Abstract
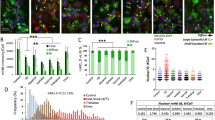
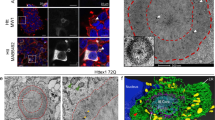
Two serine residues within the first 17 amino acid residues of huntingtin (N17) are crucial for modulation of mutant huntingtin toxicity in cell and mouse genetic models of Huntington's disease. Here we show that the stress-dependent phosphorylation of huntingtin at Ser13 and Ser16 affects N17 conformation and targets full-length huntingtin to chromatin-dependent subregions of the nucleus, the mitotic spindle and cleavage furrow during cell division. Polyglutamine-expanded mutant huntingtin is hypophosphorylated in N17 in both homozygous and heterozygous cell contexts. By high-content screening in live cells, we identified kinase inhibitors that modulated N17 phosphorylation and hence huntingtin subcellular localization. N17 phosphorylation was reduced by casein kinase-2 inhibitors. Paradoxically, IKKβ kinase inhibition increased N17 phosphorylation, affecting huntingtin nuclear and subnuclear localization. These data indicate that huntingtin phosphorylation at Ser13 and Ser16 can be modulated by small-molecule drugs, which may have therapeutic potential in Huntington's disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
The Huntington's Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 72, 971–983 (1993).
Atwal, R.S. et al. Huntingtin has a membrane association signal that can modulate huntingtin aggregation, nuclear entry and toxicity. Hum. Mol. Genet. 16, 2600–2615 (2007).
Kim, M.W., Chelliah, Y., Kim, S.W., Otwinowski, Z. & Bezprozvanny, I. Secondary structure of Huntingtin amino-terminal region. Structure 17, 1205–1212 (2009).
Tam, S. et al. The chaperonin TRiC blocks a huntingtin sequence element that promotes the conformational switch to aggregation. Nat. Struct. Mol. Biol. 16, 1279–1285 (2009).
Darnell, G., Orgel, J.P., Pahl, R. & Meredith, S.C. Flanking polyproline sequences inhibit beta-sheet structure in polyglutamine segments by inducing PPII-like helix structure. J. Mol. Biol. 374, 688–704 (2007).
Mangiarini, L. et al. Exon 1 of the HD gene with an expanded CAG repeat is sufficient to cause a progressive neurological phenotype in transgenic mice. Cell 87, 493–506 (1996).
Martín-Aparicio, E., Avila, J. & Lucas, J.J. Nuclear localization of N-terminal mutant huntingtin is cell cycle dependent. Eur. J. Neurosci. 16, 355–359 (2002).
Benn, C.L. et al. Huntingtin modulates transcription, occupies gene promoters in vivo, and binds directly to DNA in a polyglutamine-dependent manner. J. Neurosci. 28, 10720–10733 (2008).
Seong, I.S. et al. Huntingtin facilitates polycomb repressive complex 2. Hum. Mol. Genet. 19, 573–583 (2010).
Godin, J.D. et al. Huntingtin is required for mitotic spindle orientation and mammalian neurogenesis. Neuron 67, 392–406 (2010).
Steffan, J.S. et al. SUMO modification of Huntingtin and Huntington's disease pathology. Science 304, 100–104 (2004).
Aiken, C.T. et al. Phosphorylation of threonine 3: implications for Huntingtin aggregation and neurotoxicity. J. Biol. Chem. 284, 29427–29436 (2009).
Thompson, L.M. et al. IKK phosphorylates Huntingtin and targets it for degradation by the proteasome and lysosome. J. Cell Biol. 187, 1083–1099 (2009).
Lecerf, J.M. et al. Human single-chain Fv intrabodies counteract in situ huntingtin aggregation in cellular models of Huntington's disease. Proc. Natl. Acad. Sci. USA 98, 4764–4769 (2001).
Snyder-Keller, A., McLear, J.A., Hathorn, T. & Messer, A. Early or late-stage anti-N-terminal Huntingtin intrabody gene therapy reduces pathological features in B6.HDR6/1 mice. J. Neuropathol. Exp. Neurol. 69, 1078–1085 (2010).
Williamson, T.E., Vitalis, A., Crick, S.L. & Pappu, R.V. Modulation of polyglutamine conformations and dimer formation by the N-terminus of huntingtin. J. Mol. Biol. 396, 1295–1309 (2010).
Colby, D.W., Cassady, J.P., Lin, G.C., Ingram, V.M. & Wittrup, K.D. Stochastic kinetics of intracellular huntingtin aggregate formation. Nat. Chem. Biol. 2, 319–323 (2006).
