Abstract
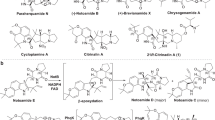
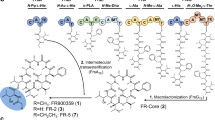
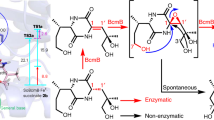
Spiroacetal compounds are ubiquitous in nature, and their stereospecific structures are responsible for diverse pharmaceutical activities. Elucidation of the biosynthetic mechanisms that are involved in spiroacetal formation will open the door to efficient generation of stereospecific structures that are otherwise hard to synthesize chemically. However, the biosynthesis of these compounds is poorly understood, owing to difficulties in identifying the responsible enzymes and analyzing unstable intermediates. Here we comprehensively describe the spiroacetal formation involved in the biosynthesis of reveromycin A, which inhibits bone resorption and bone metastases of tumor cells by inducing apoptosis in osteoclasts. We performed gene disruption, systematic metabolite analysis, feeding of labeled precursors and conversion studies with recombinant enzymes. We identified two key enzymes, dihydroxy ketone synthase and spiroacetal synthase, and showed in vitro reconstruction of the stereospecific spiroacetal structure from a stable acyclic precursor. Our findings provide insights into the creation of a variety of biologically active spiroacetal compounds for drug leads.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Weissman, K.J. & Leadlay, P.F. Combinatorial biosynthesis of reduced polyketides. Nat. Rev. Microbiol. 3, 925–936 (2005).
Hertweck, C. The biosynthetic logic of polyketide diversity. Angew. Chem. Int. Edn Engl. 48, 4688–4716 (2009).
Agtarap, A., Chamberlin, J.W., Pinkerton, M. & Steinrauf, L. The structure of monensic acid, a new biologically active compound. J. Am. Chem. Soc. 89, 5737–5739 (1967).
MacKintosh, C. & Klumpp, S. Tautomycin from the bacterium Streptomyces verticillatus. Another potent and specific inhibitor of protein phosphatases 1 and 2A. FEBS Lett. 277, 137–140 (1990).
Ishihara, H. et al. Calyculin A and okadaic acid: inhibitors of protein phosphatase activity. Biochem. Biophys. Res. Commun. 159, 871–877 (1989).
Bialojan, C. & Takai, A. Inhibitory effect of a marine-sponge toxin, okadaic acid, on protein phosphatases. Specificity and kinetics. Biochem. J. 256, 283–290 (1988).
Burg, R.W. et al. Avermectins, new family of potent anthelmintic agents: producing organism and fermentation. Antimicrob. Agents Chemother. 15, 361–367 (1979).
Niggemann, J. et al. Spirangien A and B, highly cytotoxic and antifungal spiroketals from the myxobacterium Sorangium cellulosum: isolation, structure elucidation and chemical modifications. Eur. J. Org. Chem. 2005, 5013–5018 (2005).
Osada, H., Koshino, H., Isono, K., Takahashi, H. & Kawanishi, G. Reveromycin A, a new antibiotic which inhibits the mitogenic activity of epidermal growth factor. J. Antibiot. (Tokyo) 44, 259–261 (1991).
Takahashi, H. et al. Reveromycins, new inhibitors of eukaryotic cell growth. II. Biological activities. J. Antibiot. (Tokyo) 45, 1414–1419 (1992).
Cui, Z., Hirata, D., Tsuchiya, E., Osada, H. & Miyakawa, T. The multidrug resistance-associated protein (MRP) subfamily (Yrs1/Yor1) of Saccharomyces cerevisiae is important for the tolerance to a broad range of organic anions. J. Biol. Chem. 271, 14712–14716 (1996).
Takahashi, H. et al. Inhibitory action of reveromycin A on TGF-α-dependent growth of ovarian carcinoma BG-1 in vitro and in vivo. Oncol. Res. 9, 7–11 (1997).
Miyamoto, Y. et al. Identification of Saccharomyces cerevisiae isoleucyl-tRNA synthetase as a target of the G1-specific inhibitor Reveromycin A. J. Biol. Chem. 277, 28810–28814 (2002).
Tanaka, Y. et al. Reveromycin A inhibits antigen receptor-mediated antigen presentation by B lymphoma cells via its effect on intracellular trafficking of the antigen. J. Antibiot. (Tokyo) 55, 904–913 (2002).
Woo, J.T. et al. Reveromycin A, an agent for osteoporosis, inhibits bone resorption by inducing apoptosis specifically in osteoclasts. Proc. Natl. Acad. Sci. USA 103, 4729–4734 (2006).
Muguruma, H. et al. Reveromycin A inhibits osteolytic bone metastasis of small-cell lung cancer cells, SBC-5, through an antiosteoclastic activity. Clin. Cancer Res. 11, 8822–8828 (2005).
