Abstract
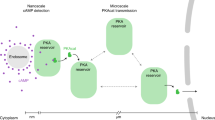
Understanding how specific cyclic AMP (cAMP) signals are organized and relayed to their effectors in different compartments of the cell to achieve functional specificity requires molecular tools that allow precise manipulation of cAMP in these compartments. Here we characterize a new method using bicarbonate-activatable and genetically targetable soluble adenylyl cyclase to control the location, kinetics and magnitude of the cAMP signal. Using this live-cell cAMP manipulation in conjunction with fluorescence imaging and mechanistic modeling, we uncovered the activation of a resident pool of protein kinase A (PKA) holoenzyme in the nuclei of HEK-293 cells, modifying the existing dogma of cAMP-PKA signaling in the nucleus. Furthermore, we show that phosphodiesterases and A-kinase anchoring proteins (AKAPs) are critical in shaping nuclear PKA responses. Collectively, our data suggest a new model in which AKAP-localized phosphodiesterases tune an activation threshold for nuclear PKA holoenzyme, thereby converting spatially distinct second messenger signals to temporally controlled nuclear kinase activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
19 April 2013
In the version of this article initially published, the domain schemes in Figure 1a were incorrectly labeled sACt (aa 1–146). The correct text is sACt (aa 1–469). The error has been corrected in the HTML and PDF versions of the article.
References
Taskén, K. & Aandahl, E.M. Localized effects of cAMP mediated by distinct routes of protein kinase A. Physiol. Rev. 84, 137–167 (2004).
Steinberg, S.F. & Brunton, L.L. Compartmentation of G protein-coupled signaling pathways in cardiac myocytes. Annu. Rev. Pharmacol. Toxicol. 41, 751–773 (2001).
Houslay, M.D. Underpinning compartmentalised cAMP signalling through targeted cAMP breakdown. Trends Biochem. Sci. 35, 91–100 (2010).
Carnegie, G.K., Means, C.K. & Scott, J.D. A-kinase anchoring proteins: from protein complexes to physiology and disease. IUBMB Life 61, 394–406 (2009).
Altarejos, J.Y. & Montminy, M. CREB and the CRTC co-activators: sensors for hormonal and metabolic signals. Nat. Rev. Mol. Cell Biol. 12, 141–151 (2011).
Kvissel, A.K. et al. Involvement of the catalytic subunit of protein kinase A and of HA95 in pre-mRNA splicing. Exp. Cell Res. 313, 2795–2809 (2007).
Martin, B.R., Deerinck, T.J., Ellisman, M.H., Taylor, S.S. & Tsien, R.Y. Isoform-specific PKA dynamics revealed by dye-triggered aggregation and DAKAP1alpha-mediated localization in living cells. Chem. Biol. 14, 1031–1042 (2007).
Harootunian, A.T. et al. Movement of the free catalytic subunit of cAMP-dependent protein kinase into and out of the nucleus can be explained by diffusion. Mol. Biol. Cell 4, 993–1002 (1993).
Mayr, B. & Montminy, M. Transcriptional regulation by the phosphorylation-dependent factor CREB. Nat. Rev. Mol. Cell Biol. 2, 599–609 (2001).
Meoli, E. et al. Protein kinase A effects of an expressed PRKAR1A mutation associated with aggressive tumors. Cancer Res. 68, 3133–3141 (2008).
Zippin, J.H. et al. Bicarbonate-responsive “soluble” adenylyl cyclase defines a nuclear cAMP microdomain. J. Cell Biol. 164, 527–534 (2004).
Jarnaess, E. et al. Splicing factor arginine/serine-rich 17A (SFRS17A) is an A-kinase anchoring protein that targets protein kinase A to splicing factor compartments. J. Biol. Chem. 284, 35154–35164 (2009).
Steegborn, C., Litvin, T.N., Levin, L.R., Buck, J. & Wu, H. Bicarbonate activation of adenylyl cyclase via promotion of catalytic active site closure and metal recruitment. Nat. Struct. Mol. Biol. 12, 32–37 (2005).