Thakur, A.K. et al. Polyglutamine disruption of the huntingtin exon 1 N terminus triggers a complex aggregation mechanism. Nat. Struct. Mol. Biol. 16, 380–389 (2009).
Wetzel, M.K. et al. p73 regulates neurodegeneration and phospho-tau accumulation during aging and Alzheimer's disease. Neuron 59, 708–721 (2008).
Gu, X. et al. Serines 13 and 16 are critical determinants of full-length human mutant huntingtin induced disease pathogenesis in HD mice. Neuron 64, 828–840 (2009).
Greenfield, N.J. Using circular dichroism spectra to estimate protein secondary structure. Nat. Protoc. 1, 2876–2890 (2006).
Schilling, G. et al. Intranuclear inclusions and neuritic aggregates in transgenic mice expressing a mutant N-terminal fragment of huntingtin. Hum. Mol. Genet. 8, 397–407 (1999).
Zhang, H. et al. Elucidating a normal function of huntingtin by functional and microarray analysis of huntingtin-null mouse embryonic fibroblasts. BMC Neurosci. 9, 38 (2008).
Bordeaux, J. et al. Antibody validation. Biotechniques 48, 197–209 (2010).
Hoffner, G., Kahlem, P. & Djian, P. Perinuclear localization of huntingtin as a consequence of its binding to microtubules through an interaction with beta-tubulin: relevance to Huntington's disease. J. Cell Sci. 115, 941–948 (2002).
Pagano, M.A. et al. 2-Dimethylamino-4,5,6,7-tetrabromo-1H-benzimidazole: a novel powerful and selective inhibitor of protein kinase CK2. Biochem. Biophys. Res. Commun. 321, 1040–1044 (2004).
Cozza, G. et al. Quinalizarin as a potent, selective and cell-permeable inhibitor of protein kinase CK2. Biochem. J. 421, 387–395 (2009).
Burke, J.R. et al. BMS-345541 is a highly selective inhibitor of IκB kinase that binds at an allosteric site of the enzyme and blocks NF-κB-dependent transcription in mice. J. Biol. Chem. 278, 1450–1456 (2003).
Arrasate, M., Mitra, S., Schweitzer, E.S., Segal, M.R. & Finkbeiner, S. Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 431, 805–810 (2004).
Truant, R., Atwal, R.S., Desmond, C., Munsie, L. & Tran, T. Huntington's disease: revisiting the aggregation hypothesis in polyglutamine neurodegenerative diseases. FEBS J. 275, 4252–4262 (2008).
Wetzel, R. Nucleation of huntingtin aggregation in cells. Nat. Chem. Biol. 2, 297–298 (2006).
Reina, C.P., Zhong, X. & Pittman, R.N. Proteotoxic stress increases nuclear localization of ataxin-3. Hum. Mol. Genet. 19, 235–249 (2009).
Mueller, T. et al. CK2-dependent phosphorylation determines cellular localization and stability of ataxin-3. Hum. Mol. Genet. 18, 3334–3343 (2009).
Fan, M.M., Zhang, H., Hayden, M.R., Pelech, S.L. & Raymond, L.A. Protective up-regulation of CK2 by mutant huntingtin in cells co-expressing NMDA receptors. J. Neurochem. 104, 790–805 (2008).
Modregger, J., DiProspero, N.A., Charles, V., Tagle, D.A. & Plomann, M. PACSIN 1 interacts with huntingtin and is absent from synaptic varicosities in presymptomatic Huntington's disease brains. Hum. Mol. Genet. 11, 2547–2558 (2002).
Plomann, M. et al. PACSIN, a brain protein that is upregulated upon differentiation into neuronal cells. Eur. J. Biochem. 256, 201–211 (1998).
Di Maira, G., Brustolon, F., Pinna, L.A. & Ruzzene, M. Dephosphorylation and inactivation of Akt/PKB is counteracted by protein kinase CK2 in HEK 293T cells. Cell. Mol. Life Sci. 66, 3363–3373 (2009).
Humbert, S. et al. The IGF-1/Akt pathway is neuroprotective in Huntington's disease and involves Huntingtin phosphorylation by Akt. Dev. Cell 2, 831–837 (2002).
Kageura, T. et al. Inhibitors from rhubarb on lipopolysaccharide-induced nitric oxide production in macrophages: structural requirements of stilbenes for the activity. Bioorg. Med. Chem. 9, 1887–1893 (2001).
Pierce, J.W. et al. Novel inhibitors of cytokine-induced IκBα phosphorylation and endothelial cell adhesion molecule expression show anti-inflammatory effects in vivo. J. Biol. Chem. 272, 21096–21103 (1997).
