Abstract
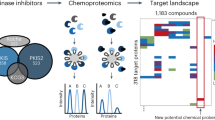
We devised a high-throughput chemoproteomics method that enabled multiplexed screening of 16,000 compounds against native protein and lipid kinases in cell extracts. Optimization of one chemical series resulted in CZC24832, which is to our knowledge the first selective inhibitor of phosphoinositide 3-kinase γ (PI3Kγ) with efficacy in in vitro and in vivo models of inflammation. Extensive target- and cell-based profiling of CZC24832 revealed regulation of interleukin-17–producing T helper cell (TH17) differentiation by PI3Kγ, thus reinforcing selective inhibition of PI3Kγ as a potential treatment for inflammatory and autoimmune diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Change history
16 May 2012
In the version of this article initially published, the name M. Sunose was misspelled in the Acknowledgements. The error has been corrected in the HTML and PDF versions of the article.
References
Vanhaesebroeck, B., Guillermet-Guibert, J., Graupera, M. & Bilanges, B. The emerging mechanisms of isoform-specific PI3K signalling. Nat. Rev. Mol. Cell Biol. 11, 329–341 (2010).
Hawkins, P.T., Anderson, K.E., Davidson, K. & Stephens, L.R. Signalling through Class I PI3Ks in mammalian cells. Biochem. Soc. Trans. 34, 647–662 (2006).
Bohnacker, T. et al. PI3Kγ adaptor subunits define coupling to degranulation and cell motility by distinct PtdIns(3,4,5)P3 pools in mast cells. Sci. Signal. 2, ra27 (2009).
Ghigo, A., Damilano, F., Braccini, L. & Hirsch, E. PI3K inhibition in inflammation: toward tailored therapies for specific diseases. Bioessays 32, 185–196 (2010).
Foukas, L.C. et al. Critical role for the p110α phosphoinositide-3-OH kinase in growth and metabolic regulation. Nature 441, 366–370 (2006).
Ji, H. et al. Inactivation of PI3Kγ and PI3Kδ distorts T-cell development and causes multiple organ inflammation. Blood 110, 2940–2947 (2007).
Ciraolo, E. et al. Phosphoinositide 3-kinase p110β activity: key role in metabolism and mammary gland cancer but not development. Sci. Signal. 1, ra3 (2008).
Maira, S.M. et al. Identification and characterization of NVP-BEZ235, a new orally available dual phosphatidylinositol 3-kinase/mammalian target of rapamycin inhibitor with potent in vivo antitumor activity. Mol. Cancer Ther. 7, 1851–1863 (2008).
Brachmann, S.M. et al. Specific apoptosis induction by the dual PI3K/mTor inhibitor NVP-BEZ235 in HER2 amplified and PIK3CA mutant breast cancer cells. Proc. Natl. Acad. Sci. USA 106, 22299–22304 (2009).
Konstantinidou, G. et al. Dual phosphoinositide 3-kinase/mammalian target of rapamycin blockade is an effective radiosensitizing strategy for the treatment of non-small cell lung cancer harboring K-RAS mutations. Cancer Res. 69, 7644–7652 (2009).
Toledo, L.I. et al. A cell-based screen identifies ATR inhibitors with synthetic lethal properties for cancer-associated mutations. Nat. Struct. Mol. Biol. 18, 721–727 (2011).
Bantscheff, M. et al. Quantitative chemical proteomics reveals mechanisms of action of clinical ABL kinase inhibitors. Nat. Biotechnol. 25, 1035–1044 (2007).
Bantscheff, M. et al. Chemoproteomics profiling of HDAC inhibitors reveals selective targeting of HDAC complexes. Nat. Biotechnol. 29, 255–265 (2011).
Kruse, U. et al. Chemoproteomics-based kinome profiling and target deconvolution of clinical multi-kinase inhibitors in primary chronic lymphocytic leukemia cells. Leukemia 25, 89–100 (2011).
Knight, Z.A. et al. A pharmacological map of the PI3-K family defines a role for p110α in insulin signaling. Cell 125, 733–747 (2006).
Vlahos, C.J., Matter, W.F., Hui, K.Y. & Brown, R.F. A specific inhibitor of phosphatidylinositol 3-kinase, 2-(4-morpholinyl)-8-phenyl-4H–1-benzopyran-4-one (LY294002). J. Biol. Chem. 269, 5241–5248 (1994).
Cansfield, A., Bergamini, G. & Neubauer, G. Selectivity profiling of PI3K interacting molecules against multiple targets. European patent EP2245181 (2011).
Jefferies, H.B. et al. A selective PIKfyve inhibitor blocks PtdIns(3,5)P(2) production and disrupts endomembrane transport and retroviral budding. EMBO Rep. 9, 164–170 (2008).
Sharma, K. et al. Proteomics strategy for quantitative protein interaction profiling in cell extracts. Nat. Methods 6, 741–744 (2009).
Berg, E.L. et al. Chemical target and pathway toxicity mechanisms defined in primary human cell systems. J. Pharmacol. Toxicol. Methods 61, 3–15 (2010).
