Abstract
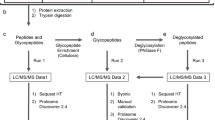
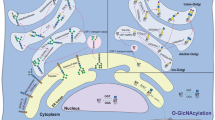
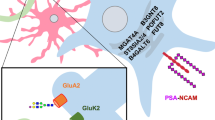
The addition of the monosaccharide β-N-acetyl-D-glucosamine to proteins (O-GlcNAc glycosylation) is an intracellular, post-translational modification that shares features with phosphorylation. Understanding the cellular mechanisms and signaling pathways that regulate O-GlcNAc glycosylation has been challenging because of the difficulty of detecting and quantifying the modification. Here, we describe a new strategy for monitoring the dynamics of O-GlcNAc glycosylation using quantitative mass spectrometry-based proteomics. Our method, which we have termed quantitative isotopic and chemoenzymatic tagging (QUIC-Tag), combines selective, chemoenzymatic tagging of O-GlcNAc proteins with an efficient isotopic labeling strategy. Using the method, we detect changes in O-GlcNAc glycosylation on several proteins involved in the regulation of transcription and mRNA translocation. We also provide the first evidence that O-GlcNAc glycosylation is dynamically modulated by excitatory stimulation of the brain in vivo. Finally, we use electron-transfer dissociation mass spectrometry to identify exact sites of O-GlcNAc modification. Together, our studies suggest that O-GlcNAc glycosylation occurs reversibly in neurons and, akin to phosphorylation, may have important roles in mediating the communication between neurons.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Khidekel, N. & Hsieh-Wilson, L.C. A 'molecular switchboard'–covalent modifications to proteins and their impact on transcription. Org. Biomol. Chem. 2, 1–7 (2004).
Greengard, P. The neurobiology of slow synaptic transmission. Science 294, 1024–1030 (2001).
Love, D.C. & Hanover, J.A. The hexosamine signaling pathway: deciphering the “O-GlcNAc code”. Sci. STKE 2005, re13 (2005).
Khidekel, N., Ficarro, S.B., Peters, E.C. & Hsieh-Wilson, L.C. Exploring the O-GlcNAc proteome: direct identification of O-GlcNAc-modified proteins from the brain. Proc. Natl. Acad. Sci. USA 101, 13132–13137 (2004).
Vosseller, K. et al. O-linked N-acetylglucosamine proteomics of postsynaptic density preparations using lectin weak affinity chromatography and mass spectrometry. Mol. Cell. Proteomics 5, 923–934 (2006).
Kneass, Z.T. & Marchase, R.B. Neutrophils exhibit rapid agonist-induced increases in protein-associated O-GlcNAc. J. Biol. Chem. 279, 45759–45765 (2004).
Zachara, N.E. et al. Dynamic O-GlcNAc modification of nucleocytoplasmic proteins in response to stress. A survival response of mammalian cells. J. Biol. Chem. 279, 30133–30142 (2004).
Cheng, X. & Hart, G.W. Alternative O-glycosylation/O-phosphorylation of serine-16 in murine estrogen receptor beta: post-translational regulation of turnover and transactivation activity. J. Biol. Chem. 276, 10570–10575 (2001).
Chou, T.Y., Hart, G.W. & Dang, C.V. c-Myc is glycosylated at threonine 58, a known phosphorylation site and a mutational hot spot in lymphomas. J. Biol. Chem. 270, 18961–18965 (1995).
Iyer, S.P. & Hart, G.W. Dynamic nuclear and cytoplasmic glycosylation: enzymes of O-GlcNAc cycling. Biochemistry 42, 2493–2499 (2003).
Cole, R.N. & Hart, G.W. Cytosolic O-glycosylation is abundant in nerve terminals. J. Neurochem. 79, 1080–1089 (2001).
Iyer, S.P.N. & Hart, G.W. Dynamic nuclear and cytoplasmic glycosylation: enzymes of O- GlcNAc cycling. Biochemistry 42, 2493–2499 (2003).
