Abstract
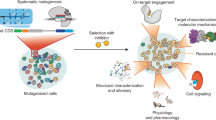
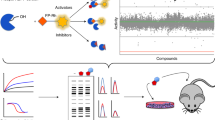
Drug discovery and chemical genetic efforts typically focus on the identification and design of inhibitors or loss-of-function probes as a means to perturb enzyme function. These tools are effective in determining the physiological consequence of ablating the activity of a specific enzyme. Remarkably, nearly a dozen examples of non-natural small molecules that activate enzyme catalysis have been identified within the past decade. In aggregate, these studies delineate four unique activation mechanisms that the small molecules exploit. These complementary gain-of-function probes offer a way to address the sufficiency of an enzyme to drive a particular cellular phenotype, and they also provide new opportunities for drug discovery. This review covers the identification and characterization of these unique small-molecule activators.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Alaimo, P.J., Shogren-Knaak, M.A. & Shokat, K.M. Chemical genetic approaches for the elucidation of signaling pathways. Curr. Opin. Chem. Biol. 5, 360–367 (2001).
Cravatt, B.F. & Sorensen, E.J. Chemical strategies for the global analysis of protein function. Curr. Opin. Chem. Biol. 4, 663–668 (2000).
Stockwell, B.R. Chemical genetics: ligand-based discovery of gene function. Nat. Rev. Genet. 1, 116–125 (2000).
Cárdenas, M.L. & Cornish-Bowden, A. Characteristics necessary for an interconvertible enzyme cascade to generate a highly sensitive response to an effector. Biochem. J. 257, 339–345 (1989).
Goldbeter, A. & Koshland, D.E. Sensitivity amplification in biochemical systems. Q. Rev. Biophys. 15, 555–591 (1982).
Szedlacsek, S.E., Cárdenas, M.L. & Cornish-Bowden, A. Response coefficients of interconvertible enzyme cascades towards effectors that act on one or both modifier enzymes. Eur. J. Biochem. 204, 807–813 (1992).
Bishop, A.C. & Chen, V.L. Brought to life: targeted activation of enzyme function with small molecules. J. Chem. Biol. 2, 1–9 (2009).
Cárdenas, M.L., Cornish-Bowden, A. & Ureta, T. Evolution and regulatory role of the hexokinases. Biochim. Biophys. Acta 1401, 242–264 (1998).
Matschinsky, F. Assessing the potential of glucokinase activators in diabetes therapy. Nat. Rev. Drug Discov. 8, 399–416 (2009).
Pal, M. Recent advances in glucokinase activators for the treatment of type 2 diabetes. Drug Discov. Today 14, 784–792 (2009).
Matschinsky, F.M. et al. The network of glucokinase-expressing cells in glucose homeostasis and the potential of glucokinase activators for diabetes therapy. Diabetes 55, 1–12 (2006).
Gidh-Jain, M. et al. Glucokinase mutations associated with non-insulin-dependent (type 2) diabetes mellitus have decreased enzymatic activity: implications for structure/function relationships. Proc. Natl. Acad. Sci. USA 90, 1932–1936 (1993).
de la Iglesia, N., Veiga-da-Cunha, M., Van Schaftingen, E., Guinovart, J.J. & Ferrer, J.C. Glucokinase regulatory protein is essential for the proper subcellular localisation of liver glucokinase. FEBS Lett. 456, 332–338 (1999).
Anderka, O. et al. Biophysical characterization of the interaction between hepatic glucokinase and its regulatory protein: impact of physiological and pharmacological effectors. J. Biol. Chem. 283, 31333–31340 (2008).
Veiga-da-Cunha, M. & Van Schaftingen, E. Identification of fructose 6-phosphate- and fructose 1-phosphate-binding residues in the regulatory protein of glucokinase. J. Biol. Chem. 277, 8466–8473 (2002).
Grimsby, J. et al. Allosteric activators of glucokinase: potential role in diabetes therapy. Science 301, 370–373 (2003).
Ralph, E.C., Thomson, J., Almaden, J. & Sun, S. Glucose modulation of glucokinase activation by small molecules. Biochemistry 47, 5028–5036 (2008).
Guertin, K.R. & Grimsby, J. Small molecule glucokinase activators as glucose lowering agents: a new paradigm for diabetes therapy. Curr. Med. Chem. 13, 1839–1843 (2006).
