Abstract
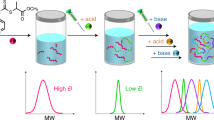
Precise control over the location of monomers in a polymer chain has been described as the ‘Holy Grail’ of polymer synthesis. Controlled chain growth polymerization techniques have brought this goal closer, allowing the preparation of multiblock copolymers with ordered sequences of functional monomers. Such structures have promising applications ranging from medicine to materials engineering. Here we show, however, that the statistical nature of chain growth polymerization places strong limits on the control that can be obtained. We demonstrate that monomer locations are distributed according to surprisingly simple laws related to the Poisson or beta distributions. The degree of control is quantified in terms of the yield of the desired structure and the standard deviation of the appropriate distribution, allowing comparison between different synthetic techniques. This analysis establishes experimental requirements for the design of polymeric chains with controlled sequence of functionalities, which balance precise control of structure with simplicity of synthesis.
Similar content being viewed by others
Introduction
How precisely can we control the location of a monomer in a synthetic polymer? This question is crucial to the synthesis of aperiodic copolymers: ‘copolymers in which monomer sequence distribution is not regular but follows the same arrangement in all chains’1. Included in this definition are polymers that contain an ordered sequence of distinct blocks, and polymers that contain specific monomers at precisely defined locations2,3,4. Such polymers could serve as a medium for information storage, as does DNA, or catalyse complex reactions, as do enzymes5. But for these applications to be realized, precise control over structure is essential, and any deviation from the desired structure must be quantified.
If only small quantities of polymer are required, near perfect control over structure can be obtained using stepwise synthesis on a solid6,7 or soluble polymer8,9 support, as in peptide synthesis. For the rapid preparation of moderate to large quantities of well-defined polymer, however, we must turn to living or controlled chain growth polymerizations10,11,12,13,14,15,16,17,18,19,20,21,22,23. But unlike stepwise techniques, chain growth polymerization is a statistical process that produces a distribution of products. In this paper, we establish a lower bound to the structural variation that exists even under ideal conditions of living polymerization. Our results are applicable to all chain polymerizations, which are carried out under living or near-living conditions, including ring-opening metathesis polymerization (ROMP)24, the various types of reversible deactivation radical polymerization (RDRP, also known as controlled/‘living’ radical polymerization)25, and ionic polymerizations26,27.
The production of aperiodic copolymers requires control over the arrangement of monomers, which should be replicated in all chains. The degree of control required can be interpreted with varying degrees of severity, corresponding to the sequence-defined, multisite and multiblock copolymers considered by Lutz1 (Fig. 1): (a) control over the absolute position of a monomer with respect to the chain end, or the absolute separation between two monomers (sequence-defined copolymers); (b) control over the relative position of a monomer with respect to the total chain length, or the relative separation between two monomers (multisite copolymers) or (c) control over the number and order of blocks in a multiblock copolymer, without regard to the distribution of each individual block (multiblock copolymers).
(a) Absolute control of monomer position and/or separation in a sequence-defined copolymer. (b) Relative control of monomer position and/or separation in a multisite copolymer. (c) Control of number and order of blocks in a multiblock copolymer. Throughout this paper, the letters A, B, C and D refer to different types of monomer. A collection of polymer chains containing on average n units of A followed by m units of B is represented as {AnBm}, whereas a single polymer chain containing exactly N units of A followed by M units of B is represented as ANBM.
The level of structural control that is actually achieved can be quantified in two ways: in terms of the fraction of chains that correspond to the desired structure (the yield), and in terms of the standard deviation, σ, of the distribution. High yields and low standard deviations imply high levels of control. In this article, we propose the standard deviation of the monomer distribution as a quantitative measure of precision; derive simple expressions for these distributions under ideal chain growth polymerization conditions and apply these expressions to a number of recent examples of multisite and multiblock copolymers. Our results demonstrate that the level of control over monomer placement is quite limited, even under ideal conditions, and that multiblock copolymers comprised of many short blocks will inevitably contain a large proportion of defective chains.
Results
Absolute position of monomers
Absolute control over the location of an individual monomer is the most demanding goal of polymer synthesis. However, techniques based on chain-growth polymerization lead, even in the best-controlled polymerizations, to a distribution of structures. For very short chains, separation of a single oligomeric structure may be possible, but requires extensive use of chromatography28,29.
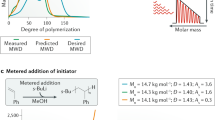
Under ideal conditions of living polymerization, the degree of polymerization obeys a Poisson distribution30, whose probability mass function is given in Table 1 (full derivations of all results in Table 1 may be found in Supplementary Note 1). In reality, unavoidable side reactions such as termination and chain transfer result in deviations from the Poisson distribution. The ideal is most closely approached by anionic polymerization31, but this technique must be carried out in dry, nonprotic solvents and is incompatible with most functional groups. RDRP techniques32,33,34,35,36,37,38,39 tolerate a wide range of functionalities, but lead to broader distributions because of termination reactions and the addition of multiple monomers per activation/deactivation cycle.