O'Donnell, N., Zachara, N.E., Hart, G.W. & Marth, J.D. Ogt-dependent X-chromosome-linked protein glycosylation is a requisite modification in somatic cell function and embryo viability. Mol. Cell. Biol. 24, 1680–1690 (2004).
Lamarre-Vincent, N. & Hsieh-Wilson, L.C. Dynamic glycosylation of the transcription factor CREB: a potential role in gene regulation. J. Am. Chem. Soc. 125, 6612–6613 (2003).
Griffith, L.S. & Schmitz, B. O-linked N-acetylglucosamine levels in cerebellar neurons respond reciprocally to perturbations of phosphorylation. Eur. J. Biochem. 262, 824–831 (1999).
Khidekel, N. et al. A chemoenzymatic approach toward the rapid and sensitive detection of O-GlcNAc posttranslational modifications. J. Am. Chem. Soc. 125, 16162–16163 (2003).
Ong, S.E., Mittler, G. & Mann, M. Identifying and quantifying in vivo methylation sites by heavy methyl SILAC. Nat. Methods 1, 119–126 (2004).
Ross, P.L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Tai, H.C., Khidekel, N., Ficarro, S.B., Peters, E.C. & Hsieh-Wilson, L.C. Parallel identification of O-GlcNAc-modified proteins from cell lysates. J. Am. Chem. Soc. 126, 10500–10501 (2004).
Cieniewski-Bernard, C. et al. Identification of O-linked N-acetylglucosamine proteins in rat skeletal muscle using two-dimensional gel electrophoresis and mass spectrometry. Mol. Cell. Proteomics 3, 577–585 (2004).
Hsu, J.L., Huang, S.Y., Chow, N.H. & Chen, S.H. Stable-isotope dimethyl labeling for quantitative proteomics. Anal. Chem. 75, 6843–6852 (2003).
Chalkley, R.J. & Burlingame, A.L. Identification of GlcNAcylation sites of peptides and alpha-crystallin using Q-TOF mass spectrometry. J. Am. Soc. Mass Spectrom. 12, 1106–1113 (2001).
Makarov, A., Denisov, E., Lange, O. & Horning, S. Dynamic range of mass accuracy in LTQ orbitrap hybrid mass spectrometer. J. Am. Soc. Mass Spectrom. 17, 977–982 (2006).
Roquemore, E.P., Chevrier, M.R., Cotter, R.J. & Hart, G.W. Dynamic O-GlcNAcylation of the small heat shock protein alpha B-crystallin. Biochemistry 35, 3578–3586 (1996).
Zhang, R., Sioma, C.S., Wang, S. & Regnier, F.E. Fractionation of isotopically labeled peptides in quantitative proteomics. Anal. Chem. 73, 5142–5149 (2001).
Haltiwanger, R.S., Grove, K. & Philipsberg, G.A. Modulation of O-linked N-acetylglucosamine levels on nuclear and cytoplasmic proteins in vivo using the peptide O-GlcNAc-beta-N-acetylglucosaminidase inhibitor O-(2-acetamido-2-deoxy-D-glucopyranosylidene)amino-N-phenylcarbamate. J. Biol. Chem. 273, 3611–3617 (1998).
Syka, J.E., Coon, J.J., Schroeder, M.J., Shabanowitz, J. & Hunt, D.F. Peptide and protein sequence analysis by electron transfer dissociation mass spectrometry. Proc. Natl. Acad. Sci. USA 101, 9528–9533 (2004).
Coon, J.J., Syka, J.E.P., Schwartz, J.C., Shabanowitz, J. & Hunt, D.F. Anion dependence in the partitioning between proton and electron transfer in ion/ion reactions. Int. J. Mass Spectrom. 236, 33–42 (2004).
Hogan, J.M., Pitteri, S.J., Chrisman, P.A. & McLuckey, S.A. Complementary structural information from a tryptic N-linked glycopeptide via electron transfer ion/ion reactions and collision-induced dissociation. J. Proteome Res. 4, 628–632 (2005).
