Abstract
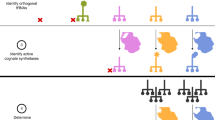
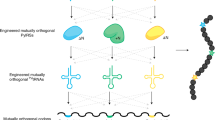
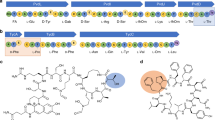
Directed evolution of orthogonal aminoacyl-tRNA synthetases (AARSs) enables site-specific installation of noncanonical amino acids (ncAAs) into proteins. Traditional evolution techniques typically produce AARSs with greatly reduced activity and selectivity compared to their wild-type counterparts. We designed phage-assisted continuous evolution (PACE) selections to rapidly produce highly active and selective orthogonal AARSs through hundreds of generations of evolution. PACE of a chimeric Methanosarcina spp. pyrrolysyl-tRNA synthetase (PylRS) improved its enzymatic efficiency (kcat/KMtRNA) 45-fold compared to the parent enzyme. Transplantation of the evolved mutations into other PylRS-derived synthetases improved yields of proteins containing noncanonical residues up to 9.7-fold. Simultaneous positive and negative selection PACE over 48 h greatly improved the selectivity of a promiscuous Methanocaldococcus jannaschii tyrosyl-tRNA synthetase variant for site-specific incorporation of p-iodo-L-phenylalanine. These findings offer new AARSs that increase the utility of orthogonal translation systems and establish the capability of PACE to efficiently evolve orthogonal AARSs with high activity and amino acid specificity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
01 December 2017
In the version of this article initially published, the label colors for p-NF and p-IF in the key for Figure 4c were transposed. The cyan bars should correspond to p-NF and the fuchsia bars to p-IF. The error has been corrected in the HTML and PDF versions of the article.
References
Liu, C.C. & Schultz, P.G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Wan, W., Tharp, J.M. & Liu, W.R. Pyrrolysyl-tRNA synthetase: an ordinary enzyme but an outstanding genetic code expansion tool. Biochim. Biophys. Acta 1844, 1059–1070 (2014).
Chin, J.W. Expanding and reprogramming the genetic code of cells and animals. Annu. Rev. Biochem. 83, 379–408 (2014).
Umehara, T. et al. N-acetyl lysyl-tRNA synthetases evolved by a CcdB-based selection possess N-acetyl lysine specificity in vitro and in vivo. FEBS Lett. 586, 729–733 (2012).
O'Donoghue, P., Ling, J., Wang, Y.-S. & Söll, D. Upgrading protein synthesis for synthetic biology. Nat. Chem. Biol. 9, 594–598 (2013).
Esvelt, K.M., Carlson, J.C. & Liu, D.R. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011).
Leconte, A.M. et al. A population-based experimental model for protein evolution: effects of mutation rate and selection stringency on evolutionary outcomes. Biochemistry 52, 1490–1499 (2013).
Carlson, J.C., Badran, A.H., Guggiana-Nilo, D.A. & Liu, D.R. Negative selection and stringency modulation in phage-assisted continuous evolution. Nat. Chem. Biol. 10, 216–222 (2014).
Dickinson, B.C., Packer, M.S., Badran, A.H. & Liu, D.R. A system for the continuous directed evolution of proteases rapidly reveals drug-resistance mutations. Nat. Commun. 5, 5352 (2014).
Hubbard, B.P. et al. Continuous directed evolution of DNA-binding proteins to improve TALEN specificity. Nat. Methods 12, 939–942 (2015).
Badran, A.H. et al. Continuous evolution of Bacillus thuringiensis toxins overcomes insect resistance. Nature 533, 58–63 (2016).
Riechmann, L. & Holliger, P. The C-terminal domain of TolA is the coreceptor for filamentous phage infection of E. coli. Cell 90, 351–360 (1997).
Yin, Y.W. & Steitz, T.A. Structural basis for the transition from initiation to elongation transcription in T7 RNA polymerase. Science 298, 1387–1395 (2002).
Tsao, M.L., Summerer, D., Ryu, Y. & Schultz, P.G. The genetic incorporation of a distance probe into proteins in Escherichia coli. J. Am. Chem. Soc. 128, 4572–4573 (2006).
Grünewald, J. et al. Immunochemical termination of self-tolerance. Proc. Natl. Acad. Sci. USA 105, 11276–11280 (2008).