The highest level of control over monomer placement achievable using chain growth polymerization techniques is obtained by addition of a non-homopolymerizing monomer (B), such as maleic anhydride, to a polymer of average length n ({An}), followed by subsequent chain extension10,11,12,13. As B cannot homopolymerize, only one monomer of B will be added to each chain. Use of an excess of B ensures that one monomer of B will be added to every chain, giving the average structure {An}B1. In a variation on this technique, addition of B may occur by a step-growth reaction, such as via azide-alkyne coupling, followed by chain extension from a suitable functional group40,41.
Numbering the monomers from the start of the chain, the expected position of B is n+1. The probability that B will be found in this position is approximately  and the standard deviation of the distribution is
and the standard deviation of the distribution is  . For example, in a polymer with average structure {A10}B1{C10}, only 12.5% of the chains will contain monomer B as the 11th unit, while in 26.6% of the chains, B will appear before position 8 or after position 14.
. For example, in a polymer with average structure {A10}B1{C10}, only 12.5% of the chains will contain monomer B as the 11th unit, while in 26.6% of the chains, B will appear before position 8 or after position 14.
In aperiodic copolymers prepared by sequential polymerizations of monomers of similar reactivity14,15,16,17,18,19,20, or by addition of alternating21,22 or very rapidly polymerizing23 monomers at intervals during the polymerization, all blocks are Poisson-distributed, at best. Thus, an {A10B1C10} polymer will include chains with no B units, as well as chains that contain 2 or more5,42. The proportion of chains that contain a unit of B in position 11 (the midpoint) can be calculated by excluding all chains that contain 11 or more units of A (41.7% of chains), as well as all chains in which the sum of units of A and B is <11 (46.0% of chains). The 12.3% of chains that remain are not significantly fewer than the corresponding proportion of {A10}B1{C10}. In {A10}B1{C10}, however, each polymer chain contained a single copy of B. In {A10B1C10}, only 4.6% of chains contain one and only one unit of B in position 11.
In general, the probability that the kth monomer of a multiblock copolymer is part of the nth block is given by the difference between two Poisson cumulative distribution functions (Table 1). As the average position of a block moves further along the polymer chain, its location becomes more diffuse. Shorter blocks are affected to a greater extent than longer blocks, as shown by Fig. 2a,c, which show the location distributions of each block of two ABACADACAB decablock copolymers. Those of the short blocks broaden rapidly and overlap significantly. The distributions of the longer blocks broaden more slowly and overlap less, but fewer changes in composition are possible in a given length of polymer. The standard deviation of the location of the block (Table 1) provides a quantitative measure of the attainable precision: in {A10B1C10}, the expected location of B is 11.5, with a standard deviation of 3.3; B occurs before position 8 or after position 14 in 29.2% of chains.
Relative position of monomers
Although absolute control over monomer placement is the ideal, it would often be acceptable to control only the relative position of a monomer with respect to the total chain length. In a diblock copolymer, {AnBm}, comprising two Poisson-distributed blocks, the relative length of the A block with respect to the entire polymer chain is closely approximated by a Beta(n,m) distribution (see Supplementary Figs 1 and 9 and Supplementary Note 2). The beta distribution is one of the most widely used distributions in statistics, and is often used to model random variables that are limited to a finite range, such as world cloud cover43 or the distribution of genetic variation44.
In the {A10}B1{C10} copolymer, the distribution of B can be approximated as the distribution of the interface between A and C blocks in an {A10C10} diblock copolymer: a Beta(10,10) distribution with expected value 0.50 and standard deviation 0.11. Thus, in ∼30% of the chains, we expect to find B outside the relative length range of 0.39–0.61—that is, outside the central fifth of the polymer. In general, the distribution of the relative location of a single inserted monomer narrows as the total length of the polymer increases, and is broadest when the monomer is inserted at the midpoint of the chain (Supplementary Fig. 2).
In an {AnBmCp} copolymer whose inserted block {Bm} is also Poisson distributed, the probability that a monomer of type B will be found in a given relative position is given by the difference between two cumulative beta distribution functions with parameters (n, m+p) and (n+m, p), respectively (Table 1). Thus, in the {A10B1C10} copolymer, the expected relative position of the B block is 0.50, and its standard deviation is 0.11—almost identical to that of the {A10}B1{C10} copolymer. Only 17.6% of chains contain a monomer of type B at the midpoint. Doubling the average length of the B block ({A10B2C10}) increases the proportion of chains that contain a monomer of type B at the midpoint to 33.6%, but the length must be increased to 11 ({A10B11C10}) to ensure that 95% of the chains contain a monomer of type B at the midpoint. The effects of average block length and position on relative position distributions are illustrated in Fig. 2 for two decablock copolymers.