Swaney, D.L. et al. Supplemental activation method for high-efficiency electron-transfer dissociation of doubly protonated peptide precursors. Anal. Chem. 79, 477–485 (2007).
Lazarus, B.D., Love, D.C. & Hanover, J.A. Recombinant O-GlcNAc transferase isoforms: identification of O-GlcNAcase, yes tyrosine kinase, and tau as isoform-specific substrates. Glycobiology 16, 415–421 (2006).
Whisenhunt, T.R. et al. Disrupting the enzyme complex regulating O-GlcNAcylation blocks signaling and development. Glycobiology 16, 551–563 (2006).
Nedivi, E., Hevroni, D., Naot, D., Israeli, D. & Citri, Y. Numerous candidate plasticity-related genes revealed by differential cDNA cloning. Nature 363, 718–722 (1993).
Ben-Ari, Y. & Cossart, R. Kainate, a double agent that generates seizures: two decades of progress. Trends Neurosci. 23, 580–587 (2000).
Beckmann, A.M., Davidson, M.S., Goodenough, S. & Wilce, P.A. Differential expression of Egr-1-like DNA-binding activities in the naive rat brain and after excitatory stimulation. J. Neurochem. 69, 2227–2237 (1997).
Nandi, A. et al. Global identification of O-GlcNAc-modified proteins. Anal. Chem. 78, 452–458 (2006).
Kamemura, K., Hayes, B.K., Comer, F.I. & Hart, G.W. Dynamic interplay between O-glycosylation and O-phosphorylation of nucleocytoplasmic proteins: alternative glycosylation/phosphorylation of THR-58, a known mutational hot spot of c-Myc in lymphomas, is regulated by mitogens. J. Biol. Chem. 277, 19229–19235 (2002).
Ludemann, N. et al. O-glycosylation of the tail domain of neurofilament protein M in human neurons and in spinal cord tissue of a rat model of amyotrophic lateral sclerosis (ALS). J. Biol. Chem. 280, 31648–31658 (2005).
Vosseller, K. et al. Quantitative analysis of both protein expression and serine/ threonine post-translational modifications through stable isotope labeling with dithiothreitol. Proteomics 5, 388–398 (2005).
Brackertz, M., Gong, Z., Leers, J. & Renkawitz, R. p66alpha and p66beta of the Mi-2/NuRD complex mediate MBD2 and histone interaction. Nucleic Acids Res. 34, 397–406 (2006).
Yang, X. et al. O-linkage of N-acetylglucosamine to Sp1 activation domain inhibits its transcriptional capability. Proc. Natl. Acad. Sci. USA 98, 6611–6616 (2001).
Gong, Z., Brackertz, M. & Renkawitz, R. SUMO modification enhances p66-mediated transcriptional repression of the Mi-2/NuRD complex. Mol. Cell. Biol. 26, 4519–4528 (2006).
Ule, J. & Darnell, R.B. RNA binding proteins and the regulation of neuronal synaptic plasticity. Curr. Opin. Neurobiol. 16, 102–110 (2006).
Elvira, G., Massie, B. & DesGroseillers, L. The zinc-finger protein ZFR is critical for Staufen 2 isoform specific nucleocytoplasmic shuttling in neurons. J. Neurochem. 96, 105–117 (2006).
Bastos, R., Lin, A., Enarson, M. & Burke, B. Targeting and function in mRNA export of nuclear pore complex protein Nup153. J. Cell Biol. 134, 1141–1156 (1996).
Jones, M.W. et al. A requirement for the immediate early gene Zif268 in the expression of late LTP and long-term memories. Nat. Neurosci. 4, 289–296 (2001).
Thiel, G. & Cibelli, G. Regulation of life and death by the zinc finger transcription factor Egr-1. J. Cell. Physiol. 193, 287–292 (2002).
