Abstract
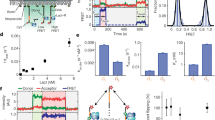
Many enzymes that react with specific sites in DNA have the property of facilitated diffusion, in which the DNA chain is used as a conduit to accelerate site location. Despite the importance of such mechanisms in gene regulation and DNA repair, there have been few viable approaches to elucidate the microscopic process of facilitated diffusion. Here we describe a new method in which a small-molecule trap (uracil) is used to clock a DNA repair enzyme as it hops and slides between damaged sites in DNA. The 'molecular clock' provides unprecedented information: the mean length for DNA sliding, the one-dimensional diffusion constant, the maximum hopping radius and the time frame for DNA hopping events. In addition, the data establish that the DNA phosphate backbone is a sufficient requirement for DNA sliding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Riggs, A.D., Bourgeois, S. & Cohn, M. The lac repressor-operator interaction. III. Kinetic studies. J. Mol. Biol. 53, 401–417 (1970).
Berg, O.G., Winter, R.B. & Von Hippel, P.H. Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory. Biochemistry 20, 6929–6948 (1981).
Gowers, D.M., Wilson, G.G. & Halford, S.E. Measurement of the contributions of 1D and 3D pathways to the translocation of a protein along DNA. Proc. Natl. Acad. Sci. USA 102, 15883–15888 (2005).
Stanford, N.P., Szczelkun, M.D., Marko, J.F. & Halford, S.E. One- and three-dimensional pathways for proteins to reach specific DNA sites. EMBO J. 19, 6546–6557 (2000).
Hedglin, M. & O'Brien, P.J. Human alkyladenine DNA glycosylase employs a processive search for DNA damage. Biochemistry 47, 11434–11445 (2008).
Sidorenko, V.S. & Zharkov, D.O. Correlated cleavage of damaged DNA by bacterial and human 8-oxoguanine-DNA glycosylases. Biochemistry 47, 8970–8976 (2008).
Porecha, R.H. & Stivers, J.T. Uracil DNA glycosylase uses DNA hopping and short-range sliding to trap extrahelical uracils. Proc. Natl. Acad. Sci. USA 105, 10791–10796 (2008).
Terry, B.J., Jack, W.E. & Modrich, P. Facilitated diffusion during catalysis by EcoRI endonuclease. Nonspecific interactions in EcoRI catalysis. J. Biol. Chem. 260, 13130–13137 (1985).
Elf, J., Li, G. & Xie, X.S. Probing transcription factor dynamics at the single-molecule level in a living cell. Science 316, 1191–1194 (2007).
Dowd, D.R. & Lloyd, R.S. Site-directed mutagenesis of the T4 endonuclease V gene: the role of arginine-3 in the target search. Biochemistry 28, 8699–8705 (1989).
Hedglin, M. & O'Brien, P.J. Hopping enables a DNA repair glycosylase to search both strands and bypass a bound protein. ACS Chem. Biol. 5, 427–436 (2010).
Tafvizi, A., Huang, F., Fersht, A.R., Mirny, L.A. & van Oijen, A.M. A single-molecule characterization of p53 search on DNA. Proc. Natl. Acad. Sci. USA 11, 563–568 (2010).
Komazin-Meredith, G., Mirchev, R., Golan, D.E., van Oijen, A.M. & Coen, D.M. Hopping of a processivity factor on DNA revealed by single-molecule assays of diffusion. Proc. Natl. Acad. Sci. USA 105, 10721–10726 (2008).
Kad, N.M., Wang, H., Kennedy, G.G., Warshaw, D.M. & Van Houten, B. Collaborative dynamic DNA scanning by nucleotide excision repair proteins investigated by single- molecule imaging of quantum-dot-labeled proteins. Mol. Cell 37, 702–713 (2010).
Gorman, J., Plys, A.J., Visnapuu, M., Alani, E. & Greene, E.C. Visualizing one-dimensional diffusion of eukaryotic DNA repair factors along a chromatin lattice. Nat. Struct. Mol. Biol. 17, 932–938 (2010).
Blainey, P.C. et al. Nonspecifically bound proteins spin while diffusing along DNA. Nat. Struct. Mol. Biol. 16, 1224–1229 (2009).
Blainey, P.C., van Oijen, A.M., Banerjee, A., Verdine, G.L. & Xie, X.S. A base-excision DNA-repair protein finds intrahelical lesion bases by fast sliding in contact with DNA. Proc. Natl. Acad. Sci. USA 103, 5752–5757 (2006).
Wang, Y.M., Austin, R.H. & Cox, E.C. Single molecule measurements of repressor protein 1D diffusion on DNA. Phys. Rev. Lett. 97, 048302 (2006).
Gorman, J. et al. Dynamic basis for one-dimensional DNA scanning by the mismatch repair complex Msh2-Msh6. Mol. Cell 28, 359–370 (2007).
Dunn, A.R., Kad, N.M., Nelson, S.R., Warshaw, D.M. & Wallace, S.S. Single Qdot-labeled glycosylase molecules use a wedge amino acid to probe for lesions while scanning along DNA. Nucleic Acids Res. 1, 7487–7498 (2011).
Wang, Y.M. & Austin,, R.H. Single-molecule imaging of LacI diffusing along nonspecific DNA. in Biophysics of DNA-Protein Interactions From Single Molecules to Biological Systems (eds. Williams, M.C. & Maher, J.L.) 9–37 (Springer, 2011).