The effect of Poisson broadening on the control that can be achieved over monomer location can be demonstrated using two recently reported multisite polymers21. The polymers were prepared by successive additions of the non-homopolymerizing monomers N-benzyl maleimide (Bz-MI), N-propyl maleimide (Pr-MI) and N-pentafluorophenyl maleimide (PFP-MI) to a single electron transfer living radical polymerization (SET-LRP) of styrene with targeted  of 65 (ref. 21). For the first polymer, maleimide solutions were added at intervals during the polymerization corresponding to 28, 46 and 70% styrene conversion, leading to the insertion of short segments of copolymer containing on average 1 unit of Bz-MI, Pr-MI or PFP-MI. The second polymer was prepared using a lower initial styrene to initiator ratio and delaying the addition of each type of maleimide until the styrene had reached high conversion. After each maleimide had been fully incorporated, additional styrene was added to generate the intervening polystyrene segments. The effect was to halve the average length of the segments containing Bz-MI and Pr-MI, from 6 in the first synthesis to 3 in the second synthesis. The average length of the PFP-MI block was 3 in both syntheses.
of 65 (ref. 21). For the first polymer, maleimide solutions were added at intervals during the polymerization corresponding to 28, 46 and 70% styrene conversion, leading to the insertion of short segments of copolymer containing on average 1 unit of Bz-MI, Pr-MI or PFP-MI. The second polymer was prepared using a lower initial styrene to initiator ratio and delaying the addition of each type of maleimide until the styrene had reached high conversion. After each maleimide had been fully incorporated, additional styrene was added to generate the intervening polystyrene segments. The effect was to halve the average length of the segments containing Bz-MI and Pr-MI, from 6 in the first synthesis to 3 in the second synthesis. The average length of the PFP-MI block was 3 in both syntheses.
The distribution of each maleimide in the polymer was then calculated, making use of the following simplifying assumptions: (i) all segments were assumed to follow a Poisson distribution; (ii) the maleimides were assumed to be evenly distributed within their respective segments. In this way, the approximate distribution of each maleimide could be obtained by calculating the distribution of each maleimide-containing segment, then normalizing with respect to the segment’s maleimide content. Differences in homo- and cross-propagation rate constants of styrene and maleimide were not explicitly considered, as it was assumed that in all cases, the rate of reversible deactivation was much greater than that of propagation. If this assumption is invalid (for example, as a result of very rapid cross-propagation occurring at a comparable rate to deactivation) then broader distributions are obtained.
Despite the much shorter segments obtained in the second synthesis, the maleimide distributions are nearly identical (Fig. 3 and Supplementary Table 2). Their standard deviations are determined primarily by the length of the surrounding blocks, with only a small contribution from the length of the block itself. The calculated distributions underestimate the true degree of structural variation, as a Poisson-distributed copolymer with  of 53 has a dispersity of 1.02, whereas the measured dispersities of the copolymers were 1.16 (sequential maleimide additions) and 1.27 (alternating maleimide and styrene additions).
of 53 has a dispersity of 1.02, whereas the measured dispersities of the copolymers were 1.16 (sequential maleimide additions) and 1.27 (alternating maleimide and styrene additions).
(a) containing benzyl maleimide (Bz-MI), propyl maleimide (Pr-MI) and pentafluorophenyl maleimide (PFP-MI) insertions, prepared using maleimide additions only (b–d) or by alternating additions of maleimide and styrene (e–g) to an SET-LRP polymerization of styrene21. The expected structures of each copolymer (a,b,e) consist of short regions containing a single maleimide unit, dispersed along a polystyrene chain. The distribution of each type of maleimide is shown in terms of absolute position (c,f) and position relative to the total chain length (d,g).
We have applied similar analyses to an icosablock copolymer prepared by sequential reversible addition-fragmentation chain transfer (RAFT) polymerizations14 (Supplementary Fig. 3), a multisite copolymer prepared by sequential additions of exo-functionalized norbornene monomers to the ROMP polymerization of an endo-functional norbornene monomer (Supplementary Fig. 4)23, and two undecablock copolymers prepared by sequential anionic polymerizations (Supplementary Fig. 5)15,16. In the RAFT and ROMP copolymers, the use of multiple short blocks (typical length 3 and 2, respectively) results in poorly defined structures. The much longer blocks obtained using anionic copolymerization (minimum block lengths 37 (ref. 15) and 130 (ref. 16)) allow greater control over the relative position of a block within the chain.
