBRA1 (ref. 11), which is identical to the B subunit of the negative elongation factor (NELF) complex16. NELF is a metazoan-specific regulator of transcription elongation that pauses RNAPII at the TSS-proximal region17,18. Although NELF was first identified as a transcription elongation repressor in vitro16, subsequent in vivo studies indicate that NELF-mediated RNAPII pausing can lead to both decreased and increased transcription18,19,20,21. In ERα+ breast cancer cells, COBRA1 interacts with ERα (ref. 22) and regulates RNAPII movement at ERα target genes23,24. While NELF-mediated RNAPII pausing has been proposed to ensure synchronous transcriptional activation of developmentally regulated genes25, the exact physiological roles of mammalian NELF have just begun to be deciphered. Published work from us and others indicates that mouse COBRA1 is critical for early embryogenesis26,27 and energy homoeostasis in adult myocardium28. However, the role of COBRA1 in adult tissue development remains unexplored. Furthermore, it is not clear whether the previously characterized physical interaction between COBRA1 and BRCA1 has any physiological significance. Using mammary epithelium-specific knockout (KO) mouse models for Brca1 and Cobra1, we provide compelling genetic evidence for a previously unrecognized functional link between BRCA1 and a transcription elongation factor in dictating the developmental outcome in mammary epithelium.
Results
Brca1 and Cobra1 complementation in ductal development
To investigate the role of COBRA1 in mammary gland development, we generated mammary epithelium-specific KO mice by breeding the MMTV-Cre strain29 with Cobra1f/f animals26, which resulted in deletion of the first four Cobra1 exons. Mammary epithelium of the resulting female MMTV-Cre,Cobra1f/f (CKO) animals was effectively depleted of COBRA1 (Supplementary Fig. 1a,b). Compared with age-matched wild-type littermates (WT, Cobra1f/f) and hemizygous mice (MMTV-Cre,Cobra1f/+), virgin CKO with homozygous deletion of Cobra1 displayed severely retarded mammary ductal growth (Fig. 1a,b; Supplementary Fig. 2a). The normal developmental phenotype of hemizygous Cobra1 knockout mice (MMTV-Cre,Cobra1f/+, Supplementary Fig. 2a) strongly indicates that the CKO-associated developmental defects are due to homozygous Cobra1 ablation, not Cre recombinase expression per se. We therefore used littermates carrying floxed WT gene alleles as the control for most of our subsequent experiments.
(a) Whole mounts of mammary glands from 8-wk virgin mice. The boundary of the ductal area is highlighted. Scale bars, 1 mm. Images are representatives of at least 6 animals. (b) Measurement of the average ductal length at four developmental time points. The numbers of animals used for each of the four time points (6, 8, 12 and 24 wks) are: WT (4, 7, 7 and 12 mice), BKO (3, 3, 4 and 4 mice), CKO (3, 6, 5 and 8 mice) and DKO (4, 7, 4 and 4 mice). Statistical analyses here and in c were conducted using Student’s t-test for comparison between CKO and WT, and between DKO and CKO. (c) Flow cytometry analysis of total live cells in various subpopulations per mammary gland from 16-wk virgin mice. Stromal cells: CD49f-EpCAM−, luminal epithelial cells: CD49fmedEpCAMhigh, myoepithelial cells: CD49fhighEpCAMmed. The numbers of animals used are: WT (4), BKO (3), CKO (3) and DKO (4). *P<0.05, **P<0.01 by Student’s t-test. Error bars represent s.e.m. wks, weeks.
The developmental defect of CKO was most profound during and shortly after puberty (6 and 8 weeks), and remained significant in older virgin mice (12 and 24 weeks, Fig. 1b). In further support, flow cytometry using established cell-surface markers for mammary epithelial cells30 showed that total live luminal (CD49fmedEpCAMhigh) and myoepithelial (CD49fhighEpCAMmed) cell populations of CKO mammary glands were equally reduced compared with their WT littermates (Fig. 1c; Supplementary Fig. 3), suggesting an overall developmental arrest of multiple mammary epithelial lineages. These data clearly indicate that mouse COBRA1 is important for adult tissue development.
Given the physical interaction between BRCA1 and COBRA1 (ref. 11), we compared the phenotypes of CKO with MMTV-Cre-mediated Brca1 KO that was conditionally deleted of Brca1 exon 11 (MMTV-Cre,Brca1f/f; BKO), and Brca1/Cobra1 double KO mice (MMTV-Cre,Brca1f/f,Cobra1f/f; DKO). Consistent with published findings31,32, BKO animals exhibited grossly normal ductal growth at puberty (Fig. 1a,b). Ductal development of DKO mice was stunted at 6 weeks (Fig. 1b; Supplementary Fig. 4), but remarkably, it approached that of WT and BKO at later stages (Fig. 1a,b). Furthermore, abundance of luminal and myoepithelial cells in DKO mammary glands was significantly higher than that in age-matched CKO (Fig. 1c). While myoepithelial cell number in DKO was still lower than that in WT (P=0.026 by Student’s t-test), there was no appreciable difference in luminal cell abundance between DKO and WT. We confirmed that COBRA1 and BRCA1 expression in DKO mice were depleted to a similar extent versus the corresponding single-gene KO animals (Supplementary Fig. 1b,c). Therefore the marked phenotypic difference between CKO and DKO reflects a bona fide genetic complementation between Brca1 and Cobra1. We interpret our finding to mean that BRCA1 imposes a developmental blockade on CKO mammary glands.