Chen, Y. et al. Soluble adenylyl cyclase as an evolutionarily conserved bicarbonate sensor. Science 289, 625–628 (2000).
Buck, J., Sinclair, M.L., Schapal, L., Cann, M.J. & Levin, L.R. Cytosolic adenylyl cyclase defines a unique signaling molecule in mammals. Proc. Natl. Acad. Sci. USA 96, 79–84 (1999).
DiPilato, L.M. & Zhang, J. The role of membrane microdomains in shaping β2-adrenergic receptor-mediated cAMP dynamics. Mol. Biosyst. 5, 832–837 (2009).
Ni, Q. et al. Signaling diversity of PKA achieved via a Ca2+-cAMP-PKA oscillatory circuit. Nat. Chem. Biol. 7, 34–40 (2011).
Terrin, A. et al. PGE1 stimulation of HEK293 cells generates multiple contiguous domains with different [cAMP]: role of compartmentalized phosphodiesterases. J. Cell Biol. 175, 441–451 (2006).
DiPilato, L.M., Cheng, X. & Zhang, J. Fluorescent indicators of cAMP and Epac activation reveal differential dynamics of cAMP signaling within discrete subcellular compartments. Proc. Natl. Acad. Sci. USA 101, 16513–16518 (2004).
Allen, M.D. & Zhang, J. Subcellular dynamics of protein kinase A activity visualized by FRET-based reporters. Biochem. Biophys. Res. Commun. 348, 716–721 (2006).
Hagiwara, M. et al. Coupling of hormonal stimulation and transcription via the cyclic AMP-responsive factor CREB is rate limited by nuclear entry of protein kinase A. Mol. Cell. Biol. 13, 4852–4859 (1993).
Burnham, K.P. & Anderson, D.R. Model Selection and Multi-Model Inference: Practical Information-Theoretic Approach Ch. 2, 49–97 (Springer, 2002).
Sugiura, N. Further analysis of data by Akaike's information criterion and finite corrections. Commun. Statist. 7, 13–26 (1978).
Shimojo, M., Paquette, A.J., Anderson, D.J. & Hersh, L.B. Protein kinase A regulates cholinergic gene expression in PC12 cells: REST4 silences the silencing activity of neuron-restrictive silencer factor/REST. Mol. Cell. Biol. 19, 6788–6795 (1999).
Lynch, M.J. et al. RNA silencing identifies PDE4D5 as the functionally relevant cAMP phosphodiesterase interacting with β arrestin to control the protein kinase A/AKAP79-mediated switching of the β2-adrenergic receptor to activation of ERK in HEK293B2 cells. J. Biol. Chem. 280, 33178–33189 (2005).
McCahill, A. et al. In resting COS1 cells a dominant negative approach shows that specific, anchored PDE4 cAMP phosphodiesterase isoforms gate the activation, by basal cyclic AMP production, of AKAP-tethered protein kinase A type II located in the centrosomal region. Cell. Signal. 17, 1158–1173 (2005).
de Rooij, J. et al. Mechanism of regulation of the Epac family of cAMP-dependent RapGEFs. J. Biol. Chem. 275, 20829–20836 (2000).
Beavo, J.A., Bechtel, P.J. & Krebs, E.G. Activation of protein kinase by physiological concentrations of cyclic AMP. Proc. Natl. Acad. Sci. USA 71, 3580–3583 (1974).
Wong, W. & Scott, J.D. AKAP signalling complexes: focal points in space and time. Nat. Rev. Mol. Cell Biol. 5, 959–970 (2004).
Dodge-Kafka, K.L. et al. The protein kinase A anchoring protein mAKAP coordinates two integrated cAMP effector pathways. Nature 437, 574–578 (2005).
Michel, J.J. & Scott, J.D. AKAP mediated signal transduction. Annu. Rev. Pharmacol. Toxicol. 42, 235–257 (2002).
Ellis-Davies, G.C. Caged compounds: photorelease technology for control of cellular chemistry and physiology. Nat. Methods 4, 619–628 (2007).