Existence of blocks in multiblock copolymers
The least restrictive goal of aperiodic polymer synthesis is simply to prepare a multiblock copolymer in which each polymer chain contains the desired number of blocks, in the correct order. If, for example, a block of monomer A encoded 1, a block of monomer B encoded 0 and monomer C served to separate consecutive blocks of A or B, a binary string could be represented using a multiblock copolymer, even without specifying the length of each block. The only condition to be fulfilled is that every chain must contain at least one monomer unit from each block.
The probability that a Poisson-distributed block of average length l contains at least one monomer is 1–e−l. For a polymer composed of n blocks, the proportion of chains that contain all blocks is obtained by multiplying the probabilities for each block. If all blocks are nominally of equal length, the minimum block length, lmin, required for a given proportion, α, of chains to contain all blocks is proportional to ln(n).

Thus, polymers consisting of many short blocks will contain a significant proportion of defective chains (see Supplementary Figs 3 and 4 for details). Attempts to insert a block of average length 3 will fail in ∼5% of chains; in a copolymer containing 18 such blocks, the majority of chains will be defective (Supplementary Fig. 3). Doubling the targeted block length to 6 results in an exponential decrease in the proportion of missing blocks, which would allow a 20-block copolymer to be prepared with only 5% of defective chains (see Supplementary Fig. 6 for details). These theoretical results were confirmed in a real system by quantifying the residual level of unreacted RAFT agent, corresponding to chains of length zero, after preparation of a series of oligomers of 4-acryloyl morpholine of average degree of polymerization 3–6 (see Supplementary Fig. 7 for details).
The validity of the Poisson distribution as a model for the number of units incorporated per chain was further demonstrated by adding a bismaleimide, bis(4-maleimidophenyl)methane, to the early stages of a RAFT polymerization of styrene (Fig. 4). The ratio of bismaleimide to RAFT agent was 1:2, such that on average one maleimide unit was added per chain. The polymerization was allowed to continue for 1 h, during which time the maleimide was entirely consumed. If exactly one maleimide had been added per chain, only the structure H shown in Fig. 4 would be obtained, with twice the molar mass of its constituent polystyrene chains. In fact, size exclusion chromatography (SEC) of the resulting polymer revealed single polymer chains (L), the linked structure (H) and aggregates of multiple chains (M). Deconvolution of the low molar mass portion of the chromatogram allowed the proportions of L and H species to be estimated at 34.4% and 15.1%, respectively. The expected proportions for a Poisson-distributed number of maleimides per chain are 36.8% L and 13.5% H. We have assumed that no looped structures are formed—a more detailed analysis taking this possibility into account can be found in Supplementary Note 3, but leads to a similar level of non-functionalized chains.
(a) After addition of the bismaleimide (0.5 equiv), polymerization was allowed to continue for 1 h, during which time the conversion of St increased from 17 to 21% and the bismaleimide was completely consumed. The resulting copolymer contained, on average, 1 maleimide unit per polystyrene chain (with an estimated standard deviation of 1). The distribution of maleimide units resulted in the formation of a mixture of linear chains (L), double chains (H) and multiply linked structures (M). (b) Distribution of chain lengths obtained after addition of 0.5 equiv. of bismaleimide. Linear chains and multiply linked structures are present in addition to H. The distribution immediately before bismaleimide addition is also shown. Deconvolution (c) of the low molar mass portion of the peak using Gaussian components (thick grey line represents best fit) allows estimation of the relative mass fractions of each structure (e), which are in good agreement with the predicted distribution assuming that the maleimide content of each chain follows a Poisson distribution (d).
This result conflicts with previous reports that only 10% of chains contain no maleimide when an average of 1 maleimide is incorporated per chain42,45. That value was obtained from a matrix-assisted laser desorption/ionization–time of flight mass spectrum in which polystyrene species containing 1 or 2 units of Bz-MI formed the most intense peaks45, while species containing zero or more than 3 units of Bz-MI accounted for 10% and 15% of the sequence distribution, respectively42. This implies that the average number of maleimides per chain is greater than 1, however, and so it seems likely that the frequency of zero-maleimide chains was underestimated. The incorporation of a polar maleimide unit may affect the ability of the polymer to form a charged complex, which can be observed by mass spectroscopy, biasing the results.
These results, applied to the SET-LRP multisite polymers of Fig. 3 (ref. 21), suggest that the yield of copolymer chains that contain at least one of each type of maleimide is only 25%, whereas 5% of chains contain no maleimide at all. Better results are obtained for the ROMP multisite polymer23 (Supplementary Fig. 4): the use of, on average, two functional monomers per site yields 56% of chains with all four types of functional monomer (the yield of chains with all nine blocks, including spacer blocks, is only 20%). A further increase in average block length to 3, as has been recommended by Lutz and co-workers42,46, raises the yield of chains containing all four functionalities to over 81%, whereas an average block length of 6 allows the formation of 20-block copolymers with 95% fidelity (Supplementary Fig. 6).