Rescue of alveologenesis and lactogenesis in DKO
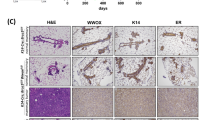
Despite the partial ductal growth in older virgin CKO (Fig. 1b), all pups of CKO dams died shortly after birth from obvious lack of nursing, suggesting a profound and persistent functional defect rather than transient ductal growth delay in CKO mammary glands. In support, mammary glands of CKO at postpartum were largely devoid of alveolar structure (Fig. 2a–c) and milk proteins (Fig. 2d). In contrast to CKO, hemizygous Cobra1 knockout dams (MMTV-Cre,Cobra1f/+) displayed alveologenesis (Supplementary Fig. 2b) and nursing ability comparable to their WT littermates (Cobra1f/f). Thus, we conclude that COBRA1 is required for normal mammary gland functions.
(a,b) Whole mounts of mammary glands from 16 to 20-week mice 1-day postpartum. Scale bar, 1 mm in a and 500 μm in b. (c) H&E stain of the lobular-alveolar structure in mammary glands of 16–20-week mice 1-day postpartum. Scale bar, 100 μm. (d) Immunohistochemistry for total milk proteins in mammary glands of 16–20-week mice 1-day postpartum. Scale bar, 50 μm. Images in this figure are representatives of at least 4 animals in each genotype.
Consistent with published reports31,32, we observed alveologenic and lactogenic defects in BKO mammary glands, albeit much milder than CKO (Fig. 2). In stark contrast, DKO dams with simultaneous deletion of Brca1 and Cobra1 underwent efficient alveologenesis and lactogenesis, as evidenced by the normal alveolar structure (Fig. 2a–c) and abundant milk proteins (Fig. 2d). Collectively, these genetic data unequivocally demonstrate a functional interaction between Brca1 and Cobra1 in mammary gland development and function.
Gene-specific genetic interaction between Brca1 and Cobra1
To determine how specific the genetic complementation is between Brca1 and Cobra1, we first asked whether genetic ablation of other growth-arresting tumour suppressor genes could rescue the developmental defects associated with CKO. Tumour suppressor genes Ink4-Arf play a critical role in oncogene-induced senescence, and co-deletion of the Ink4a/Arf locus restored developmental defect associated with the loss of Bmi1, which encodes a transcriptional regulator of stem cell renewal33. Likewise, deletion of tumour suppressor gene Trp53 partially rescued early embryonic lethality associated with Brca1 deficiency34,35,36. We therefore combined CKO with whole-body deletion of Ink4-Arf or mammary gland-specific deletion of Trp53. In contrast to Brca1 deletion, neither Ink4-Arf nor Trp53 deficiency rescued the ductal growth defect of virgin CKO (Supplementary Figs 5 and 6), suggesting a gene-specific genetic interaction between Brca1 and Cobra1. In a separate experiment, we did not find any sign of mutual rescue of embryonic lethality associated with Brca1 or Cobra1 deletion (Supplementary Table 1). Taken together, these findings underscore the specificity of the genetic interaction between Cobra1 and Brca1 in mammary epithelium.
DKO exhibits altered epithelial homoeostasis
While DKO had functionally restored mammary glands, its epithelial layers were appreciably thickened when compared with both WT and the single-gene KOs (Fig. 3a). This was largely due to an increased population of epithelial cells that expressed Keratin 8 (K8), an established luminal marker (Fig. 3b). In contrast, DKO still maintained a single layer of Keratin 14 (K14)-positive myoepithelial cells (Fig. 3b). However, we reproducibly observed a small number of K14-positive cells uniquely residing in the luminal epithelial compartment of DKO mammary ducts (arrows in Fig. 3b). Bromodeoxyuridine (BrdU) pulse-labelling and TUNEL analysis indicate both elevated cell proliferation and apoptosis rates, respectively, in DKO mammary epithelium (Supplementary Fig. 7), indicating an increased epithelial turnover. Thus, despite developmental and functional rescue, DKO mammary glands exhibit distinct epithelial cell dynamics and organization.
(a) H&E staining of mammary ducts from 8-week WT and KO animals. Scale bar, 50 μm. (b) Immunofluorescence staining with luminal epithelial and myoepithelial markers K8 and K14, respectively, from 8-week animals. Scale bar, 50 μm. (c) Enumeration of mature luminal (CD49fmedEpCAMhighCD49b−) and progenitor cells (CD49fmedEpCAMhighCD49b+). The numbers of animals used are: WT (4), BKO (3), CKO (3) and DKO (4). *P<0.05, **P<0.01 by Student’s t-test. Error bars represent s.e.m.
To assess further tissue homoeostasis in DKO mammary glands, we conducted flow cytometry to distinguish luminal progenitor cells from mature luminal cells in 8-week-old virgin animals after puberty, using CD49b as an established luminal progenitor marker30. Consistent with published findings37,38,39, we found that BKO had more luminal progenitor cells versus WT animals (Fig. 3c). In contrast, CKO mammary glands contained reduced pools of both mature (CD49b−) and progenitor (CD49b+) cells in the luminal epithelial compartment (Fig. 3c), again suggesting inhibition of mammary epithelial cells of all lineages and differentiation stages upon Cobra1 ablation. Intriguingly, the flow cytometry profiles of DKO were distinct from those of BKO and CKO. In particular, the luminal progenitor cell population in DKO was substantially smaller than that in BKO (Fig. 3c). There was a concomitant upward trend in mature luminal cell abundance in DKO versus BKO (Fig. 3c). This result suggests that antagonism between BRCA1 and COBRA1 influences the relative abundance of mature and progenitor cells in the luminal compartment.
DKO remains defective in HR-mediated DSB repair
Given the established role of BRCA1 in HR-mediated DSB repair, we asked whether the Brca1/Cobra1 interaction affected DSB repair efficiency. First, we used a green fluorescence protein (GFP)-based reporter assay in vitro, in which repair of site-specific DSB through the HR-dependent pathway gives rise to a functional GFP gene40 (Fig. 4a). As expected, short interfering RNA (siRNA)-mediated knockdown of BRCA1 in HeLa cells significantly compromised HR efficiency, as indicated by the reduced percentage of GFP+ cells (Fig. 4b,c; Supplementary Fig. 8). Depletion of COBRA1 alone did not affect HR efficiency, nor did it rescue the HR defect in BRCA1-depleted cells (Fig. 4b,c; Supplementary Fig. 8), suggesting that COBRA1 was not involved in BRCA1-mediated DSB repair in vitro.
