Abstract
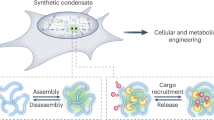
We have developed a system for producing a supramolecular scaffold that permeates the entire Escherichia coli cytoplasm. This cytoscaffold is constructed from a three-component system comprising a bacterial microcompartment shell protein and two complementary de novo coiled-coil peptides. We show that other proteins can be targeted to this intracellular filamentous arrangement. Specifically, the enzymes pyruvate decarboxylase and alcohol dehydrogenase have been directed to the filaments, leading to enhanced ethanol production in these engineered bacterial cells compared to those that do not produce the scaffold. This is consistent with improved metabolic efficiency through enzyme colocation. Finally, the shell-protein scaffold can be directed to the inner membrane of the cell, demonstrating how synthetic cellular organization can be coupled with spatial optimization through in-cell protein design. The cytoscaffold has potential in the development of next-generation cell factories, wherein it could be used to organize enzyme pathways and metabolite transporters to enhance metabolic flux.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Polka, J.K., Hays, S.G. & Silver, P.A. Building spatial synthetic biology with compartments, scaffolds, and communities. Cold Spring Harb. Perspect. Biol. 8, a024018 (2016).
Zhang, Y. et al. Using unnatural protein fusions to engineer resveratrol biosynthesis in yeast and Mammalian cells. J. Am. Chem. Soc. 128, 13030–13031 (2006).
Grinkova, Y.V., Denisov, I.G. & Sligar, S.G. Engineering extended membrane scaffold proteins for self-assembly of soluble nanoscale lipid bilayers. Protein Eng. Des. Sel. 23, 843–848 (2010).
Delebecque, C.J., Silver, P.A. & Lindner, A.B. Designing and using RNA scaffolds to assemble proteins in vivo. Nat. Protoc. 7, 1797–1807 (2012).
Zalatan, J.G. et al. Engineering complex synthetic transcriptional programs with CRISPR RNA scaffolds. Cell 160, 339–350 (2015).
Agapakis, C.M., Boyle, P.M. & Silver, P.A. Natural strategies for the spatial optimization of metabolism in synthetic biology. Nat. Chem. Biol. 8, 527–535 (2012).
Dueber, J.E. et al. Synthetic protein scaffolds provide modular control over metabolic flux. Nat. Biotechnol. 27, 753–759 (2009).
Poshyvailo, L., von Lieres, E. & Kondrat, S. Does metabolite channeling accelerate enzyme-catalyzed cascade reactions? PLoS One 12, e0172673 (2017).
Wheeldon, I. et al. Substrate channelling as an approach to cascade reactions. Nat. Chem. 8, 299–309 (2016).
Chowdhury, C., Sinha, S., Chun, S., Yeates, T.O. & Bobik, T.A. Diverse bacterial microcompartment organelles. Microbiol. Mol. Biol. Rev. 78, 438–468 (2014).
Frank, S., Lawrence, A.D., Prentice, M.B. & Warren, M.J. Bacterial microcompartments moving into a synthetic biological world. J. Biotechnol. 163, 273–279 (2013).
Kerfeld, C.A. & Erbilgin, O. Bacterial microcompartments and the modular construction of microbial metabolism. Trends Microbiol. 23, 22–34 (2015).
Cameron, J.C., Wilson, S.C., Bernstein, S.L. & Kerfeld, C.A. Biogenesis of a bacterial organelle: the carboxysome assembly pathway. Cell 155, 1131–1140 (2013).
Kerfeld, C.A., Heinhorst, S. & Cannon, G.C. Bacterial microcompartments. Annu. Rev. Microbiol. 64, 391–408 (2010).
Bobik, T.A., Havemann, G.D., Busch, R.J., Williams, D.S. & Aldrich, H.C. The propanediol utilization (pdu) operon of Salmonella enterica serovar Typhimurium LT2 includes genes necessary for formation of polyhedral organelles involved in coenzyme B12-dependent 1,2-propanediol degradation. J. Bacteriol. 181, 5967–5975 (1999).