Discussion
The results presented in this communication assume the ideal case of Poisson-distributed polymer segments, an approximation that has been validated for anionic polymerization, but which does not hold for all controlled chain-growth polymerization techniques. Side reactions such as radical coupling and chain transfer will lead to broader distributions, as will the addition of more than one monomer during each activation–deactivation cycle in RDRP. In such cases, more realistic distributions could be obtained through the use of simulations, or using alternate distribution functions such as the negative binomial distribution which allow for greater variability. An example is given in Supplementary Note 4 for a multiblock polymer with overall dispersity of 1.1 (see also Supplementary Figs 10–14, Supplementary Table 3). The idealized results presented here are valuable because they show the maximum level of precision that can be obtained using chain polymerization, and provide indications of how that precision can be maximized.
We have quantified the maximum level of control over monomer placement that can be obtained using controlled chain growth polymerization techniques. The most ambitious goal of aperiodic polymer synthesis is precise control over the absolute position or separation of introduced monomers. We have shown that, regardless of the technique used, chain growth polymerization is poorly suited to this task, as uncertainty in the position of a given monomer grows in proportion to the square root of the length of the polymer chain. The less ambitious goal of controlling relative position with respect to the total chain length is more achievable, with improved control achieved for longer chains, and as the desired functionality approaches either chain end. Finally, if one desires simply that all blocks of a multiblock copolymer be present in a large majority of chains, this is possible so long as the individual blocks are of sufficient length. In all cases, we have given simple formulae for the probability distributions, expected values and standard deviations of the positions of inserted blocks.
Our research clearly shows the limitations of chain polymerization in the synthesis of very precisely controlled structures—the synthetic analogues of DNA or enzymes. Fortunately, the natural world provides numerous examples of less precisely controlled structures—proteins such as mucins, collagens and elastins—which play vital physiological roles. These proteins are characterized by complex yet variable structures that present multiple functionalities. In many cases, structural complexity enables self-assembly, generating fibrils or gels via a bottom-up process. It is these biomolecules that should provide the inspiration for precision chain polymerization.
Indeed, self-assembly of complex multiblock polymer architectures prepared via chain polymerization has already been used to develop biomimetic stress-stiffening gels47, and thermoresponsive, flower-like micelles which display multiple functionalities14. A recent review by Bates et al.48 highlights the enormous range of phase-separated structures that can be generated by polymers containing a relatively small number of blocks; faced with a bewildering array of possibilities, the challenge is to select the architecture which will give the desired properties. Having done so, it is equally important to know the precision with which that architecture can be prepared. This knowledge will allow the design of a new family of materials that offer an optimal balance of precision in the placement of functionalities and simplicity of production, for a wide range of chemical, biomedical and engineering applications.
Methods
Chemicals and reagents
Tetrahydrofuran (THF, Ajax Finechem, 99%), N,N-dimethylformamide (DMF, Merck, HPLC-grade), dimethyl 2,2′-azobis(2-methylpropanoate) (V-601), 2,2′-Azobis[2-(2-imidazolin-2-yl)propane]dihydrochloride (VA-044, Wako), 1,4-dioxane (Sigma-Aldrich, ≥99%) and 4-acryloylmorpholine (NAM, Sigma-Aldrich, 97%) were used without further purification. Styrene (Sigma-Aldrich, ≥99%) was filtered through a basic aluminium oxide (activated, basic, Brockmann I, standard grade, ∼150 mesh, 58A) column before use. All polymerizations were carried out under a nitrogen atmosphere. The RAFT agent, 2-(((butylthio)carbonothiolyl)thio)propanoic acid was prepared according to a reported procedure49.
NMR spectroscopy
1H NMR spectra were recorded on Bruker DPX-300 spectrometer using deuterated solvents obtained from Aldrich. Chemical shift values (δ) are reported in p.p.m. The residual proton signal of the solvent was used as internal standard.
Size exclusion chromatography
Number-average molar masses (Mn,SEC) and dispersity values (Ð) were determined using THF-SEC performed on an Agilent 390-MDS, comprising of an autosampler and a PLgel 5.0 μm bead-size guard column (50 × 7.5 mm2), followed by two linear 5.0 μm bead-size PLgel Mixed D columns (300 × 7.5 mm2) and a differential refractive index detector using THF (2% (v/v), TEA) as the eluent at 30 °C with a flow rate of 1 ml min−1. The SEC system was calibrated with linear PS EasiVial standards (Agilent Ltd.) ranging from 162 to 105 g mol−1. All samples were passed through 0.45 μm PTFE filter before SEC analysis. Molar masses were obtained by conventional calibration using ASTRA software.