(a) Diagram of the GFP reporter assay for measuring HR efficiency. I-Scel: restriction enzyme. iGFP: internal GFP fragment as the template for HR. (b) Immunoblot of COBRA1 and BRCA1 for assessing siRNA knockdown efficiency with control (Con) oligos or ones targeting human BRCA1 (-B) and COBRA1 (-C) in HeLa cells. (c) Percentage of GFP+ cells as a result of HR-mediated DSB repair. Results are average of three independent experiments, and were normalized to the control group. *P<0.05 by Student’s t-test. (d) Mice of 8-wk old were pulse-labelled with BrdU, irradiated (20 Gy), and mammary glands were harvested 3 h later for immunostaining for γH2AX and BrdU. Scale bar, 5 μm. (e) The same samples as shown in d were stained for Rad51 and BrdU. Scale bar, 5 μm. (f) Percentage of Rad51+/BrdU+ mammary epithelial cells. *P<0.05 by Student’s t-test. The numbers of animals used are indicated below the graph. Error bars represent s.e.m. wk, week.
Next, we examined HR efficiency in vivo following ionizing radiation. HR repair predominantly occurs in proliferating cells during late S and G2 phases of the cell cycle, when sister chromatids are available as the homologous templates for HR-mediated repair41. We therefore tracked proliferating cells in irradiated mice by pulse labelling them with BrdU. DSB damage was monitored 3 h after irradiation by immunofluorescence staining for γH2AX. As expected, ionizing radiation-induced γH2AX nuclear foci were present in both BrdU+ and BrdU− cells of WT and KO animals (Fig. 4d). To assess efficiency of HR-dependent DSB repair, we enumerated BrdU+ mammary epithelial cells with ionizing radiation-induced nuclear foci of the well-established HR marker Rad51. In particular, published studies demonstrated that recruitment of Rad51 to DSB sites is facilitated by BRCA1 (refs 42, 43). Consistent with the in vitro findings, irradiated BKO animals exhibited substantially lower Rad51+/BrdU+ ratios versus WT (Fig. 4e,f). CKO mammary glands had similar Rad51+/BrdU+ ratios versus WT control, again suggesting that COBRA1 is not required for ionizing radiation-induced DSB repair per se. Notably, HR repair in DKO mice remained as deficient as that in BKO (Fig. 4e,f). Taken together, the genetic complementation between Brca1 and Cobra1 does not affect DSB repair function of BRCA1.
DKO has restored pubertal transcription
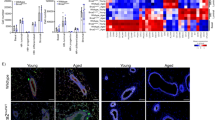
To gain molecular insight into the Brca1/Cobra1 genetic complementation during ductal development at puberty, we carried out gene expression profiling of total mammary epithelial cells from virgin WT, BKO, CKO and DKO at 4, 6 and 8 weeks. Consistent with their normal ductal growth (Fig. 1a,b) and previously reported gene expression profiling of the same animal model32, BKO mice exhibited relatively few transcriptionally affected genes compared with their WT controls (Supplementary Data 1). In contrast, the gene expression profiles of CKO were significantly different from their WT littermates, with the most significant transcriptional aberration observed at the early (4 week) and mid-pubertal (6 week) stages (Fig. 5a; Supplementary Data 1). Furthermore, these CKO-affected genes were enriched with previously identified pubertal genes44 (P=7.65 × 10−13 for 6 weeks by Fisher’s exact test), and oestrogen (P=7.73 × 10−6 by Fisher’s exact test) and progesterone-responsive genes45 (P=5.00 × 10−5 by Fisher’s exact test) in mammary epithelium (Supplementary Data 2). Strikingly, ∼80% of the CKO-affected genes at 4 and 6 weeks were either partially or completely rescued in DKO mammary glands (Fig. 5a; Supplementary Data 2). Likewise, the DKO-rescued genes were enriched with puberty-related (P=2.34 × 10−9 for 6 weeks by Fisher’s exact test) and oestrogen (P=2.09 × 10−5 by Fisher’s exact test) and progesterone-responsive genes (P=7.64 × 10−4 by Fisher’s exact test, Supplementary Data 2). For example, expression of Gata3 and Prlr, two known pubertal genes, was disrupted by Cobra1 ablation but partially restored in DKO (Fig. 5b). We also confirmed the microarray result for several pubertal genes by gene-specific RT-PCR (PCR with reverse transcription; Fig. 5c). Of note, while the transcriptional rescue in DKO occurred as early as 4 weeks (Fig. 5a), restoration of ductal growth in DKO was not apparent until 8 weeks (Fig. 1a,b; Supplementary Fig. 4), likely due to incomplete transcriptional rescue of CKO-affected genes. The fact that transcriptional rescue precedes developmental rescue suggests that the former is likely a cause, rather than consequence, of the restored ductal morphogenesis. Collectively, the antagonistic activity of BRCA1 in pubertal gene expression provides a reasonable explanation for its development-arresting function in CKO mammary glands.
(a) Heatmap illustrates the gene expression changes in total mammary epithelial cells of CKO and DKO as compared with their corresponding WT littermates (n=3) at three time points (4, 6 and 8 wks). The gene expression levels in WT are set as 1. (b) Expression patterns for two representative pubertal genes that are affected by CKO and partially rescued by DKO. The lowest expression level in each graph is set at 1. (c) Confirmation of the microarray data by gene-specific RT-PCR for a number of pubertal genes, using RNA samples from 6-wk old mice. The result is average of values from 3 animals in each genotype. β-Actin mRNA was used for normalization. *P<0.05 by Student’s t-test. Statistical analysis was conducted between WT and CKO, and between CKO and DKO. Error bars represent s.e.m. wk, week.