Kamata, K., Mitsuya, M., Nishimura, T., Eiki, J.-I. & Nagata, Y. Structural basis for allosteric regulation of the monomeric allosteric enzyme human glucokinase. Structure 12, 429–438 (2004).
Milne, J.C. & Denu, J.M. The Sirtuin family: therapeutic targets to treat diseases of aging. Curr. Opin. Chem. Biol. 12, 11–17 (2008).
Longo, V.D. & Kennedy, B.K. Sirtuins in aging and age-related disease. Cell 126, 257–268 (2006).
Kaeberlein, M., McVey, M. & Guarente, L. The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes Dev. 13, 2570–2580 (1999).
Lin, S.J., Defossez, P.A. & Guarente, L. Requirement of NAD and SIR2 for life-span extension by calorie restriction in Saccharomyces cerevisiae. Science 289, 2126–2128 (2000).
Cohen, H.Y. et al. Calorie restriction promotes mammalian cell survival by inducing the SIRT1 deacetylase. Science 305, 390–392 (2004).
Lavu, S., Boss, O., Elliott, P.J. & Lambert, P.D. Sirtuins–novel therapeutic targets to treat age-associated diseases. Nat. Rev. Drug Discov. 7, 841–853 (2008).
Kaeberlein, M., Kirkland, K.T., Fields, S. & Kennedy, B.K. Sir2-independent life span extension by calorie restriction in yeast. PLoS Biol. 2, E296 ( 2004).
Finkel, T., Deng, C.-X. & Mostoslavsky, R. Recent progress in the biology and physiology of sirtuins. Nature 460, 587–591 (2009).
Anderson, R.M., Bitterman, K.J., Wood, J.G., Medvedik, O. & Sinclair, D.A. Nicotinamide and PNC1 govern lifespan extension by calorie restriction in Saccharomyces cerevisiae. Nature 423, 181–185 (2003).
Kim, J.-E., Chen, J. & Lou, Z. DBC1 is a negative regulator of SIRT1. Nature 451, 583–586 (2008).
Zhao, W. et al. Negative regulation of the deacetylase SIRT1 by DBC1. Nature 451, 587–590 (2008).
Kim, E.-J., Kho, J.-H., Kang, M.-R. & Um, S.-J. Active regulator of SIRT1 cooperates with SIRT1 and facilitates suppression of p53 activity. Mol. Cell 28, 277–290 (2007).
Smith, B.C., Hallows, W.C. & Denu, J.M. Mechanisms and molecular probes of sirtuins. Chem. Biol. 15, 1002–1013 (2008).
Howitz, K.T. et al. Small molecule activators of sirtuins extend Saccharomyces cerevisiae lifespan. Nature 425, 191–196 (2003).
Kaeberlein, M. et al. Substrate-specific activation of sirtuins by resveratrol. J. Biol. Chem. 280, 17038–17045 (2005).
Milne, J.C. et al. Small molecule activators of SIRT1 as therapeutics for the treatment of type 2 diabetes. Nature 450, 712–716 (2007).
Pacholec, M. et al. SRT1720, SRT2183, SRT1460, and resveratrol are not direct activators of SIRT1. J. Biol. Chem. published online, doi:10.1074/jbc.M109.088682 (8 January 2010).
Frödin, M. et al. A phosphoserine/threonine-binding pocket in AGC kinases and PDK1 mediates activation by hydrophobic motif phosphorylation. EMBO J. 21, 5396–5407 (2002).
Huse, M. & Kuriyan, J. The conformational plasticity of protein kinases. Cell 109, 275–282 (2002).
Biondi, R.M. Phosphoinositide-dependent protein kinase 1, a sensor of protein conformation. Trends Biochem. Sci. 29, 136–142 (2004).
Biondi, R.M. et al. High resolution crystal structure of the human PDK1 catalytic domain defines the regulatory phosphopeptide docking site. EMBO J. 21, 4219–4228 (2002).
Biondi, R.M. et al. Identification of a pocket in the PDK1 kinase domain that interacts with PIF and the C-terminal residues of PKA. EMBO J. 19, 979–988 (2000).
Engel, M. et al. Allosteric activation of the protein kinase PDK1 with low molecular weight compounds. EMBO J. 25, 5469–5480 (2006).
Stroba, A. et al. 3,5-Diphenylpent-2-enoic acids as allosteric activators of the protein kinase PDK1: structure-activity relationships and thermodynamic characterization of binding as paradigms for PIF-binding pocket-targeting compounds. J. Med. Chem. 52, 4683–4693 (2009).