RAFT polymerization of styrene with a controlled addition of bismaleimide
2-(((Butylthio)carbonothiolyl)thio)propanoic acid (114 mg, 0.480 mmol, 1 equiv.), styrene (2 g, 19.2 mmol, 40 equiv.), V-601 (11 mg, 0.048 mmol, 0.1 equiv.) and DMF (0.540 ml) were placed in a flask equipped with a magnetic stirring bar. The tube was then sealed with a rubber septum, degassed with nitrogen and then immersed in an oil bath thermostated at 65 °C. A degassed solution of bismaleimide (91 mg, 0.240 mmol, 0.5 equiv.) in 0.630 ml of DMF was added with a degassed syringe 3 h after the polymerization started (17% styrene conversion at this stage). After 1 more hour of polymerization, a sample was taken for NMR and THF-SEC analysis to ensure the full incorporation of the bismaleimide in the growing copolymer chain (around 4% of styrene conversion took place in this interval). The THF-SEC chromatogram shows multiple bismaleimide insertion.
RAFT homopolymerization of NAM
For a targeted degree of polymerization ( Xn) of 3. RAFT agent (281 mg, 1.181 mmol, 1 equiv.), NAM (0.5 g, 3.541 mmol, 3 equiv.), VA-044 (3.82 mg, 0.012 mmol, 0.01 equiv.), dioxane (0.477 ml) and H2O (0.257 ml) were placed in a flask equipped with a magnetic stirring bar. The tube was then sealed with a rubber septum, degassed with nitrogen and then immersed in an oil bath thermostated at 70 °C. After 2 h, a sample is withdrawn from the polymerization medium for 1H NMR (in dimethylsulphoxide (DMSO-d6)). Conversion was determined by 1H NMR to be 94%, corresponding to an  of 2.82.
of 2.82.
For a targeted Xn of 4. RAFT agent (211 mg, 0.885 mmol, 1 equiv.), NAM (0.5 g, 3.541 mmol, 4 equiv.), VA-044 (2.20 mg, 0.007 mmol, 0.0077, equiv.), dioxane (0.441 ml) and H2O (0.294 ml) were placed in a flask equipped with a magnetic stirring bar. The tube was then sealed with a rubber septum, degassed with nitrogen and then immersed in an oil bath thermostated at 70 °C. After 2 h, a sample was withdrawn from the polymerization medium for 1H NMR (in DMSO-d6). Conversion was determined by 1H NMR to be 97%, corresponding to an  of 3.89.
of 3.89.
For a targeted Xn of 5. RAFT agent (169 mg, 0.708 mmol, 1 equiv.), NAM (0.5 g, 3.541 mmol, 5 equiv.), VA-044 (1.53 mg, 0.005 mmol, 0.007 equiv.), dioxane (0.404 ml) and H2O (0.331 ml) were placed in a flask equipped with a magnetic stirring bar. The tube was then sealed with a rubber septum, degassed with nitrogen and then immersed in an oil bath thermostated at 70 °C. After 2 h, a sample is withdrawn from the polymerization medium for 1H NMR (in DMSO-d6). Conversion was determined by 1H NMR to be 98%, corresponding to an  of 4.92.
of 4.92.
For a targeted Xn of 6. RAFT agent (141 mg, 0.590 mmol, 1 equiv.), NAM (0.5 g, 3.541 mmol, 6 equiv.), VA-044 (0.95 mg, 0.003 mmol, 0.005 equiv.), dioxane (0.367 ml) and H2O (0.367 ml) were placed in a flask equipped with a magnetic stirring bar. The tube was then sealed with a rubber septum, degassed with nitrogen and then immersed in an oil bath thermostated at 70 °C. After 2 h, a sample is withdrawn from the polymerization medium for 1H NMR (in DMSO-d6). Conversion was determined by 1H NMR to be 99%, corresponding to an  of 5.92.
of 5.92.
Additional information
How to cite this article: Gody, G. et al. The limits of precision monomer placement in chain growth polymerization. Nat. Commun. 7:10514 doi: 10.1038/ncomms10514 (2016).
References
Lutz, J.-F. Aperiodic copolymers. ACS Macro Lett. 3, 1020–1023 (2014).
Schmidt, B. V. K. J. & Barner-Kowollik, C. Polymer chemistry: macromolecules made to order. Nat. Chem. 5, 990–992 (2013).
Lutz, J.-F., Ouchi, M., Liu, D. R. & Sawamoto, M. Sequence-controlled polymers. Science 341, 6146 (2013).
Lutz, J.-F. Sequence-controlled polymerizations: the next Holy Grail in polymer science? Polym. Chem. 1, 55–62 (2010).
Lutz, J.-F. Writing on polymer chains. Acc. Chem. Res. 46, 2696–2705 (2013).
Merrifield, R. B. Solid phase peptide synthesis. I. The synthesis of a tetrapeptide. J. Am. Chem. Soc. 85, 2149–2154 (1963).
Merrifield, R. B. Solid phase synthesis (nobel lecture). Angew. Chem. Int. Ed. 24, 799–810 (1985).