Discussion
Our mouse genetic study unequivocally demonstrates an important role of the COBRA1 in mammary gland development. Furthermore, our work uncovers genetic complementation between Brca1 and Cobra1 in the context of mammary gland development. Historically, genetic suppression analysis has provided a highly valuable tool used in various simple model organisms for identifying functional interactions between genes that act in a common pathway or two related ones46. Although multiple mechanisms can explain a genetic suppression phenotype, the specific genetic complementation between Brca1 and Cobra1 is most likely due to an antagonistic action of these two gene products in a common pathway. First of all, the observed genetic suppression during DKO mammary gland development is associated with a similar functional rescue in pubertal transcriptional programme, but not in DSB repair. Second, mutual suppression, which was observed in alveologenesis and lactogenesis of DKO mammary glands (Fig. 2), often occurs between two genes that encode interacting proteins46. In this regard, the genetic finding serves as a satisfying functional validation of our previously characterized physical interaction between human BRCA1 and COBRA1 (ref. 11). Lastly, CKO-associated developmental defects were only suppressed by deletion of Brca1, not other tumour suppressor genes tested, including Trp53. This gene-specific genetic suppression most likely reflects a COBRA1-opposing action of BRCA1 in a specific functional pathway, rather than a more general, p53-like DNA damage checkpoint function in cell cycle arrest and/or apoptosis.
The MMTV-Cre system has been widely used in tissue-specific characterization of genes involved in mouse mammary gland development and functions47,48. However, recent reports indicated that MMTV-Cre Line A, which was used in the current study, exhibited moderate impairment in alveologenesis and lactogenesis49,50. This reported side effect of MMTV-Cre unlikely affects our conclusions on BRCA1 and COBRA1 for the following reasons. First, unlike age-matched CKO, hemizygous MMTV-Cre,Cobra1f/+ dams displayed grossly normal alveologenesis and nursing capability. In contrast to the previous reports49,50, the absence of any appreciable MMTV-Cre-associated lactational defects in our study could be due to the more heterogeneous strain background in our work. Furthermore, virgin CKO animals, but not MMTV-Cre,Cobra1f/+, display a dramatic defect in mammary ductal development at all age groups examined (6–24 weeks of age, Fig. 1a,b). In this regard, no ductal defects were reported for virgin MMTV-Cre Line A animals in previously published studies49,50. Lastly, CKO, BKO and DKO, which were assessed in the same MMTV-Cre Line A background, gave rise to distinct alveologenic and lactogenic phenotypes. Taken together, these data strongly suggest that usage of MMTV-Cre Line A alone is not the cause of the profound developmental defects in CKO or the striking genetic rescue observed in DKO.
Our genetic findings clearly indicate that DNA repair-independent antagonism between BRCA1 and COBRA1 plays a critical role in pubertal mammary gland development and maintenance of mammary epithelial homoeostasis. Loss of COBRA1 manifests the BRCA1-mediated inhibition of ductal morphogenesis as observed in CKO, and conversely, BKO mice without functional BRCA1 exhibit enlarged luminal progenitor cell population and defective alveolar development as a result of unopposed COBRA1 actions. Absence of both BRCA1 and COBRA1 in DKO mammary glands could reach a quasi-balanced state that allows for gross tissue development and restored mammary gland functions. While functions of DKO mammary glands are largely restored to a level comparable to WT, the inner layer of DKO epithelial ducts tend to have an expanded K8+ cell population juxtaposed with some K14+ cells. Furthermore, the ratio of CD49b+ over CD49b− luminal cells in DKO is significantly lower in DKO versus WT. Lastly, DKO epithelium experiences both increased proliferation and apoptosis, which could explain why the absolute number of total live cells as measured by flow cytometry is similar to that in WT animals (Fig. 1c). We speculate that DKO luminal epithelium may have the propensity for precocious differentiation, thus resulting in exhaustion of the luminal progenitor subpopulation. As luminal progenitor cells are thought to be the cells of origin for BRCA1-associated tumours37,38,39, investigation is under way to interrogate the impact of Cobra1 ablation on tumorigenesis in Brca1-deficient mouse mammary epithelium. Our current genetic work underscores the importance of interrogating functional interactions between Brca1 and Cobra1 in the physiologically relevant tissue context and developmental window.
Methods
Mice
Cobra1f/f mice have been described previously26. MMTV-Cre,Cobra1f/f mice were generated by breeding MMTV-Cre Line A animals (from Dr Anthony Wynshaw-Boris) with Cobra1f/f mice. Trp53f/f (Trp53tm1Brn), Ink4-Arf KO, and Brca1f/f mice31 were obtained from Mouse Model of Human Cancer Consortium (MMHCC), National Cancer Institute. EIIa-Cre was purchased from the Jackson Laboratory, and used to generate the whole-body hemizygous deletion strain Brca1+/−,Cobra1+/− per previously described procedures26. All mouse strains were in a similarly mixed genetic background (129SvEv/SvJae/C57BL6/FVB). Mutant mice were analysed with their corresponding littermate controls. All procedures performed on animals were in compliance with ethical regulations and were approved by the Institutional Animal Care and Use Committee (IACUC) at the University of Texas Health Science Center at San Antonio.
Whole mount and immunostaining
Inguinal mammary glands from mice of different age groups as indicated in individual figures were used for whole-mount staining. Inguinal fat pads were gently isolated and spread onto a glass slide. Glands were fixed in Carnoy’s fixative (ethanol:chloroform:glacial acetic acid, 60:30:10) overnight at room temperature. Glands were rehydrated in descending grades of alcohol (70, 50 and 30%) for 15 min each, then washed with distilled water before overnight staining in Carmine alum (1 g carmine, 2.5 g aluminium potassium sulfate boiled for 20 min in distilled water, filtered and brought to a final volume of 500 ml). Stained glands were dehydrated in ascending grades of alcohol (70, 70, 90, 95, 100 and 100%) for 15 min each, and cleared with Citrisolv reagent (Fisher, Cat no. 22-143975). Samples were examined under a Nikon SMZ1000 dissection microscope. Duct length was measured from calibrated images using Eclipse software. Average length of three longest ducts from nipple region in inguinal mammary glands was taken as the ductal length of each animal. The value shown in Fig. 1b is mean of ductal lengths for multiple animals of a given genotype at a given age±s.e.m.