Yanagisawa, T. et al. Multistep engineering of pyrrolysyl-tRNA synthetase to genetically encode Nε-(o-azidobenzyloxycarbonyl) lysine for site-specific protein modification. Chem. Biol. 15, 1187–1197 (2008).
Lubkowski, J., Hennecke, F., Plückthun, A. & Wlodawer, A. The structural basis of phage display elucidated by the crystal structure of the N-terminal domains of g3p. Nat. Struct. Biol. 5, 140–147 (1998).
Lubkowski, J., Hennecke, F., Plückthun, A. & Wlodawer, A. Filamentous phage infection: crystal structure of g3p in complex with its coreceptor, the C-terminal domain of TolA. Structure 7, 711–722 (1999).
Deng, L.W. & Perham, R.N. Delineating the site of interaction on the pIII protein of filamentous bacteriophage fd with the F-pilus of Escherichia coli. J. Mol. Biol. 319, 603–614 (2002).
Boeke, J.D. & Model, P. A prokaryotic membrane anchor sequence: carboxyl terminus of bacteriophage f1 gene III protein retains it in the membrane. Proc. Natl. Acad. Sci. USA 79, 5200–5204 (1982).
Boeke, J.D., Model, P. & Zinder, N.D. Effects of bacteriophage f1 gene III protein on the host cell membrane. Mol. Gen. Genet. 186, 185–192 (1982).
Rapoza, M.P. & Webster, R.E. The filamentous bacteriophage assembly proteins require the bacterial SecA protein for correct localization to the membrane. J. Bacteriol. 175, 1856–1859 (1993).
Badran, A.H. & Liu, D.R. Development of potent in vivo mutagenesis plasmids with broad mutational spectra. Nat. Commun. 6, 8425 (2015).
Guo, L.T. et al. Polyspecific pyrrolysyl-tRNA synthetases from directed evolution. Proc. Natl. Acad. Sci. USA 111, 16724–16729 (2014).
Kavran, J.M. et al. Structure of pyrrolysyl-tRNA synthetase, an archaeal enzyme for genetic code innovation. Proc. Natl. Acad. Sci. USA 104, 11268–11273 (2007).
Wong, M.L., Guzei, I.A. & Kiessling, L.L. An asymmetric synthesis of l-pyrrolysine. Org. Lett. 14, 1378–1381 (2012).
Jiang, R. & Krzycki, J.A. PylSn and the homologous N-terminal domain of pyrrolysyl-tRNA synthetase bind the tRNA that is essential for the genetic encoding of pyrrolysine. J. Biol. Chem. 287, 32738–32746 (2012).
Herring, S. et al. The amino-terminal domain of pyrrolysyl-tRNA synthetase is dispensable in vitro but required for in vivo activity. FEBS Lett. 581, 3197–3203 (2007).
Neumann, H. et al. A method for genetically installing site-specific acetylation in recombinant histones defines the effects of H3 K56 acetylation. Mol. Cell 36, 153–163 (2009).
Neumann, H., Peak-Chew, S.Y. & Chin, J.W. Genetically encoding Nε-acetyllysine in recombinant proteins. Nat. Chem. Biol. 4, 232–234 (2008).
Ho, J.M. et al. Efficient reassignment of a frequent serine codon in wild-type Escherichia coli. ACS Synth. Biol. 5, 163–171 (2016).
Adhin, M.R. & van Duin, J. Scanning model for translational reinitiation in eubacteria. J. Mol. Biol. 213, 811–818 (1990).
Andrè, A. et al. Reinitiation of protein synthesis in Escherichia coli can be induced by mRNA cis-elements unrelated to canonical translation initiation signals. FEBS Lett. 468, 73–78 (2000).
Cabantous, S., Rogers, Y., Terwilliger, T.C. & Waldo, G.S. New molecular reporters for rapid protein folding assays. PLoS One 3, e2387 (2008).
Nozawa, K. et al. Pyrrolysyl-tRNA synthetase-tRNA(Pyl) structure reveals the molecular basis of orthogonality. Nature 457, 1163–1167 (2009).
Xie, J. et al. The site-specific incorporation of p-iodo-l-phenylalanine into proteins for structure determination. Nat. Biotechnol. 22, 1297–1301 (2004).