Pfeifer, S., Zarafshani, Z., Badi, N. & Lutz, J.-F. Liquid-phase synthesis of block copolymers containing sequence-ordered segments. J. Am. Chem. Soc. 131, 9195–9197 (2009).
Tong, X., Guo, B.-H. & Huang, Y. Toward the synthesis of sequence-controlled vinyl copolymers. Chem. Commun. 47, 1455–1457 (2011).
Feng, X.-S. & Pan, C.-Y. Synthesis of amphiphilic miktoarm ABC star copolymers by RAFT mechanism using maleic anhydride as linking agent. Macromolecules 35, 4888–4893 (2002).
Henry, S. M., Convertine, A. J., Benoit, D. S. W., Hoffman, A. S. & Stayton, P. S. End-Functionalized polymers and junction-functionalized diblock copolymers via RAFT chain extension with maleimido monomers. Bioconj. Chem. 20, 1122–1128 (2009).
Han, D.-H. & Pan, C.-Y. A novel strategy for synthesis of amphiphilic π-shaped copolymers by RAFT polymerization. Eur. Polym. J. 42, 507–515 (2006).
Zhou, C. et al. A well-defined amphiphilic polymer co-network from precise control of the end-functional groups of linear RAFT polymers. RSC Adv 4, 8144–8156 (2014).
Gody, G., Maschmeyer, T., Zetterlund, P. B. & Perrier, S. Rapid and quantitative one-pot synthesis of sequence-controlled polymers by radical polymerization. Nat. Commun. 4, 3505 (2013).
Fleury, G. & Bates, F. S. Structure and properties of hexa- and undecablock terpolymers with hierarchical molecular architectures. Macromolecules 42, 3598–3610 (2009).
Nagata, Y. et al. Preparation and characterization of a styrene−isoprene undecablock copolymer and its hierarchical microdomain structure in bulk. Macromolecules 38, 10220–10225 (2005).
Minoda, M., Sawamoto, M. & Higashimura, T. Sequence-regulated oligomers and polymers by living cationic polymerization. 2. Principle of sequence regulation and synthesis of sequence-regulated oligomers of functional vinyl ethers and styrene derivatives. Macromolecules 23, 4889–4895 (1990).
Soeriyadi, A. H., Boyer, C., Nyström, F., Zetterlund, P. B. & Whittaker, M. R. High-order multiblock copolymers via iterative Cu(0)-mediated radical polymerizations (SET-LRP): toward biological precision. J. Am. Chem. Soc. 133, 11128–11131 (2011).
Anastasaki, A. et al. Photoinduced sequence-control via one pot living radical polymerization of acrylates. Chem. Sci. 5, 3536–3542 (2014).
Fetsch, C. & Luxenhofer, R. Highly defined multiblock copolypeptoids: pushing the limits of living nucleophilic ring-opening polymerization. Macromol. Rapid Commun. 33, 1708–1713 (2012).
Zamfir, M. & Lutz, J.-F. Ultra-precise insertion of functional monomers in chain-growth polymerizations. Nat. Commun. 3, 1138 (2012).
Chan-Seng, D., Zamfir, M. & Lutz, J.-F. Polymer-chain encoding: synthesis of highly complex monomer sequence patterns by using automated protocols. Angew. Chem. Int. Ed. 51, 12254–12257 (2012).
Moatsou, D., Hansell, C. F. & O'Reilly, R. K. Precision polymers: a kinetic approach for functional poly(norbornenes). Chem. Sci. 5, 2246–2250 (2014).
Grubbs, R. & Tumas, W. Polymer synthesis and organotransition metal chemistry. Science 243, 907–915 (1989).
Jenkins, A. D., Jones, R. G. & Moad, G. Terminology for reversible-deactivation radical polymerization previously called ‘controlled’ radical or ‘living’ radical polymerization (IUPAC Recommendations 2010). Pure Appl. Chem. 82, 483 (2010).
Smid, J., Van Beylen, M. & Hogen-Esch, T. E. Perspectives on the contributions of Michael Szwarc to living polymerization. Prog. Polym. Sci. 31, 1041–1067 (2006).
Aoshima, S. & Kanaoka, S. A renaissance in living cationic polymerization. Chem. Rev. 109, 5245–5287 (2009).
Houshyar, S. et al. The scope for synthesis of macro-RAFT agents by sequential insertion of single monomer units. Polym. Chem. 3, 1879–1889 (2012).
Vandenbergh, J., Reekmans, G., Adriaensens, P. & Junkers, T. Synthesis of sequence controlled acrylate oligomers via consecutive RAFT monomer additions. Chem. Commun. 49, 10358–10360 (2013).
Flory, P. J. Molecular size distribution in ethylene oxide polymers. J. Am. Chem. Soc. 62, 1561–1565 (1940).