Havemann, G.D. & Bobik, T.A. Protein content of polyhedral organelles involved in coenzyme B12-dependent degradation of 1,2-propanediol in Salmonella enterica serovar Typhimurium LT2. J. Bacteriol. 185, 5086–5095 (2003).
Sampson, E.M. & Bobik, T.A. Microcompartments for B12-dependent 1,2-propanediol degradation provide protection from DNA and cellular damage by a reactive metabolic intermediate. J. Bacteriol. 190, 2966–2971 (2008).
Fan, C. & Bobik, T.A. The N-terminal region of the medium subunit (PduD) packages adenosylcobalamin-dependent diol dehydratase (PduCDE) into the Pdu microcompartment. J. Bacteriol. 193, 5623–5628 (2011).
Fan, C. et al. Short N-terminal sequences package proteins into bacterial microcompartments. Proc. Natl. Acad. Sci. USA 107, 7509–7514 (2010).
Fan, C., Cheng, S., Sinha, S. & Bobik, T.A. Interactions between the termini of lumen enzymes and shell proteins mediate enzyme encapsulation into bacterial microcompartments. Proc. Natl. Acad. Sci. USA 109, 14995–15000 (2012).
Lawrence, A.D. et al. Solution structure of a bacterial microcompartment targeting peptide and its application in the construction of an ethanol bioreactor. ACS Synth. Biol. 3, 454–465 (2014).
Jakobson, C.M., Tullman-Ercek, D., Slininger, M.F. & Mangan, N.M. A systems-level model reveals that 1,2-Propanediol utilization microcompartments enhance pathway flux through intermediate sequestration. PLOS Comput. Biol. 13, e1005525 (2017).
Sutter, M., Greber, B., Aussignargues, C. & Kerfeld, C.A. Assembly principles and structure of a 6.5-MDa bacterial microcompartment shell. Science 356, 1293–1297 (2017).
Cho, H. The role of cytoskeletal elements in shaping bacterial cells. J. Microbiol. Biotechnol. 25, 307–316 (2015).
Cabeen, M.T. & Jacobs-Wagner, C. Bacterial cell shape. Nat. Rev. Microbiol. 3, 601–610 (2005).
Cabeen, M.T. & Jacobs-Wagner, C. The bacterial cytoskeleton. Annu. Rev. Genet. 44, 365–392 (2010).
Parsons, J.B. et al. Synthesis of empty bacterial microcompartments, directed organelle protein incorporation, and evidence of filament-associated organelle movement. Mol. Cell 38, 305–315 (2010).
Chowdhury, C. et al. Selective molecular transport through the protein shell of a bacterial microcompartment organelle. Proc. Natl. Acad. Sci. USA 112, 2990–2995 (2015).
Crowley, C.S. et al. Structural insight into the mechanisms of transport across the Salmonella enterica Pdu microcompartment shell. J. Biol. Chem. 285, 37838–37846 (2010).
Pang, A., Frank, S., Brown, I., Warren, M.J. & Pickersgill, R.W. Structural insights into higher order assembly and function of the bacterial microcompartment protein PduA. J. Biol. Chem. 289, 22377–22384 (2014).
Thomas, F., Boyle, A.L., Burton, A.J. & Woolfson, D.N. A set of de novo designed parallel heterodimeric coiled coils with quantified dissociation constants in the micromolar to sub-nanomolar regime. J. Am. Chem. Soc. 135, 5161–5166 (2013).
Lee, M.J., Brown, I.R., Juodeikis, R., Frank, S. & Warren, M.J. Employing bacterial microcompartment technology to engineer a shell-free enzyme-aggregate for enhanced 1,2-propanediol production in Escherichia coli. Metab. Eng. 36, 48–56 (2016).