Hindie, V. et al. Structure and allosteric effects of low-molecular-weight activators on the protein kinase PDK1. Nat. Chem. Biol. 5, 758–764 (2009).
Biondi, R.M., Kieloch, A., Currie, R.A., Deak, M. & Alessi, D.R. The PIF-binding pocket in PDK1 is essential for activation of S6K and SGK, but not PKB. EMBO J. 20, 4380–4390 (2001).
Pellicena, P. & Kuriyan, J. Protein-protein interactions in the allosteric regulation of protein kinases. Curr. Opin. Struct. Biol. 16, 702–709 (2006).
Tappan, E. & Chamberlin, A.R. Activation of protein phosphatase 1 by a small molecule designed to bind to the enzyme's regulatory site. Chem. Biol. 15, 167–174 (2008).
Balasubramanyam, K., Swaminathan, V., Ranganathan, A. & Kundu, T.K. Small molecule modulators of histone acetyltransferase p300. J. Biol. Chem. 278, 19134–19140 (2003).
Reineke, J. et al. Autocatalytic cleavage of Clostridium difficile toxin B. Nature 446, 415–419 (2007).
Egerer, M., Giesemann, T., Jank, T., Satchell, K.J.F. & Aktories, K. Auto-catalytic cleavage of Clostridium difficile toxins A and B depends on cysteine protease activity. J. Biol. Chem. 282, 25314–25321 (2007).
Egerer, M., Giesemann, T., Herrmann, C. & Aktories, K. Autocatalytic processing of Clostridium difficile toxin B. Binding of inositol hexakisphosphate. J. Biol. Chem. 284, 3389–3395 (2009).
Lupardus, P.J., Shen, A., Bogyo, M. & Garcia, K.C. Small molecule-induced allosteric activation of the Vibrio cholerae RTX cysteine protease domain. Science 322, 265–268 (2008).
Shen, A. et al. Mechanistic and structural insights into the proteolytic activation of Vibrio cholerae MARTX toxin. Nat. Chem. Biol. 5, 469–478 (2009).
Prochazkova, K. & Satchell, K.J.F. Structure-function analysis of inositol hexakisphosphate-induced autoprocessing of the Vibrio cholerae multifunctional autoprocessing RTX toxin. J. Biol. Chem. 283, 23656–23664 (2008).
Prochazkova, K. et al. Structural and molecular mechanism for autoprocessing of MARTX toxin of Vibrio cholerae at multiple sites. J. Biol. Chem. 284, 26557–26568 (2009).
Pop, C. & Salvesen, G. Human caspases: activation, specificity and regulation. J. Biol. Chem. 284, 21777–21781 (2009).
Creagh, E.M., Conroy, H. & Martin, S.J. Caspase-activation pathways in apoptosis and immunity. Immunol. Rev. 193, 10–21 (2003).
Yi, C.H. & Yuan, J. The Jekyll and Hyde functions of caspases. Dev. Cell 16, 21–34 (2009).
Stennicke, H.R. & Salvesen, G.S. Caspases - controlling intracellular signals by protease zymogen activation. Biochim. Biophys. Acta 1477, 299–306 (2000).
Earnshaw, W.C., Martins, L.M. & Kaufmann, S.H. Mammalian caspases: structure, activation, substrates, and functions during apoptosis. Annu. Rev. Biochem. 68, 383–424 (1999).
Salvesen, G.S. & Riedl, S.J. Caspase mechanisms. Adv. Exp. Med. Biol. 615, 13–23 (2008).
Roy, S. et al. Maintenance of caspase-3 proenzyme dormancy by an intrinsic “safety catch” regulatory tripeptide. Proc. Natl. Acad. Sci. USA 98, 6132–6137 (2001).
Hardy, J.A., Lam, J., Nguyen, J.T., O'Brien, T. & Wells, J.A. Discovery of an allosteric site in the caspases. Proc. Natl. Acad. Sci. USA 101, 12461–12466 (2004).
Scheer, J.M., Romanowski, M.J. & Wells, J.A. A common allosteric site and mechanism in caspases. Proc. Natl. Acad. Sci. USA 103, 7595–7600 (2006).
Gao, J., Sidhu, S.S. & Wells, J.A. Two-state selection of conformation-specific antibodies. Proc. Natl. Acad. Sci. USA 106, 3071–3076 (2009).
Chai, J. et al. Crystal structure of a procaspase-7 zymogen: mechanisms of activation and substrate binding. Cell 107, 399–407 (2001).