Stierl, M. et al. Light modulation of cellular cAMP by a small bacterial photoactivated adenylyl cyclase, bPAC, of the soil bacterium Beggiatoa. J. Biol. Chem. 286, 1181–1188 (2011).
Schröder-Lang, S. et al. Fast manipulation of cellular cAMP level by light in vivo. Nat. Methods 4, 39–42 (2007).
Kholodenko, B.N., Hancock, J.F. & Kolch, W. Signalling ballet in space and time. Nat. Rev. Mol. Cell Biol. 11, 414–426 (2010).
Chuang, H.Y., Hofree, M. & Ideker, T. A decade of systems biology. Annu. Rev. Cell Dev. Biol. 26, 721–744 (2010).
Tay, S. et al. Single-cell NF-κB dynamics reveal digital activation and analogue information processing. Nature 466, 267–271 (2010).
Paliwal, S. et al. MAPK-mediated bimodal gene expression and adaptive gradient sensing in yeast. Nature 446, 46–51 (2007).
Rich, T.C. et al. Cellular mechanisms underlying prostaglandin-induced transient cAMP signals near the plasma membrane of HEK-293 cells. Am. J. Physiol. Cell Physiol. 292, C319–C331 (2007).
Xin, W., Tran, T.M., Richter, W., Clark, R.B. & Rich, T.C. Roles of GRK and PDE4 activities in the regulation of β2 adrenergic signaling. J. Gen. Physiol. 131, 349–364 (2008).
Saucerman, J.J. et al. Systems analysis of PKA-mediated phosphorylation gradients in live cardiac myocytes. Proc. Natl. Acad. Sci. USA 103, 12923–12928 (2006).
Greenberg, M.E. & Bender, T.P. Identification of newly transcribed RNA. Curr. Protoc. Mol. Biol. 4, 4.10 (2007).
Acknowledgements
We thank L. Levin (Weill Cornell Medical College, Cornell University) for the gift of sACt cDNA. We thank M. Houslay (Institute of Biomedical and Life Sciences, University of Glasgow) and K. Xiang (University of Illinois, Urbana-Champaign) for the gift of dnPDE4 isoforms. We thank L. Hersh (University of Kentucky College of Medicine) for giving us the A126.1B2 and A126.1B2 Catβ cell line. We also thank S. Mehta, G. Mo, T. Ueno, C. Pohlmeyer and T. Inoue for their technical help. This work was funded by US National Institutes of Health (NIH) grants R01 DK073368, DP1 OD006419 (to J.Z.), F31 DK074381 (to L.M.D.) and R01 HL094476, American Heart Association grant 0830470N (to J.J.S.) and NIH grant GM08715 (to J.H.Y.).
Author information
Authors and Affiliations
Contributions
Q.N. conceived the original idea for SMICUS. V.S., L.M.D., Q.N. and J.Z. designed the experimental aspects of the project. J.H.Y. and J.J.S. designed the modeling aspects of the project. V.S. and L.M.D. carried out the experiments. J.H.Y. developed the mathematical model and carried out the simulations. V.S., L.M.D., J.H.Y., Q.N., J.J.S. and J.Z. analyzed the data. J.Z., L.M.D., V.S. and J.H.Y. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 3675 kb)
Rights and permissions
About this article
Cite this article
Sample, V., DiPilato, L., Yang, J. et al. Regulation of nuclear PKA revealed by spatiotemporal manipulation of cyclic AMP. Nat Chem Biol 8, 375–382 (2012). https://doi.org/10.1038/nchembio.799
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.799
This article is cited by
-
Cyclic AMP induces reversible EPAC1 condensates that regulate histone transcription
Nature Communications (2023)
-
Location-specific inhibition of Akt reveals regulation of mTORC1 activity in the nucleus
Nature Communications (2020)
-
cAMP-PKA signal transduction specificity in Saccharomyces cerevisiae
Current Genetics (2020)
-
Single-fluorophore biosensors for sensitive and multiplexed detection of signalling activities
Nature Cell Biology (2018)
-
FRET biosensor uncovers cAMP nano-domains at β-adrenergic targets that dictate precise tuning of cardiac contractility
Nature Communications (2017)