Yano, A. et al. Inhibition of Hsp90 activates osteoclast c-Src signaling and promotes growth of prostate carcinoma cells in bone. Proc. Natl. Acad. Sci. USA 105, 15541–15546 (2008).
Shimizu, T., Masuda, T., Hiramoto, K. & Nakata, T. Total synthesis of reveromycin A. Org. Lett. 2, 2153–2156 (2000).
Paterson, I., Findlay, A.D. & Noti, C. Total synthesis of (–)-spirangien A, an antimitotic polyketide isolated from the myxobacterium Sorangium cellulosum. Chem. Asian J. 4, 594–611 (2009).
Sheppeck, J.E. II, Liu, W. & Chamberlin, A.R. Total synthesis of the serine/threonine-specific protein phosphatase inhibitor tautomycin (1). J. Org. Chem. 62, 387–398 (1997).
Oliynyk, M. et al. Analysis of the biosynthetic gene cluster for the polyether antibiotic monensin in Streptomyces cinnamonensis and evidence for the role of monB and monC genes in oxidative cyclization. Mol. Microbiol. 49, 1179–1190 (2003).
Bhatt, A. et al. Accumulation of an E,E,E-triene by the monensin-producing polyketide synthase when oxidative cyclization is blocked. Angew. Chem. Int. Edn. Engl. 44, 7075–7078 (2005).
Gallimore, A.R. et al. Evidence for the role of the monB genes in polyether ring formation during monensin biosynthesis. Chem. Biol. 13, 453–460 (2006).
Li, W., Ju, J., Rajski, S.R., Osada, H. & Shen, B. Characterization of the tautomycin biosynthetic gene cluster from Streptomyces spiroverticillatus unveiling new insights into dialkylmaleic anhydride and polyketide biosynthesis. J. Biol. Chem. 283, 28607–28617 (2008).
Ikeda, H., Nonomiya, T., Usami, M., Ohta, T. & Omura, S. Organization of the biosynthetic gene cluster for the polyketide anthelmintic macrolide avermectin in Streptomyces avermitilis. Proc. Natl. Acad. Sci. USA 96, 9509–9514 (1999).
Frank, B. et al. Spiroketal polyketide formation in Sorangium: identification and analysis of the biosynthetic gene cluster for the highly cytotoxic spirangienes. Chem. Biol. 14, 221–233 (2007).
Ishikawa, J. & Hotta, K. FramePlot: a new implementation of the frame analysis for predicting protein-coding regions in bacterial DNA with a high G + C content. FEMS Microbiol. Lett. 174, 251–253 (1999).
Staunton, J. & Weissman, K.J. Polyketide biosynthesis: a millennium review. Nat. Prod. Rep. 18, 380–416 (2001).
Bevitt, D.J., Cortes, J., Haydock, S.F. & Leadlay, P.F. 6-Deoxyerythronolide-B synthase 2 from Saccharopolyspora erythraea. Cloning of the structural gene, sequence analysis and inferred domain structure of the multifunctional enzyme. Eur. J. Biochem. 204, 39–49 (1992).
Donadio, S. & Katz, L. Organization of the enzymatic domains in the multifunctional polyketide synthase involved in erythromycin formation in Saccharopolyspora erythraea. Gene 111, 51–60 (1992).
Bisang, C. et al. A chain initiation factor common to both modular and aromatic polyketide synthases. Nature 401, 502–505 (1999).
Haydock, S.F. et al. Divergent sequence motifs correlated with the substrate specificity of (methyl)malonyl-CoA:acyl carrier protein transacylase domains in modular polyketide synthases. FEBS Lett. 374, 246–248 (1995).
Aparicio, J.F. et al. Organization of the biosynthetic gene cluster for rapamycin in Streptomyces hygroscopicus: analysis of the enzymatic domains in the modular polyketide synthase. Gene 169, 9–16 (1996).
Keatinge-Clay, A.T. & Stroud, R.M. The structure of a ketoreductase determines the organization of the beta-carbon processing enzymes of modular polyketide synthases. Structure 14, 737–748 (2006).
Reid, R. et al. A model of structure and catalysis for ketoreductase domains in modular polyketide synthases. Biochemistry 42, 72–79 (2003).
Caffrey, P. Conserved amino acid residues correlating with ketoreductase stereospecificity in modular polyketide synthases. ChemBioChem 4, 654–657 (2003).
Keatinge-Clay, A. Crystal structure of the erythromycin polyketide synthase dehydratase. J. Mol. Biol. 384, 941–953 (2008).
Kwan, D.H. et al. Prediction and manipulation of the stereochemistry of enoylreduction in modular polyketide synthases. Chem. Biol. 15, 1231–1240 (2008).