Riedl, S.J. et al. Structural basis for the activation of human procaspase-7. Proc. Natl. Acad. Sci. USA 98, 14790–14795 (2001).
Putt, K.S. et al. Small-molecule activation of procaspase-3 to caspase-3 as a personalized anticancer strategy. Nat. Chem. Biol. 2, 543–550 (2006).
Denault, J.-B. et al. Small molecules not direct activators of caspases. Nat. Chem. Biol. 3, 519 (2007); author reply 3, 520 (2007).
Peterson, Q.P. et al. PAC-1 activates procaspase-3 in vitro through relief of zinc-mediated inhibition. J. Mol. Biol. 388, 144–158 (2009).
Wolan, D.W., Zorn, J.A., Gray, D.C. & Wells, J.A. Small-molecule activators of a proenzyme. Science 326, 853–858 (2009).
Svingen, P.A. et al. Components of the cell death machine and drug sensitivity of the National Cancer Institute Cell Line Panel. Clin. Cancer Res. 10, 6807–6820 (2004).
Zhang, B.B., Zhou, G. & Li, C. AMPK: an emerging drug target for diabetes and the metabolic syndrome. Cell Metab. 9, 407–416 (2009).
Hardie, D.G. Minireview: the AMP-activated protein kinase cascade: the key sensor of cellular energy status. Endocrinology 144, 5179–5183 (2003).
Hardie, D.G. AMP-activated protein kinase as a drug target. Annu. Rev. Pharmacol. Toxicol. 47, 185–210 (2007).
Amodeo, G.A., Rudolph, M.J. & Tong, L. Crystal structure of the heterotrimer core of Saccharomyces cerevisiae AMPK homologue SNF1. Nature 449, 492–495 (2007).
Crute, B.E., Seefeld, K., Gamble, J., Kemp, B.E. & Witters, L.A. Functional domains of the alpha1 catalytic subunit of the AMP-activated protein kinase. J. Biol. Chem. 273, 35347–35354 (1998).
Chen, L. et al. Structural insight into the autoinhibition mechanism of AMP-activated protein kinase. Nature 459, 1146–1149 (2009).
Xiao, B. et al. Structural basis for AMP binding to mammalian AMP-activated protein kinase. Nature 449, 496–500 (2007).
Townley, R. & Shapiro, L. Crystal structures of the adenylate sensor from fission yeast AMP-activated protein kinase. Science 315, 1726–1729 (2007).
Kahn, B.B., Alquier, T., Carling, D. & Hardie, D.G. AMP-activated protein kinase: ancient energy gauge provides clues to modern understanding of metabolism. Cell Metab. 1, 15–25 (2005).
Sabina, R.L., Holmes, E.W. & Becker, M.A. The enzymatic synthesis of 5-amino-4-imidazolecarboxamide riboside triphosphate (ZTP). Science 223, 1193–1195 (1984).
Corton, J.M., Gillespie, J.G., Hawley, S.A. & Hardie, D.G. 5-aminoimidazole-4-carboxamide ribonucleoside. A specific method for activating AMP-activated protein kinase in intact cells? Eur. J. Biochem. 229, 558–565 (1995).
Cool, B. et al. Identification and characterization of a small molecule AMPK activator that treats key components of type 2 diabetes and the metabolic syndrome. Cell Metab. 3, 403–416 (2006).
Göransson, O. et al. Mechanism of action of A-769662, a valuable tool for activation of AMP-activated protein kinase. J. Biol. Chem. 282, 32549–32560 (2007).
Scott, J.W. et al. Thienopyridone drugs are selective activators of AMP-activated protein kinase beta1-containing complexes. Chem. Biol. 15, 1220–1230 (2008).
Sanders, M.J. et al. Defining the mechanism of activation of AMP-activated protein kinase by the small molecule A-769662, a member of the thienopyridone family. J. Biol. Chem. 282, 32539–32548 (2007).
Zhao, G. et al. Discovery and SAR development of thienopyridones: a class of small molecule AMPK activators. Bioorg. Med. Chem. Lett. 17, 3254–3257 (2007).
Pang, T. et al. Small molecule antagonizes autoinhibition and activates AMP-activated protein kinase in cells. J. Biol. Chem. 283, 16051–16060 (2008).
Kim, C., Cheng, C.Y., Saldanha, S.A. & Taylor, S.S. PKA-I holoenzyme structure reveals a mechanism for cAMP-dependent activation. Cell 130, 1032–1043 (2007).