Hirsch, E. et al. Central role for G protein-coupled phosphoinositide 3-kinase γ in inflammation. Science 287, 1049–1053 (2000).
Jones, G.E. et al. Requirement for PI 3-kinase γ in macrophage migration to MCP-1 and CSF-1. Exp. Cell Res. 290, 120–131 (2003).
Savitski, M.M. et al. Targeted data acquisition for improved reproducibility and robustness of proteomic mass spectrometry assays. J. Am. Soc. Mass Spectrom. 21, 1668–1679 (2010).
Williams, O. et al. Discovery of dual inhibitors of the immune cell PI3Ks p110δ and p110γ: a prototype for new anti-inflammatory drugs. Chem. Biol. 17, 123–134 (2010).
Mitsdoerffer, M. et al. Proinflammatory T helper type 17 cells are effective B-cell helpers. Proc. Natl. Acad. Sci. USA 107, 14292–14297 (2010).
Zhou, L. et al. IL-6 programs TH-17 cell differentiation by promoting sequential engagement of the IL-21 and IL-23 pathways. Nat. Immunol. 8, 967–974 (2007).
Patrucco, E. et al. PI3Kγ modulates the cardiac response to chronic pressure overload by distinct kinase-dependent and -independent effects. Cell 118, 375–387 (2004).
Rommel, C., Camps, M. & Ji, H. PI3K δ and PI3K γ: partners in crime in inflammation in rheumatoid arthritis and beyond? Nat. Rev. Immunol. 7, 191–201 (2007).
Rückle, T., Schwarz, M.K. & Rommel, C. PI3Kγ inhibition: towards an 'aspirin of the 21st century'? Nat. Rev. Drug Discov. 5, 903–918 (2006).
Martin, D. et al. PI3Kγ mediates Kaposi's sarcoma–associated herpesvirus vGPCR-induced sarcomagenesis. Cancer Cell 19, 805–813 (2011).
Schmid, M.C. et al. Receptor tyrosine kinases and TLR/IL1Rs unexpectedly activate myeloid cell PI3Kγ, a single convergent point promoting tumor inflammation and progression. Cancer Cell 19, 715–727 (2011).
Becattini, B. et al. PI3Kγ within a nonhematopoietic cell type negatively regulates diet-induced thermogenesis and promotes obesity and insulin resistance. Proc. Natl. Acad. Sci. USA 108, E854–E863 (2011).
Kobayashi, N. et al. Blockade of class IB phosphoinositide-3 kinase ameliorates obesity-induced inflammation and insulin resistance. Proc. Natl. Acad. Sci. USA 108, 5753–5758 (2011).
Fougerat, A. et al. Genetic and pharmacological targeting of phosphoinositide 3-kinase-γ reduces atherosclerosis and favors plaque stability by modulating inflammatory processes. Circulation 117, 1310–1317 (2008).
Fadden, P. et al. Application of chemoproteomics to drug discovery: identification of a clinical candidate targeting hsp90. Chem. Biol. 17, 686–694 (2010).
Graves, P.R. et al. Discovery of novel targets of quinoline drugs in the human purine binding proteome. Mol. Pharmacol. 62, 1364–1372 (2002).
Camps, M. et al. Blockade of PI3Kγ suppresses joint inflammation and damage in mouse models of rheumatoid arthritis. Nat. Med. 11, 936–943 (2005).
Bilancio, A. et al. Key role of the p110δ isoform of PI3K in B-cell antigen and IL-4 receptor signaling: comparative analysis of genetic and pharmacologic interference with p110δ function in B cells. Blood 107, 642–650 (2006).
Condliffe, A.M. et al. Sequential activation of class IB and class IA PI3K is important for the primed respiratory burst of human but not murine neutrophils. Blood 106, 1432–1440 (2005).
Vecchione, C. et al. Protection from angiotensin II–mediated vasculotoxic and hypertensive response in mice lacking PI3Kγ. J. Exp. Med. 201, 1217–1228 (2005).
Haylock-Jacobs, S. et al. PI3Kδ drives the pathogenesis of experimental autoimmune encephalomyelitis by inhibiting effector T cell apoptosis and promoting TH17 differentiation. J. Autoimmun. 36, 278–287 (2011).
Konrad, S. et al. Phosphoinositide 3-kinases γ and δ, linkers of coordinate C5a receptor-Fcγ receptor activation and immune complex-induced inflammation. J. Biol. Chem. 283, 33296–33303 (2008).
Lubberts, E. et al. Treatment with a neutralizing anti–murine interleukin-17 antibody after the onset of collagen-induced arthritis reduces joint inflammation, cartilage destruction, and bone erosion. Arthritis Rheum. 50, 650–659 (2004).
Genovese, M.C. et al. LY2439821, a humanized anti–interleukin-17 monoclonal antibody, in the treatment of patients with rheumatoid arthritis: a phase I randomized, double-blind, placebo-controlled, proof-of-concept study. Arthritis Rheum. 62, 929–939 (2010).