Maul, R.W. & Gearhart, P.J. AID and somatic hypermutation. in Advances in Immunology (ed. Frederick W. Alt) 159–191 (Academic Press, 2010).
Griller, D. & Ingold, K.U. Free-radical clocks. Acc. Chem. Res. 13, 317–323 (1980).
Fishbein, J.C. & Jencks, W.P. Elimination reactions of β-cyano thioethers: evidence for a carbanion intermediate and a change in rate-limiting step. J. Am. Chem. Soc. 110, 5075–5086 (1988).
Amyes, T.L. & Jencks, W.P. Lifetimes of oxocarbenium ions in aqueous solution from common ion inhibition of the solvolysis of α-azido ethers by added azide ion. J. Am. Chem. Soc. 111, 7888–7900 (1989).
Xiao, G. et al. Crystal structure of Escherichia coli uracil DNA glycosylase and its complexes with uracil and glycerol: Structure and glycosylase mechanism revisited. Proteins 35, 13–24 (1999).
Jiang, Y.L., Krosky, D.J., Seiple, L. & Stivers, J.T. Uracil-directed ligand tethering: an efficient strategy for uracil DNA glycosylase (UNG) inhibitor development. J. Am. Chem. Soc. 127, 17412–17420 (2005).
Berg, H.C. Random Walks in Biology (Princeton University Press, 1993).
Belotserkovskii, B.P. & Zarling, D.A. Analysis of a one-dimensional random walk with irreversible losses at each step: applications for protein movement on DNA. J. Theor. Biol. 226, 195–203 (2004).
Halford, S.E. & Marko, J.F. How do site-specific DNA-binding proteins find their targets? Nucleic Acids Res. 32, 3040–3052 (2004).
Schurr, J.M. The one-dimensional diffusion coefficient of proteins absorbed on DNA: hydrodynamic considerations. Biophys. Chem. 9, 413–414 (1979).
Bagchi, B., Blainey, P.C. & Xie, X.S. Diffusion constant of a nonspecifically bound protein undergoing curvilinear motion along DNA. J. Phys. Chem. B 112, 6282–6284 (2008).
Bruner, S.D., Norman, D.P. & Verdine, G.L. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866 (2000).
Kuznetsov, N.A., Koval, V. & Fedorova, O. Mechanism of recognition and repair of damaged DNA by human 8-oxoguanine DNA glycosylase hOGG1. Biochemistry (Mosc.) 76, 118–130 (2011).
Kuznetsov, N.A. et al. Kinetic conformational analysis of human 8-oxoguanine-DNA glycosylase. J. Biol. Chem. 282, 1029–1038 (2007).
Cao, C., Jiang, Y.L., Stivers, J.T. & Song, F. Dynamic opening of DNA during the enzymatic search for a damaged base. Nat. Struct. Mol. Biol. 11, 1230–1236 (2004).
Parker, J.B. et al. Enzymatic capture of an extrahelical thymine in the search for uracil in DNA. Nature 449, 433–437 (2007).
Friedman, J.I., Majumdar, A. & Stivers, J.T. Nontarget DNA binding shapes the dynamic landscape for enzymatic recognition of DNA damage. Nucleic Acids Res. 37, 3493–3500 (2009).
Parker, J.B. & Stivers, J.T. Dynamics of uracil and 5-fluorouracil in DNA. Biochemistry 50, 612–617 (2011).
Kampmann, M. Obstacle bypass in protein motion along DNA by two-dimensional rather than one-dimensional sliding. J. Biol. Chem. 279, 38715–38720 (2004).
Nishida, K., Ando, Y. & Kawamura, H. Diffusion coefficients of anticancer drugs and compounds having a similar structure at 30 °C. Colloid Polym. Sci. 261, 70–73 (1983).
Young, M.A., Ravishanker, G. & Beveridge, D.L. A 5-nanosecond molecular dynamics trajectory for B-DNA: analysis of structure, motions, and solvation. Biophys. J. 73, 2313–2336 (1997).
Krosky, D.J., Song, F. & Stivers, J.T. The origins of high-affinity enzyme binding to an extrahelical DNA base. Biochemistry 44, 5949–5959 (2005).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Acknowledgements
We thank J. Parker (Harvard Medical School) for purified hUNG used in this study. This work was supported by US National Institutes of Health grant GM056834.
Author information
Authors and Affiliations
Contributions
J.D.S. performed all experiments. J.D.S. and J.T.S. analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 976 kb)
Rights and permissions
About this article
Cite this article
Schonhoft, J., Stivers, J. Timing facilitated site transfer of an enzyme on DNA. Nat Chem Biol 8, 205–210 (2012). https://doi.org/10.1038/nchembio.764
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.764
This article is cited by
-
“Flexible hinge” dynamics in mismatched DNA revealed by fluorescence correlation spectroscopy
Journal of Biological Physics (2022)
-
Shaping of the 3D genome by the ATPase machine cohesin
Experimental & Molecular Medicine (2020)
-
Dynamic DNA binding licenses a repair factor to bypass roadblocks in search of DNA lesions
Nature Communications (2016)
-
Kinetic gating mechanism of DNA damage recognition by Rad4/XPC
Nature Communications (2015)
-
Single-molecule methods for studying gene regulation in vivo
Pflügers Archiv - European Journal of Physiology (2013)