Taylor, S.S. et al. Signaling through cAMP and cAMP-dependent protein kinase: diverse strategies for drug design. Biochim. Biophys. Acta 1784, 16–26 (2008).
Yang, J. et al. Contribution of non-catalytic core residues to activity and regulation in protein kinase A. J. Biol. Chem. 284, 6241–6248 (2009).
Saldanha, S.A., Kaler, G., Cottam, H.B., Abagyan, R. & Taylor, S.S. Assay principle for modulators of protein-protein interactions and its application to non-ATP-competitive ligands targeting protein kinase A. Anal. Chem. 78, 8265–8272 (2006).
Budas, G.R., Koyanagi, T., Churchill, E.N. & Mochly-Rosen, D. Competitive inhibitors and allosteric activators of protein kinase C isoenzymes: a personal account and progress report on transferring academic discoveries to the clinic. Biochem. Soc. Trans. 35, 1021–1026 (2007).
Csukai, M. & Mochly-Rosen, D. Pharmacologic modulation of protein kinase C isozymes: the role of RACKs and subcellular localisation. Pharmacol. Res. 39, 253–259 (1999).
Churchill, E.N., Qvit, N. & Mochly-Rosen, D. Rationally designed peptide regulators of protein kinase C. Trends Endocrinol. Metab. 20, 25–33 (2009).
Silverman, R.H. Implications for RNase L in prostate cancer biology. Biochemistry 42, 1805–1812 (2003).
Wreschner, D.H., McCauley, J.W., Skehel, J.J. & Kerr, I.M. Interferon action–sequence specificity of the ppp(A2′p)nA-dependent ribonuclease. Nature 289, 414–417 (1981).
Tanaka, N. et al. Structural basis for recognition of 2′,5′-linked oligoadenylates by human ribonuclease L. EMBO J. 23, 3929–3938 (2004).
Tanaka, N. et al. Molecular basis for recognition of 2′,5′-linked oligoadenylates by the N-terminal ankyrin repeat domain of human ribonuclease L. Nucleic Acids Symp. Ser. (Oxf) 323–324 (2005).
Nakanishi, M., Goto, Y. & Kitade, Y. 2–5A induces a conformational change in the ankyrin-repeat domain of RNase L. Proteins 60, 131–138 (2005).
Nakanishi, M. et al. Functional characterization of 2′,5′-linked oligoadenylate binding determinant of human RNase L. J. Biol. Chem. 280, 41694–41699 (2005).
Thakur, C.S. et al. Small-molecule activators of RNase L with broad-spectrum antiviral activity. Proc. Natl. Acad. Sci. USA 104, 9585–9590 (2007).
Papa, F.R., Zhang, C., Shokat, K. & Walter, P. Bypassing a kinase activity with an ATP-competitive drug. Science 302, 1533–1537 (2003).
McPherson, J.D. et al. A physical map of the human genome. Nature 409, 934–941 (2001).
Venter, J.C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Chen, C.-H. et al. Activation of aldehyde dehydrogenase-2 reduces ischemic damage to the heart. Science 321, 1493–1495 (2008).
Okuzumi, T. et al. Inhibitor hijacking of Akt activation. Nat. Chem. Biol. 5, 484–493 (2009).
Sergina, N.V. et al. Escape from HER-family tyrosine kinase inhibitor therapy by the kinase-inactive HER3. Nature 445, 437–441 (2007).
Acknowledgements
We thank J. Sadowsky, S. Mahrus and D. Wolan for helpful discussions and N. Agard, J. Diaz, D. Wildes and D. Wolan for critically reviewing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Zorn, J., Wells, J. Turning enzymes ON with small molecules. Nat Chem Biol 6, 179–188 (2010). https://doi.org/10.1038/nchembio.318
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.318
This article is cited by
-
Deubiquitinase-targeting chimeras for targeted protein stabilization
Nature Chemical Biology (2022)
-
Body weight regulation via MT1-MMP-mediated cleavage of GFRAL
Nature Metabolism (2022)
-
Chemical activation of divergent protein tyrosine phosphatase domains with cyanine-based biarsenicals
Scientific Reports (2019)
-
Pharmacologic treatment with CPI-613 and PS48 decreases mitochondrial membrane potential and increases quantity of autolysosomes in porcine fibroblasts
Scientific Reports (2019)
-
Correcting glucose-6-phosphate dehydrogenase deficiency with a small-molecule activator
Nature Communications (2018)