Lee, W. et al. Molecular Weight Distribution of Polystyrene Made by Anionic Polymerization. Macromolecules 33, 5111–5115 (2000).
Matyjaszewski, K. & Xia, J. Atom transfer radical polymerization. Chem. Rev. 101, 2921–2990 (2001).
Rosen, B. M. & Percec, V. Single-electron transfer and single-electron transfer degenerative chain transfer living radical polymerization. Chem. Rev. 109, 5069–5119 (2009).
Konkolewicz, D. et al. Reversible-deactivation radical polymerization in the presence of metallic copper. a critical assessment of the SARA ATRP and SET-LRP mechanisms. Macromolecules 46, 8749–8772 (2013).
Keddie, D. J. A guide to the synthesis of block copolymers using reversible-addition fragmentation chain transfer (RAFT) polymerization. Chem. Soc. Rev. 43, 496–505 (2014).
Hurtgen, M., Detrembleur, C., Jerome, C. & Debuigne, A. Insight into organometallic-mediated radical polymerization. Polym. Rev. 51, 188–213 (2011).
Yamago, S. Precision polymer synthesis by degenerative transfer controlled/living radical polymerization using organotellurium, organostibine, and organobismuthine chain-transfer agents. Chem. Rev. 109, 5051–5068 (2009).
Kamigaito, M., Ando, T. & Sawamoto, M. Metal-catalyzed living radical polymerization. Chem. Rev. 101, 3689–3746 (2001).
Hawker, C. J., Bosman, A. W. & Harth, E. New polymer synthesis by nitroxide mediated living radical polymerizations. Chem. Rev. 101, 3661–3688 (2001).
Altintas, O., Krolla-Sidenstein, P., Gliemann, H. & Barner-Kowollik, C. Single-chain folding of diblock copolymers driven by orthogonal h-donor and acceptor units. Macromolecules 47, 5877–5888 (2014).
Altintas, O., Lejeune, E., Gerstel, P. & Barner-Kowollik, C. Bioinspired dual self-folding of single polymer chains via reversible hydrogen bonding. Polym. Chem. 3, 640–651 (2012).
Lutz, J.-F., Schmidt, B. V. K. J. & Pfeifer, S. Tailored polymer microstructures prepared by atom transfer radical copolymerization of styrene and N-substituted maleimides. Macromol. Rapid Commun. 32, 127–135 (2011).
Falls, L. W. The beta distribution: a statistical model for world cloud cover. J. Geophys. Res. 79, 1261–1264 (1974).
Wright, S. Evolution in mendelian populations. Genetics 16, 97–159 (1931).
Pfeifer, S. & Lutz, J.-F. Development of a library of N-substituted maleimides for the local functionalization of linear polymer chains. Chem. Eur. J 14, 10949–10957 (2008).
Pfeifer, S. & Lutz, J.-F. A facile procedure for controlling monomer sequence distribution in radical chain polymerizations. J. Am. Chem. Soc. 129, 9542–9543 (2007).
Kouwer, P. H. J. et al. Responsive biomimetic networks from polyisocyanopeptide hydrogels. Nature 493, 651–655 (2013).
Bates, F. S. et al. Multiblock polymers: Panacea or Pandora’s Box? Science 336, 434–440 (2012).
Ferguson, C. J. et al. Ab initio emulsion polymerization by RAFT-controlled self-assembly. Macromolecules 38, 2191–2204 (2005).
Acknowledgements
We thank M. Destarac for his comments on an early version of the manuscript. The Centre National de Recherche Scientifique (S.H.), the Royal Society Wolfson Merit Award (WM130055; S.P.), the Monash-Warwick Alliance (G.G., S.P.) and the Australian Research Council (Future Fellowship, P.B.Z.) are acknowledged for financial support.
Author information
Authors and Affiliations
Contributions
S.H. developed the model and wrote the manuscript. G.G. and S.P. designed the experiments, which were conducted by G.G. Interpretation of the results was carried out by G.G., S.P., P.B.Z. and S.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-14, Supplementary Tables 1-3, Supplementary Notes 1-4 and Supplementary References (PDF 1280 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Gody, G., Zetterlund, P., Perrier, S. et al. The limits of precision monomer placement in chain growth polymerization. Nat Commun 7, 10514 (2016). https://doi.org/10.1038/ncomms10514
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms10514
This article is cited by
-
Acid-triggered radical polymerization of vinyl monomers
Nature Synthesis (2024)
-
Concurrent control over sequence and dispersity in multiblock copolymers
Nature Chemistry (2022)
-
A comparison of RAFT and ATRP methods for controlled radical polymerization
Nature Reviews Chemistry (2021)
-
Visualization and design of the functional group distribution during statistical copolymerization
Nature Communications (2019)
-
Encoding information into polymers
Nature Reviews Chemistry (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.