Shaul, J.D., Farina, A. & Huxford, T. The human IKKβ subunit kinase domain displays CK2-like phosphorylation specificity. Biochem. Biophys. Res. Commun. 374, 592–597 (2008).
McElhinny, J.A., Trushin, S.A., Bren, G.D., Chester, N. & Paya, C.V. Casein kinase II phosphorylates IκBα at S-283, S-289, S-293, and T-291 and is required for its degradation. Mol. Cell. Biol. 16, 899–906 (1996).
Witt, J. et al. Mechanism of PP2A-mediated IKKβ dephosphorylation: a systems biological approach. BMC Syst. Biol. 3, 71 (2009).
Metzler, M. et al. Phosphorylation of huntingtin at Ser421 in YAC128 neurons is associated with protection of YAC128 neurons from NMDA-mediated excitotoxicity and is modulated by PP1 and PP2A. J. Neurosci. 30, 14318–14329 (2010).
Pardo, R. et al. Inhibition of calcineurin by FK506 protects against polyglutamine-huntingtin toxicity through an increase of huntingtin phosphorylation at S421. J. Neurosci. 26, 1635–1645 (2006).
Trushin, S.A., Pennington, K.N., Algeciras-Schimnich, A. & Paya, C.V. Protein kinase C and calcineurin synergize to activate IκB kinase and NF-κB in T lymphocytes. J. Biol. Chem. 274, 22923–22931 (1999).
Bamborough, P. et al. Progress towards the development of anti-inflammatory inhibitors of IKKβ. Curr. Top. Med. Chem. 9, 623–639 (2009).
Trettel, F. et al. Dominant phenotypes produced by the HD mutation in STHdh(Q111) striatal cells. Hum. Mol. Genet. 9, 2799–2809 (2000).
Acknowledgements
This work was supported by operating grants from the Canadian Institutes of Health Research (MOP-165174), the Krembil Foundation and the CHDI Foundation (to R.T.). We thank M. Prakesch (Ontario Institute for Cancer Research) for supplying multiple CK2 and IKKβ inhibitors, I. Bezprozvanny (University of Texas Southwestern) and G. Hajnocsky (Thomas Jefferson University) for the gift of huntingtin-null MEF cell lines, M.E. MacDonald (Massachusetts General Hospital) for the gift of mouse striatal STHdh cell lines, and Raquel and Richard Epand for assistance and advice with CD spectroscopy.
Author information
Authors and Affiliations
Contributions
R.S.A. conceived and performed most experiments. C.R.D., N.C., J.X. and T.M. performed additional experiments. R.T. conceived experiments, prepared final figures and wrote the manuscript with R.S.A., C.R.D., N.C. and T.M. S.S. shared preliminary data to conceive experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Methods (PDF 3622 kb)
Supplementary Video 1
Huntingtin1-171 Q142-eYFP expression in STHdh cells. 24 hours. This cell population forms inclusions that coalesce into a few large inclusions by a soluble exchange, yet cell death is not evident. (MOV 1003 kb)
Supplementary Video 2
Huntingtin1-171 Q142-eYFP expression in STHdh cells II. 24 hours. This cell population does not form large coalesced inclusions, only small multiple inclusions that rapidly lead to cell death. (MOV 911 kb)
Supplementary Video 3
Huntingtin1-171 (Q142) S13A S16A-eYFP expression in STHdh cells. 24 hours. This cell population rapidly forms multiple inclusions relative to the wild-type N17 constructs, but cell death is delayed. Relative to the wild-type N17 1-171 Q142, this suggests that the two different populations seen with the wild-type protein may correspond to cell signaling in one population versus another. When all huntingtin fragments are phosphomimicked, inclusions are large and often only 1-2 spots, cell death is delayed, suggesting that phospho-N17 inclusions may be innocuous to the cell. (MOV 808 kb)
Rights and permissions
About this article
Cite this article
Atwal, R., Desmond, C., Caron, N. et al. Kinase inhibitors modulate huntingtin cell localization and toxicity. Nat Chem Biol 7, 453–460 (2011). https://doi.org/10.1038/nchembio.582
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.582
This article is cited by
-
The cytoprotective role of GM1 ganglioside in Huntington disease cells
Molecular Biology Reports (2022)
-
Exploring the role of high-mobility group box 1 (HMGB1) protein in the pathogenesis of Huntington’s disease
Journal of Molecular Medicine (2020)
-
DNA Damage Repair in Huntington's Disease and Other Neurodegenerative Diseases
Neurotherapeutics (2019)
-
Therapeutic approaches to Huntington disease: from the bench to the clinic
Nature Reviews Drug Discovery (2018)
-
Proteostasis in Huntington's disease: disease mechanisms and therapeutic opportunities
Acta Pharmacologica Sinica (2018)