Witkowski, A., Naggert, J., Wessa, B. & Smith, S. A catalytic role for histidine 237 in rat mammary gland thioesterase II. J. Biol. Chem. 266, 18514–18519 (1991).
Benach, J. et al. Structure of bacterial 3β/17β-hydroxysteroid dehydrogenase at 1.2 A resolution: a model for multiple steroid recognition. Biochemistry 41, 14659–14668 (2002).
Shafqat, N. et al. Expanded substrate screenings of human and Drosophila type 10 17β-hydroxysteroid dehydrogenases (HSDs) reveal multiple specificities in bile acid and steroid hormone metabolism: characterization of multifunctional 3α/7α/7β/17β/20β/21-HSD. Biochem. J. 376, 49–60 (2003).
Shimizu, T., Satoh, T., Murakoshi, K. & Sodeoka, M. Asymmetric total synthesis of (−)-spirofungin A and (+)-spirofungin B. Org. Lett. 7, 5573–5576 (2005).
Höltzel, A. et al. Spirofungin, a new antifungal antibiotic from Streptomyces violaceusniger Tu 4113. J. Antibiot. (Tokyo) 51, 699–707 (1998).
Persson, B. et al. The SDR (short-chain dehydrogenase/reductase and related enzymes) nomenclature initiative. Chem. Biol. Interact. 178, 94–98 (2009).
Mayerhofer, L.E., Conway de Macario, E., Yao, R. & Macario, A.J. Structure, organization, and expression of genes coding for envelope components in the archaeon Methanosarcina mazei S-6. Arch. Microbiol. 169, 339–345 (1998).
Jing, H. et al. Archaeal surface layer proteins contain beta propeller, PKD, and beta helix domains and are related to metazoan cell surface proteins. Structure 10, 1453–1464 (2002).
Koshino, H., Takahashi, H., Osada, H. & Isono, K. Reveromycins, new inhibitors of eukaryotic cell growth. III. Structures of reveromycins A, B, C and D. J. Antibiot. (Tokyo) 45, 1420–1427 (1992).
Ubukata, M., Koshino, H., Osada, H. & Isono, K. Absolute configuration of reveromycin A, an inhibitor of the signal transduction of epidermal growth factor. J. Chem. Soc. Chem. Commun. 1994, 1877–1878 (1994).
Acknowledgements
We thank T. Shimizu for discussions on spiroacetal formation; K. Wierzba for reading this paper; T. Nakamura for mass analysis; N. Dohmae for MALDI-TOF and N-terminal sequencing; and E. Oowada for technical assistance. We thank all the technical staff of the Sequence Technology Team at RIKEN Genomic Sciences Center for their assistance. This work was supported in part by the Naito Foundation, the Strategic Programs for R&D of RIKEN, a Grant-in-Aid for Creative Scientific Research and a Grant-in-Aid for Scientific Research (C) from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Contributions
S.T.: the identification of RM gene cluster, genetic analysis, enzyme characterization and writing of the paper; A.T. and Y. Sakaki: sequencing of Streptomyces sp. SN-593 genome; Y. Sekiyama: the 13C-labeling experiments; H.T. and R.S.: the isolation of RM derivatives; T.N., M.U. and H.K.: the structural analyses; T.N. and M.U: the writing of structural analysis; S.P.: the analysis of the border genes; T.K.: the characterization of RevJ; T.D.: the RT-PCR analysis; J.I.: the development of the gene annotation system; H.I.: discussion about the gene transformation system for Streptomyces sp. SN-593; H.O.: the integration of all research. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 12412 kb)
Rights and permissions
About this article
Cite this article
Takahashi, S., Toyoda, A., Sekiyama, Y. et al. Reveromycin A biosynthesis uses RevG and RevJ for stereospecific spiroacetal formation. Nat Chem Biol 7, 461–468 (2011). https://doi.org/10.1038/nchembio.583
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.583
This article is cited by
-
Wakodecaline C, new tetrahydrofuran-fused decalin metabolite isolated from fungus Pyrenochaetopsis sp. RK10-F058
The Journal of Antibiotics (2023)
-
Studies on Streptomyces sp. SN-593: reveromycin biosynthesis, β-carboline biomediator activating LuxR family regulator, and construction of terpenoid biosynthetic platform
The Journal of Antibiotics (2022)
-
β-carboline chemical signals induce reveromycin production through a LuxR family regulator in Streptomyces sp. SN-593
Scientific Reports (2020)
-
β-carboline biomediators induce reveromycin production in Streptomyces sp. SN-593
Scientific Reports (2019)
-
An integrated screening system for the selection of exemplary substrates for natural and engineered cytochrome P450s
Scientific Reports (2019)