Primary antibodies used were anti-COBRA1 (1:50) (ref. 26), anti-milk protein (Nordic Immunology, RAM/MSP, 1:10,000), anti-K8 (Developmental Studies Hybridoma Bank, TROMA-1, 1:100), anti-K14 (Covance, PRB-155P, 1:5,000), anti-BrdU (GE Healthcare, RPN20, 1:10,000), anti-γH2AX (Cell Signaling, 9718, 1:500), and anti-Rad51 (Santa Cruz, sc-8349, 1:100).
Mammary glands were fixed in 10% neutral buffered formalin for 18 h at 4 °C and paraffin embedded. Sections of 2 or 3 μM in thickness were used for hematoxylin–eosin (H&E) staining, immunofluorescent staining and immunohistochemistry. Samples were baked at 70 °C for 15 min, then de-paraffinized by three 5-min extractions in 100% xylene, followed by 3-min each of descending grade of alcohol (100% twice, 95, 70 and 50%). Samples were washed briefly with phosphate-buffered saline (PBS) before transferring to boiling antigen-unmasking solution (Vector Labs, H-3300) for 20 min. For immunohistochemistry, sections were pre-treated with 3% hydrogen peroxide for 10 min before blocking. Blocking was performed with 10% normal goat serum in PBS for 1 h at room temperature followed by primary antibody incubation overnight at 4 °C. For detection with primary antibody using the immune enzymatic method, the ABC peroxidase detection system (Vector Labs, PK-6105) was used with 3, 3′-diaminobenzidine (DAB) as substrate (Vector Labs, SK-4105) according to manufacturer’s instruction.
For immunofluorescence staining, sections were incubated with Alexa-488 and Alexa-546-conjugated secondary antibodies (Life Technologies), mounted with Vectashield mounting medium with DAPI (Vector Labs, H-1200), and examined with an Olympus FV1000 confocal microscope or Nikon Eclipse Ni fluorescent microscope. For BrdU pulse-labelling, mice were intraperitoneally injected with cell proliferation labelling reagent (GE Healthcare, RPN201) at 16.7 ml kg−1. For BrdU/Rad51 and BrdU/γΗ2ΑX double staining, mice were first injected with BrdU and then X-rayed at 20 Gy using a Faxitron cabinet X-ray system (Model 43855F). Mammary glands were harvested 3 h after labelling.
Flow cytometry
Thoracic and inguinal mammary glands from virgin mice were isolated in sterile condition and lymph nodes from inguinal glands were removed. Single cells were prepared using published protocol51 with minor modifications. All reagents were purchased from StemCell Technologies (Vancouver, Canada), unless otherwise indicated. Briefly, isolated glands were minced using scissors and digested for 15–18 h at 37 °C in DMEM F-12 (Cat. no. 36254) containing 2% fetal bovine serum (FBS), insulin (5 mg ml−1), penicillin–streptomycin and a final concentration of 1 mg ml−1 collagenase and 100 U ml−1 hyaluronidase (Cat. no. 07919). After vortexing, epithelial organoids were collected by centrifugation at 600g for 4 min. Red blood cells (RBCs) in the resulting pellets were lysed with 0.8% NH4Cl. Epithelial organoids were then digested by pipetting with 2 ml of 0.05% pre-warmed Trypsin (Life Technologies, 25300) for 2 min, followed by washing in ice-cold Hanks Balanced Salt Solution (Cat. no. 37150) with 2% FBS (HF). Cells were resuspended in 5 mg ml−1 Dispase (Cat. no. 07913) with 0.1 mg ml−1 DNAse I (Sigma-Aldrich, D4513). After trituration for 1–2 min, cells were resuspended in ice-cold HF, and single cells were prepared by filtering the cell suspension through a 40-μm cell strainer (Fisher, Cat. no. 22363547). Cells were counted and resuspended in HF at a concentration of 1 × 106 cells per 100 μl. Cell were incubated for 10 min on ice with 10% rat serum (Jackson Laboratories, 012-000-120) and Fc receptor antibody (BD Biosciences, 553141). After blocking, cells were incubated for 20 min with antibodies for the following cell-surface markers: Ep-CAM-PE (BioLegend, 118206, 0.5 μl per 100 μl), CD49f-FITC (BD Biosciences, 555735, 2 μl per 100 μl), CD31-Biotin (BD Bioscience, 553371, 1 μl per 100 μl), CD45 biotin (BioLegend, 103103, 1 μl per 100 μl), TER-119 Biotin (BioLegend, 103511, 1 μl per 100 μl), and CD49b-Alexa Fluor 647 (BioLegend, 104317, 0.5 μl per 100 μl) followed by Streptavidin-Pacific Blue (Invitrogen, S11222, 0.5 μl per 100 μl) incubation. 7-AAD (BD Biosciences, 559925, 5 μl per 100 μl) was added 10 min before analysis. CD49b+ cells were gated using a fluorescent-minus-one control, in which all antibodies except CD49b-Alexa 647 were used. Sorting was performed with a Moflo Astrios cell sorter (Beckmen Coulter). Data were analysed using FACSDiva software. Purity of the stromal, luminal, and myoepithelial populations were verified by RT-PCR analysis of Vimentin (stromal), K18 (luminal), K5 (myoepithelial) and K14 (myoepithelial) mRNA.