Kageyama, Y., Kobayashi, H. & Kato, N. Infliximab treatment reduces the serum levels of interleukin-23 in patients with rheumatoid arthritis. Mod. Rheumatol. 19, 657–662 (2009).
Savitski, M.M. et al. Delayed fragmentation and optimized isolation width settings for improvement of protein identification and accuracy of isobaric mass tag quantification on Orbitrap-type mass spectrometers. Anal. Chem. 83, 8959–8967 (2011).
Savitski, M.M., Scholten, A., Sweetman, G., Mathieson, T. & Bantscheff, M. Evaluation of data analysis strategies for improved mass spectrometry-based phosphoproteomics. Anal. Chem. 82, 9843–9849 (2010).
Kunkel, E.J. et al. An integrative biology approach for analysis of drug action in models of human vascular inflammation. FASEB J. 18, 1279–1281 (2004).
Kunkel, E.J. et al. Rapid structure-activity and selectivity analysis of kinase inhibitors by BioMAP analysis in complex human primary cell-based models. Assay Drug Dev. Technol. 2, 431–441 (2004).
Acknowledgements
We are grateful to M. Sunhose and W. Miller for chemistry and cheminformatics support; MS, information technology, biochemistry and biology groups for technical support; G. Creighton-Gutteridge for experimental support; F. Weisbrodt for assistance with graphics; G. Bennett (Cellzome Ltd.) and O. Azzolino (University of Torino) for animal data; Y. Abraham for generation of the cladogram; R. Hale and T. Edwards for general advice; and M. Wymann, D. Simmons and A. Watt for stimulating discussions and valuable comments.
Author information
Authors and Affiliations
Contributions
G.B., S.S., K.M. and J.P. developed and performed assays; K.B., A.C. and K.E. designed and synthesized compounds; C.R., F. Reinhard, D.L. and R.M. performed screening; F. Rharbaoui contributed to animal studies; M.M.S., T.M. and M.B. developed MS technology; C.D. supported data analysis and display; A.O. and I.P. contributed cellular activity profiles; T.W. designed and performed MS experiments; C.H. supervised biochemistry work; G.D. contributed to the manuscript and gave advice; E.H. gave general advice and contributed animal data; N.R. led the chemistry efforts; O.R. contributed to data analysis and manuscript preparation; and G.B., M.B. and G.N. designed and supervised the study, analyzed data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.B., S.S., T.W., K.M., J.P., C.R., A.H., C.D., T.M., F. Rharbaoui, F. Reinhard, M.M.S., G.D., M.B. and G.N. are employees of Cellzome AG; K.B., A.C., K.E., D.L., R.M., N.R. and O.R. are employees of Cellzome Ltd.; and A.M. and I.P. are employees of BioSeek Inc.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 11964 kb)
Supplementary Data Set 1
Complementary kinase capturing of CZC15098 and CZC15292 derived bead matrices in different cell extracts (XLSX 256 kb)
Supplementary Data Set 2
Concentration-inhibition profiles for all reference compounds and CZC19945 and CZC24832 in different cell extracts (XLSX 902 kb)
Supplementary Data Set 3
Correction factors for calculation of apparent dissociation constants from IC50s obtained in profiling experiments using kinobeads and HL60 cell extracts (XLSX 175 kb)
Supplementary Data Set 4
Correction factors for calculation of apparent dissociation constants from IC50s obtained in profiling experiments using LK beads and HL60 cell extracts (XLSX 163 kb)
Supplementary Data Set 5
Proteomic binding profiles using an immobilized linkable analogue of CZC19945/CZC24832 as probe matrix and HeLa, HL60, and PBMC cell extracts (XLSX 175 kb)
Supplementary Data Set 6
Correction factors for calculation of apparent dissociation constants from IC50s obtained in profiling experiments using LK beads and mixed raw264.7, HL60 cell extracts (XLSX 163 kb)
Supplementary Data Set 7
Correction factors for calculation of apparent dissociation constants from IC50s obtained in profiling experiments using LK beads and mixed RBL-1, HL60 cell extracts (XLSX 163 kb)
Rights and permissions
About this article
Cite this article
Bergamini, G., Bell, K., Shimamura, S. et al. A selective inhibitor reveals PI3Kγ dependence of TH17 cell differentiation. Nat Chem Biol 8, 576–582 (2012). https://doi.org/10.1038/nchembio.957
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.957
This article is cited by
-
PI3Kγ controls IL-17A expression and attenuates alveolar bone loss in an experimental periodontitis model
Inflammation Research (2023)
-
The role of PI3Kγ in the immune system: new insights and translational implications
Nature Reviews Immunology (2022)
-
The emerging role of mass spectrometry-based proteomics in drug discovery
Nature Reviews Drug Discovery (2022)
-
Chemical proteomics reveals target selectivity of clinical Jak inhibitors in human primary cells
Scientific Reports (2019)
-
High-throughput screening of the Plasmodium falciparum cGMP-dependent protein kinase identified a thiazole scaffold which kills erythrocytic and sexual stage parasites
Scientific Reports (2019)