Turner, J.M., Graziano, J., Spraggon, G. & Schultz, P.G. Structural plasticity of an aminoacyl-tRNA synthetase active site. Proc. Natl. Acad. Sci. USA 103, 6483–6488 (2006).
Yanagisawa, T. et al. Crystallographic studies on multiple conformational states of active-site loops in pyrrolysyl-tRNA synthetase. J. Mol. Biol. 378, 634–652 (2008).
Wang, H.H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Amiram, M. et al. Evolution of translation machinery in recoded bacteria enables multi-site incorporation of nonstandard amino acids. Nat. Biotechnol. 33, 1272–1279 (2015).
Lajoie, M.J. et al. Genomically recoded organisms expand biological functions. Science 342, 357–360 (2013).
Schmied, W.H., Elsässer, S.J., Uttamapinant, C. & Chin, J.W. Efficient multisite unnatural amino acid incorporation in mammalian cells via optimized pyrrolysyl tRNA synthetase/tRNA expression and engineered eRF1. J. Am. Chem. Soc. 136, 15577–15583 (2014).
Fan, C., Xiong, H., Reynolds, N.M. & Söll, D. Rationally evolving tRNAPyl for efficient incorporation of noncanonical amino acids. Nucleic Acids Res. 43, e156 (2015).
Young, T.S., Ahmad, I., Yin, J.A. & Schultz, P.G. An enhanced system for unnatural amino acid mutagenesis in E. coli. J. Mol. Biol. 395, 361–374 (2010).
Guo, J., Melançon, C.E. III, Lee, H.S., Groff, D. & Schultz, P.G. Evolution of amber suppressor tRNAs for efficient bacterial production of proteins containing nonnatural amino acids. Angew. Chem. Int. Edn. Engl. 48, 9148–9151 (2009).
Li, Z. et al. Nitrilase-activatable noncanonical amino acid precursors for cell-selective metabolic labeling of proteomes. ACS Chem. Biol. 11, 3273–3277 (2016).
Acknowledgements
The authors thank S. Trauger at the Small Molecule Mass Spectrometry Laboratory at Harvard University for providing expertise with intact protein mass spectrometry analysis. This work was supported by the Defense Advanced Research Projects Agency N66001-12-C-4207 (D.R.L.), the US National Institutes of Health (NIH) R01 EB022376 (D.R.L.), R35 GM118062 (D.R.L.), R21 AI119813 (C.F.), R01 GM022854 (D.S.), and R35 GM122560 (D.S.), the Department of Energy FG02-98ER2031 (D.S.), and the Howard Hughes Medical Institute (D.R.L.). D.I.B. is supported by a Ruth L. Kirschstein National Research Service Award (F32 GM106621).
Author information
Authors and Affiliations
Contributions
D.I.B. designed the research, performed experiments, analyzed data, and wrote the manuscript. D.R.L. designed and supervised the research and wrote the manuscript. D.S. designed and supervised the research. C.F. performed protein purification, in vitro aminoacylation assays, aided with in vivo amber suppression assays, and analyzed data. L.-T.G. designed the chimeric chPylRS variant for evolution in PACE, performed protein purification, performed in vitro aminoacylation assays, and analyzed data. C.M. aided in mutation analysis of evolved chPylRS variants from PACE. All authors contributed to editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have filed a provisional patent application on the PACE system and related improvements.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–19, Supplementary Tables 1–8 and Supplementary Note 1 (PDF 8892 kb)
Rights and permissions
About this article
Cite this article
Bryson, D., Fan, C., Guo, LT. et al. Continuous directed evolution of aminoacyl-tRNA synthetases. Nat Chem Biol 13, 1253–1260 (2017). https://doi.org/10.1038/nchembio.2474
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2474
This article is cited by
-
Tracking endogenous proteins based on RNA editing-mediated genetic code expansion
Nature Chemical Biology (2024)
-
Deciphering functional roles of protein succinylation and glutarylation using genetic code expansion
Nature Chemistry (2024)
-
A non-canonical nucleophile unlocks a new mechanistic pathway in a designed enzyme
Nature Communications (2024)
-
An evolved pyrrolysyl-tRNA synthetase with polysubstrate specificity expands the toolbox for engineering enzymes with incorporation of noncanonical amino acids
Bioresources and Bioprocessing (2023)
-
Applications of synthetic biology in medical and pharmaceutical fields
Signal Transduction and Targeted Therapy (2023)