In vitro HR-based DSB repair assay
The homology-directed repair assay was performed using established methods52. The recombination substrate, pDR-GFP, contains two inactive GFP genes, one of which is due to the presence of an I-SceI endonuclease recognition sequence. This DNA is integrated into a single site in HeLa cells. On day 1, siRNAs specific for a control sequence, COBRA1, and BRCA1 were transfected, using Oligofectamine (Invitrogen), into wells containing HeLa-DR-GFP cells. On day 3, the cells were re-transfected with the same siRNAs plus a plasmid for the expression of the I-SceI endonuclease using the Lipofectamine 2000 transfection reagent (Invitrogen). On day 6, cells were released from the monolayer using trypsin and the fraction of GFP+ cells was determined using a FACSCalibur analytical flow cytometry instrument. Results were normalized to the per cent of GFP+ cells in the sample in which the control siRNA was transfected.
Gene expression profiling and bioinformatics analysis
Triplicates of RNA samples from live Lin-/CD24+ mammary epithelial cells of WT and mutant mice were labelled using the Illumina TotalPrepTM RNA amplification kit (Ambion, Cat. no. AMIL1791) and subsequently hybridized to Illumina mouse whole genome gene expression BeadChips (MouseRef-8 version 2.0, Illumina). BeadChips were scanned on an iScan Reader (Illumina) using iScan software (version 3.3.29, Illumina). For further analysis, the scanned data were uploaded into GenomeStudio software (version 1.9.0, Illumina) via the gene expression module (Direct Hyb). The genomic data are available at NIH Gene Expression Omnibus site (GSE67440).
For each of the time points, we identified genes that are affected by Cobra1 KO (CKO-affected) and those that are eventually rescued by double KO (DKO-rescued). We define CKO-affected genes as the genes that show ≥2.0 fold enrichment (either up or down) in CKO mice compared with corresponding WT control mice, with P≤0.05. DKO-rescued genes are defined as those CKO-affected genes that had either (1) ≤1.5 fold enrichment (either up or down, P<0.05) in DKO versus WT control mice, or (2) fold of changes in DKO versus WT (P<0.05) no more than 50% of those in CKO versus WT, or (3) any fold of changes in DKO versus WT with P value larger than 0.05. Supplementary Data 2 shows the total number of CKO-affected and DKO-rescued genes for the indicated time points.
Pubertal, oestrogen and progesterone signature genes were extracted from previously published studies44,45 to identify the overlap with CKO-affected or DKO-rescued genes. Supplementary Data 2 shows the overlap among CKO-affected/DKO-rescued genes with pubertal, oestrogen and progesterone genes. The statistical significance (P value) of the overlap was calculated using the Fisher’s exact test:

where N is the total number of genes in the experiment; m,n is the selected affected/rescued and previously published signature genes respectively and o is the overlap among those genes. C(n,k) is the binomial coefficient.
The microarray data have been submitted to the NIH database (accession number GSE67440).
Primer sequences
For RT-PCR: 18sRNA-F: 5′-GAATTCCCAGTAAGTGCGGG-3′, 18sRNA-R: 5′-GGGCAGGGACTTAATCAACG-3′. Cobra1-F: 5′-ACAACTTCTTCAGCCCTTCCC-3′, Cobra1-R: 5′-TCTGCACCACCTCTCCTTGG-3′. Brca1-F: 5′-AGCAAACAGCCTGGCATAGC-3′, Brca1-R: 5′-ACTTGCAGCCCATCTGCTCT-3′. p16Ink4a-F: 5′-GAACTCTTTCGGTCGTACCCC-3′, p16Ink4a-R: 5′-CGTGAACGTTGCCCATCAT-3′. p19Arf-F: 5′-CTTGAGAAGAGGGCCGCAC-3′, p19Arf-R: 5′-AACGTTGCCCATCATCATCA-3′. p53-F: 5′-GAGACAGCAGGGCTCACTCC-3′, p53-R: 5′-TGGCCCTTCTTGGTCTTCAG-3′. Ctse-F: 5′-ATTGGCAGATTGCCCTGGAT-3′, Ctse-R: 5′-GCCTTCGGAGCAGAACATCA-3′. Prom2-F: 5′-TGACCTGGATAAGCACCTGG-3′, Prom2-R: 5′-AAGCTCTGAAGCTCCTGCTG-3′. Acot1-F: 5′-ATGGCAGCAGCTCCAGACTT-3′, Acot1-R: 5′-CCCAACCTCCAAACCATCAT-3′. Ramp2-F: 5′-GCCTCATCCCGTTCCTTGTT-3′, Ramp2-R: 5′-CCTGGGCATCGCTGTCTTTA-3′. Vwf-F: 5′-CGACCTGGAGTGTATGAGCC-3′, Vwf-R: 5′-ACACACTTGTTTTCGTGCCG-3′. Gata3-F: 5′-GATGTAAGTCGAGGCCCAAG-3′, Gata3-R: 5′-GCAGGCATTGCAAAGGTAGT-3′. K18-F: 5′- ACTCCGCAAGGTGGTAGATGA-3′, K18-R: 5′- TCCACTTCCACAGTCAATCCA-3′, K14-F: 5′- AGCGGCAAGAGTGAGATTTCT-3′, K14-R: 5′- CCTCCAGGTTATTCTCCAGGG -3′, K5-F: 5′-GAGATCGCCACCTACAGGAA-3′, K5-R: 5′-TCCTCCGTAGCCAGAAGAGA-3′, Vimentin-F: 5′-CGGCTGCGAGAGAAATTGC-3′, Vimentin-R: 5′- CCACTTTCCGTTCAAGGTCAAG-3′, β-Actin-F: 5′-CGGTTCCGATGCCCTGAGGCTCTT-3′, β-Actin-R: 5′-CGTCACACTTCATGATGGAATTGA-3′.