Fletcher, J.M. et al. Self-assembling cages from coiled-coil peptide modules. Science 340, 595–599 (2013).
Weber, B. et al. Automated tracing of microtubules in electron tomograms of plastic embedded samples of Caenorhabditis elegans embryos. J. Struct. Biol. 178, 129–138 (2012).
Johnson, E. et al. Correlative in-resin super-resolution and electron microscopy using standard fluorescent proteins. Sci. Rep. 5, 9583 (2015).
Mueller-Reichert, T. & Verkade, P. Correlative Light and Electron Microscopy II; Methods in Cell Biology Vol. 124 (Academic Press, 2014).
Szeto, T.H., Rowland, S.L., Habrukowich, C.L. & King, G.F. The MinD membrane targeting sequence is a transplantable lipid-binding helix. J. Biol. Chem. 278, 40050–40056 (2003).
Noël, C.R., Cai, F. & Kerfeld, C.A. Purification and characterization of protein nanotubes assembled from a single bacterial microcompartment shell subunit. Adv. Mater. Interfaces 3, 1500295 (2015).
Lewicka, A.J. et al. Fusion of pyruvate decarboxylase and alcohol dehydrogenase increases ethanol production in Escherichia coli. ACS Synth. Biol. 3, 976–978 (2014).
Kremer, J.R., Mastronarde, D.N. & McIntosh, J.R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Mastronarde, D.N. Dual-axis tomography: an approach with alignment methods that preserve resolution. J. Struct. Biol. 120, 343–352 (1997).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Paul-Gilloteaux, P. et al. eC-CLEM: flexible multidimensional registration software for correlative microscopies. Nat. Methods 14, 102–103 (2017).
Acknowledgements
We are grateful to the Biotechnology and Biological Sciences Research Council of the UK for a strategic LoLa Award to M.J.W., D.N.W., P.V. and W.-F.X. (BB/M002969/1). D.N.W. holds a Royal Society Wolfson Research Merit Award. We thank the Wolfson Bioimaging Facility and BrisSynBio, a BBSRC/EPSRC-funded Synthetic Biology Research Centre (L01386X), for access to confocal and electron microscopes; K. Howland for assistance with GC–MS analysis; R. Sessions and I. Uddin for preparing images used in Supplementary Figure 1; L. Harrington and P. Schwille for advice on the MinD system; and the entire BMC-SAGE LoLa group for helpful discussions.
Author information
Authors and Affiliations
Contributions
M.J.L. made constructs, prepared samples for TEM and confocal analysis, imaged samples by TEM, purified nanotubes and analyzed them by TEM and AFM and conducted the ethanol production experiments and analyses. J.M. undertook tomography and 3D reconstructions. L.H. undertook CLEM sample preparation and imaging. D.A. undertook confocal imaging. I.R.B. sectioned samples for TEM analysis. W.-F.X. assisted with AFM and statistical analysis. M.J.L., J.M., L.H., J.M.F., S.F., P.V., D.N.W. and M.J.W. designed the experiments. All authors contributed to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–4, Supplementary Figures 1–14 (PDF 2536 kb)
Rights and permissions
About this article
Cite this article
Lee, M., Mantell, J., Hodgson, L. et al. Engineered synthetic scaffolds for organizing proteins within the bacterial cytoplasm. Nat Chem Biol 14, 142–147 (2018). https://doi.org/10.1038/nchembio.2535
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2535
This article is cited by
-
Assembling membraneless organelles from de novo designed proteins
Nature Chemistry (2024)
-
Engineering synthetic biomolecular condensates
Nature Reviews Bioengineering (2023)
-
Synthetic biology promotes the capture of CO2 to produce fatty acid derivatives in microbial cell factories
Bioresources and Bioprocessing (2022)
-
Metabolic pathway assembly using docking domains from type I cis-AT polyketide synthases
Nature Communications (2022)
-
Intracellular artificial supramolecules based on de novo designed Y15 peptides
Nature Communications (2021)