James, A.B., Conway, A.M. & Morris, B.J. Genomic profiling of the neuronal target genes of the plasticity-related transcription factor – Zif268. J. Neurochem. 95, 796–810 (2005).
Wang, Q., Yu, S., Simonyi, A., Sun, G.Y. & Sun, A.Y. Kainic acid-mediated excitotoxicity as a model for neurodegeneration. Mol. Neurobiol. 31, 3–16 (2005).
Marin, P. et al. Glutamate-dependent phosphorylation of elongation factor-2 and inhibition of protein synthesis in neurons. J. Neurosci. 17, 3445–3454 (1997).
Datta, R., Choudhury, P., Ghosh, A. & Datta, B. A glycosylation site, 60SGTS63, of p67 is required for its ability to regulate the phosphorylation and activity of eukaryotic initiation factor 2α. Biochemistry 42, 5453–5460 (2003).
Collins, M.O. et al. Proteomic analysis of in vivo phosphorylated synaptic proteins. J. Biol. Chem. 280, 5972–5982 (2005).
Gu, Y., Hamajima, N. & Ihara, Y. Neurofibrillary tangle-associated collapsin response mediator protein-2 (CRMP-2) is highly phosphorylated on Thr-509, Ser-518, and Ser-522. Biochemistry 39, 4267–4275 (2000).
Liu, F., Iqbal, K., Grundke-Iqbal, I., Hart, G.W. & Gong, C.X. O-GlcNAcylation regulates phosphorylation of tau: a mechanism involved in Alzheimer's disease. Proc. Natl. Acad. Sci. USA 101, 10804–10809 (2004).
Gama, C.I. et al. Sulfation patterns of glycosaminoglycans encode molecular recognition and activity. Nat. Chem. Biol. 2, 467–473 (2006).
Acknowledgements
We thank P. Qasba and B. Ramakrishnan for the generous gift of the GalT plasmid, S. Whiteheart for the OGA antibody, T.C. Neo for assistance with synthesis of ketogalactose probe 1, and A. Su for technical discussions. This work was supported by the US National Institutes of Health (RO1 NS045061), the National Science Foundation CAREER Award (CHE-0239861) and the Parson's Foundation (N.K.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Expression levels of EGR-1, GRASP55 and eIF4G following kainic acid treatment of rats. (PDF 22 kb)
Supplementary Fig. 2
Annotated CAD MS4 and ETD MS/MS mass spectra for all sequenced peptides. (PDF 1955 kb)
Supplementary Table 1
Mean ratios of individual peptides from α-crystallin and OGT, and mean ratios of all peptides. (PDF 85 kb)
Supplementary Table 2
Identification and quantification of changes in O-GlcNAc glycosylation induced by kainic acid. (PDF 9 kb)
Supplementary Table 3
O-GlcNAc glycosylated proteins identified from the cerebral cortex of kainic acid-stimulated rats. (PDF 10 kb)
Rights and permissions
About this article
Cite this article
Khidekel, N., Ficarro, S., Clark, P. et al. Probing the dynamics of O-GlcNAc glycosylation in the brain using quantitative proteomics. Nat Chem Biol 3, 339–348 (2007). https://doi.org/10.1038/nchembio881
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio881
This article is cited by
-
Parallel pathways for serotonin biosynthesis and metabolism in C. elegans
Nature Chemical Biology (2023)
-
Loss of O-GlcNAcylation on MeCP2 at Threonine 203 Leads to Neurodevelopmental Disorders
Neuroscience Bulletin (2022)
-
Hexosamine biosynthetic pathway and O-GlcNAc-processing enzymes regulate daily rhythms in protein O-GlcNAcylation
Nature Communications (2021)
-
PKCγ-Mediated Phosphorylation of CRMP2 Regulates Dendritic Outgrowth in Cerebellar Purkinje Cells
Molecular Neurobiology (2020)
-
Fueling the fire: emerging role of the hexosamine biosynthetic pathway in cancer
BMC Biology (2019)