Statistical analysis
All data are expressed as arithmetic mean±s.e.m. Differences between relevant test and control group means were tested using an unpaired two-tailed Student’s t-test unless otherwise indicated. For measures repeated at multiple ages, comparisons between test and control were performed on matched data from each time point. We considered a P value of<0.05 as statistically significant. Analytical approaches for microarray data are given in a separate section above.
Additional information
Accession codes: The microarray data have been deposited in the NIH database under the accession number GSE67440.
How to cite this article: Nair, S. J. et al. Genetic suppression reveals DNA repair-independent antagonism between BRCA1 and COBRA1 in mammary gland development. Nat. Commun. 7:10913 doi: 10.1038/ncomms10913 (2016).
Accession codes
References
King, M. C. "The race" to clone BRCA1. Science 343, 1462–1465 (2014).
Venkitaraman, A. R. Cancer suppression by the chromosome custodians, BRCA1 and BRCA2. Science 343, 1470–1475 (2014).
Rosen, E. M., Fan, S. & Ma, Y. BRCA1 regulation of transcription. Cancer Lett. 236, 175–185 (2006).
Lee, E. Y. & Abbondante, S. Tissue-specific tumor suppression by BRCA1. Proc. Natl Acad. Sci. USA 111, 4353–4354 (2014).
Scully, R. et al. BRCA1 is a component of the RNA polymerase II holoenzyme. Proc. Natl Acad. Sci. USA 94, 5605–5610 (1997).
Fan, S. et al. BRCA1 inhibition of estrogen receptor signaling in transfected cells. Science 284, 1354–1356 (1999).
Tkocz, D. et al. BRCA1 and GATA3 corepress FOXC1 to inhibit the pathogenesis of basal-like breast cancers. Oncogene 31, 3667–3678 (2012).
Hu, Y. F., Hao, Z. L. & Li, R. Chromatin remodeling and activation of chromosomal DNA replication by an acidic transcriptional activation domain from BRCA1. Genes Dev. 13, 637–642 (1999).
Hu, Y.-F., Miyake, T., Ye, Q. & Li, R. Characterization of a novel trans-activation domain of BRCA1 that functions in concert with the BRCA1 C-terminal (BRCT) domain. J. Biol. Chem. 275, 40910–40915 (2000).
Miyake, T., Hu, Y.-F., Yu, D. S. & Li, R. A functional comparison of BRCA1 C-terminal domains in transcription activation and chromatin remodeling. J. Biol. Chem. 275, 40169–40173 (2000).
Ye, Q. et al. BRCA1-induced large-scale chromatin unfolding and allele-specific effects of cancer-predisposing mutations. J. Cell Biol. 155, 911–921 (2001).
Hu, Y.-F. & Li, R. JunB potentiates function of BRCA1 activation domain 1(AD1) through a coiled-coil-mediated interaction. Genes Dev. 16, 1509–1517 (2002).
Gorski, J. J. et al. Profiling of the BRCA1 transcriptome through microarray and ChIP-chip analysis. Nucleic Acids Res. 39, 9536–9548 (2011).
Consortium, E. P.. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Gardini, A., Baillat, D., Cesaroni, M. & Shiekhattar, R. Genome-wide analysis reveals a role for BRCA1 and PALB2 in transcriptional co-activation. EMBO J. 33, 890–905 (2014).
Yamaguchi, Y. et al. NELF, a multisubunit complex containing RD, cooperates with DSIF to repress RNA polymerase II elongation. Cell 97, 41–51 (1999).
Kwak, H. & Lis, J. T. Control of transcriptional elongation. Annu. Rev. Genet. 47, 483–508 (2013).
Adelman, K. & Lis, J. T. Promoter-proximal pausing of RNA polymerase II: emerging roles in metazoans. Nat. Rev. Genet. 13, 720–731 (2012).
Gilchrist, D. A. et al. Pausing of RNA polymerase II disrupts DNA-specified nucleosome organization to enable precise gene regulation. Cell 143, 540–551 (2010).
Sun, J. & Li, R. Human negative elongation factor activates transcription and regulates alternative transcription initiation. J. Biol. Chem. 285, 6443–6452 (2010).
Sun, J. et al. Genetic and genomic analyses of RNA polymerase II-pausing factor in regulation of mammalian transcription and cell growth. J. Biol. Chem. 286, 36248–36257 (2011).
Aiyar, S. E. et al. Attenuation of estrogen receptor alpha-mediated transcription through estrogen-stimulated recruitment of a negative elongation factor. Genes Dev. 18, 2134–2146 (2004).
Kininis, M., Isaacs, G. D., Core, L. J., Hah, N. & Kraus, W. L. Postrecruitment regulation of RNA polymerase II directs rapid signaling responses at the promoters of estrogen target genes. Mol. Cell Biol. 29, 1123–1133 (2009).
Danko, C. G. et al. Signaling pathways differentially affect RNA polymerase II initiation, pausing, and elongation rate in cells. Mol. Cell 50, 212–222 (2013).
Levine, M. Paused RNA polymerase II as a developmental checkpoint. Cell 145, 502–511 (2011).
Amleh, A. et al. Mouse cofactor of BRCA1 (Cobra1) is required for early embryogenesis. PloS ONE 4, e5034 (2009).
Williams, L. H. et al. Pausing of RNA polymerase II regulates mammalian developmental potential through control of signaling networks. Mol. Cell 58, 311–322 (2015).
Pan, H. et al. RNA polymerase II pausing factor NELF controls energy homeostasis in cardiomyocytes. Cell Rep. 7, 79–85 (2014).
Wagner, K. U. et al. Cre-mediated gene deletion in the mammary gland. Nucleic Acids Res. 27, 4323–4330 (1997).
Shehata, M. et al. Phenotypic and functional characterisation of the luminal cell hierarchy of the mammary gland. Breast Cancer Res. 14, R134 (2012).
Xu, X. et al. Conditional mutation of Brca1 in mammary epithelial cells results in blunted ductal morphogenesis and tumour formation. Nat. Genet. 22, 37–43 (1999).
Smart, C. E. et al. Analysis of Brca1-deficient mouse mammary glands reveals reciprocal regulation of Brca1 and c-kit. Oncogene 30, 1597–1607 (2011).
Kim, W. Y. & Sharpless, N. E. The regulation of INK4/ARF in cancer and aging. Cell 127, 265–275 (2006).
Hakem, R. et al. The tumor suppressor gene Brca1 is required for embryonic cellular proliferation in the mouse. Cell 85, 1009–1023 (1996).
Ludwig, T., Chapman, D. L., Papaioannou, V. E. & Efstratiadis, A. Targeted mutations of breast cancer susceptibility gene homologs in mice: lethal phenotypes of Brca1, Brca2, Brca1/Brca2, Brca1/p53, and Brca2/p53 nullizygous embryos. Genes Dev. 11, 1226–1241 (1997).
Xu, X. et al. Genetic interactions between tumor suppressors BRCA1 and p53 in apoptosis, cell cycle and tumorigenesis. Nat. Genet. 28, 266–271 (2001).
Lim, E. et al. Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat. Med. 15, 907–913 (2009).
Molyneux, G. et al. BRCA1 basal-like breast cancers originate from luminal epithelial progenitors and not from basal stem cells. Cell Stem Cell 7, 403–417 (2010).
Proia, T. A. et al. Genetic predisposition directs breast cancer phenotype by dictating progenitor cell fate. Cell Stem Cell 8, 149–163 (2011).
Pierce, A. J., Hu, P., Han, M., Ellis, N. & Jasin, M. Ku DNA end-binding protein modulates homologous repair of double-strand breaks in mammalian cells. Genes Dev. 15, 3237–3242 (2001).
Ira, G. et al. DNA end resection, homologous recombination and DNA damage checkpoint activation require CDK1. Nature 431, 1011–1017 (2004).
Scully, R. et al. Association of BRCA1 with Rad51 in mitotic and meiotic cells. Cell 88, 265–275 (1997).
Parameswaran, B. et al. Damage-induced BRCA1 phosphorylation by Chk2 contributes to the timing of end resection. Cell Cycle 4, 437–448 (2015).
McBryan, J., Howlin, J., Kenny, P. A., Shioda, T. & Martin, F. ERalpha-CITED1 co-regulated genes expressed during pubertal mammary gland development: implications for breast cancer prognosis. Oncogene 26, 6406–6419 (2007).
Lu, S. et al. Transcriptional responses to estrogen and progesterone in mammary gland identify networks regulating p53 activity. Endocrinology 149, 4809–4820 (2008).
Prelich, G. Suppression mechanisms: themes from variations. Trends Genet. 15, 261–266 (1999).
Wagner, K. U. et al. Spatial and temporal expression of the Cre gene under the control of the MMTV-LTR in different lines of transgenic mice. Transgenic Res. 10, 545–553 (2001).
Zhang, Q. et al. Temporally and spatially controlled expression of transgenes in embryonic and adult tissues. Transgenic Res. 19, 499–509 (2010).
Robinson, G. W. & Hennighausen, L. MMTV-Cre transgenes can adversely affect lactation: considerations for conditional gene deletion in mammary tissue. Anal. Biochem. 412, 92–95 (2011).
Yuan, T., Wang, Y., Pao, L., Anderson, S. M. & Gu, H. Lactation defect in a widely used MMTV-Cre transgenic line of mice. PLoS ONE 6, e19233 (2011).
Stingl, J. et al. Purification and unique properties of mammary epithelial stem cells. Nature 439, 993–997 (2006).
Ransburgh, D. J., Chiba, N., Ishioka, C., Toland, A. E. & Parvin, J. D. Identification of breast tumor mutations in BRCA1 that abolish its function in homologous DNA recombination. Cancer Res. 70, 988–995 (2010).
Acknowledgements
We thank Drs Anthony Wynshaw-Boris for the MMTV-Cre mouse strain, Jonathan Gelfond for assistance in statistical analysis, and Tyler Curiel for critical reading of the manuscript. The work was supported by grants to Y.H. from the National Institutes of Health (NIH CA170306), to J.R. from NIH (G12MD007591, P20CA165589), to J.D.P. from NIH (CA141090), and to R.L. from NIH (CA161349 and CA184084), the Tom C. & H. Frost Endowment, the Max and Minnie Tomerlin Voelcker Fund, and Avon Foundation for Women. S.J.N was supported by a Department of Defense Predoctoral Fellowship (W81XWH-09-1-0014). We also thank generous support from Cancer Therapy and Research Center at University of Texas Health Science Center at San Antonio (P30CA054174).
Author information
Authors and Affiliations
Contributions
R.L. and Y.H. co-managed the project. R.L., Y.H., S.J.N., X.Z. and H-C.C. designed the experiments, S.J.N., X.Z., H-C.C., P.G., C.A., N.S., and T.B. carried out the experiments, M.J.J., Y.W., F.S.A., J.R., J-B.F., J.D.P. and V.X.J. analysed the data, R.L., Y.H., S.J.N., X.Z. and H-C.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-8 and Supplementary Table 1 (PDF 1006 kb)
Supplementary Data 1
Lists of genes that are differentially expressed between wild-type control mice and mutant mice at 4, 6 and 8 weeks of ages. (XLSX 100 kb)
Supplementary Data 2
Lists of total and puberty-related genes that are affected by CKO and rescued by DKO. (XLSX 149 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Nair, S., Zhang, X., Chiang, HC. et al. Genetic suppression reveals DNA repair-independent antagonism between BRCA1 and COBRA1 in mammary gland development. Nat Commun 7, 10913 (2016). https://doi.org/10.1038/ncomms10913
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms10913
This article is cited by
-
RNA polymerase II pausing factor NELF in CD8+ T cells promotes antitumor immunity
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.