Abstract
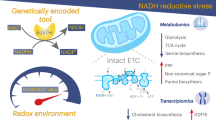
The redox coenzymes NADH and NADPH are broadly required for energy metabolism, biosynthesis and detoxification. Despite detailed knowledge of specific enzymes and pathways that utilize these coenzymes, a holistic understanding of the regulation and compartmentalization of NADH- and NADPH-dependent pathways is lacking, partly because of a lack of tools with which to investigate these processes in living cells. We have previously reported the use of the naturally occurring Lactobacillus brevis H2O-forming NADH oxidase (LbNOX) as a genetic tool for manipulation of the NAD+/NADH ratio in human cells. Here, we present triphosphopyridine nucleotide oxidase (TPNOX), a rationally designed and engineered mutant of LbNOX that is strictly specific to NADPH. We characterized the effects of TPNOX expression on cellular metabolism and used it in combination with LbNOX to show how the redox states of mitochondrial NADPH and NADH pools are connected.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Williamson, D.H., Lund, P. & Krebs, H.A. The redox state of free nicotinamide-adenine dinucleotide in the cytoplasm and mitochondria of rat liver. Biochem. J. 103, 514–527 (1967).
Veech, R.L., Eggleston, L.V. & Krebs, H.A. The redox state of free nicotinamide-adenine dinucleotide phosphate in the cytoplasm of rat liver. Biochem. J. 115, 609–619 (1969).
Sies, H. Metabolic compartmentation (Academic Press, 1982).
Klingenberg, M. & Buecher, T. Biological oxidations. Annu. Rev. Biochem. 29, 669–708 (1960).
Pollak, N., Dölle, C. & Ziegler, M. The power to reduce: pyridine nucleotides—small molecules with a multitude of functions. Biochem. J. 402, 205–218 (2007).
MacDonald, M.J. Feasibility of a mitochondrial pyruvate malate shuttle in pancreatic islets: further implication of cytosolic NADPH in insulin secretion. J. Biol. Chem. 270, 20051–20058 (1995).
Ronnebaum, S.M. et al. A pyruvate cycling pathway involving cytosolic NADP-dependent isocitrate dehydrogenase regulates glucose-stimulated insulin secretion. J. Biol. Chem. 281, 30593–30602 (2006).
Freeman, H.C., Hugill, A., Dear, N.T., Ashcroft, F.M. & Cox, R.D. Deletion of nicotinamide nucleotide transhydrogenase: a new quantitive trait locus accounting for glucose intolerance in C57BL/6J mice. Diabetes 55, 2153–2156 (2006).
Pandolfi, P.P. et al. Targeted disruption of the housekeeping gene encoding glucose 6-phosphate dehydrogenase (G6PD): G6PD is dispensable for pentose synthesis but essential for defense against oxidative stress. EMBO J. 14, 5209–5215 (1995).
Ducker, G.S. et al. Reversal of cytosolic one-carbon flux compensates for loss of the mitochondrial folate pathway. Cell Metab. 23, 1140–1153 (2016).
Chen, D. et al. Tissue-specific regulation of SIRT1 by calorie restriction. Genes Dev. 22, 1753–1757 (2008).
Sahlin, K., Katz, A. & Henriksson, J. Redox state and lactate accumulation in human skeletal muscle during dynamic exercise. Biochem. J. 245, 551–556 (1987).
O'Neill, J.S. & Reddy, A.B. Circadian clocks in human red blood cells. Nature 469, 498–503 (2011).
Schwartz, J.P., Passonneau, J.V., Johnson, G.S. & Pastan, I. The effect of growth conditions on NAD+ and NADH concentrations and the NAD+:NADH ratio in normal and transformed fibroblasts. J. Biol. Chem. 249, 4138–4143 (1974).
Braidy, N. et al. Age related changes in NAD+ metabolism oxidative stress and Sirt1 activity in wistar rats. PLoS One 6, e19194 (2011).
Zhu, X.H., Lu, M., Lee, B.Y., Ugurbil, K. & Chen, W. In vivo NAD assay reveals the intracellular NAD contents and redox state in healthy human brain and their age dependences. Proc. Natl. Acad. Sci. USA 112, 2876–2881 (2015).
Verdin, E. NAD+ in aging, metabolism, and neurodegeneration. Science 350, 1208–1213 (2015).
Zhao, Y. et al. SoNar, a highly responsive NAD+/NADH sensor, allows high-throughput metabolic screening of anti-tumor agents. Cell Metab. 21, 777–789 (2015).
Hung, Y.P., Albeck, J.G., Tantama, M. & Yellen, G. Imaging cytosolic NADH-NAD+ redox state with a genetically encoded fluorescent biosensor. Cell Metab. 14, 545–554 (2011).
Cameron, W.D. et al. Apollo-NADP+: a spectrally tunable family of genetically encoded sensors for NADP+. Nat. Methods 13, 352–358 (2016).
Bilan, D.S. & Belousov, V.V. Genetically encoded probes for NAD+/NADH monitoring. Free Radic. Biol. Med. 100, 32–42 (2016).
Fan, J. et al. Quantitative flux analysis reveals folate-dependent NADPH production. Nature 510, 298–302 (2014).
Liu, L. et al. Malic enzyme tracers reveal hypoxia-induced switch in adipocyte NADPH pathway usage. Nat. Chem. Biol. 12, 345–352 (2016).
Lewis, C.A. et al. Tracing compartmentalized NADPH metabolism in the cytosol and mitochondria of mammalian cells. Mol. Cell 55, 253–263 (2014).
Jiang, L. et al. Reductive carboxylation supports redox homeostasis during anchorage-independent growth. Nature 532, 255–258 (2016).
Titov, D.V. et al. Complementation of mitochondrial electron transport chain by manipulation of the NAD+/NADH ratio. Science 352, 231–235 (2016).
Cahn, J.K. et al. A general tool for engineering the NAD/NADP cofactor preference of oxidoreductases. ACS Synth. Biol. 6, 326–333 (2017).
Brown, D.M., Upcroft, J.A. & Upcroft, P. A H2O-producing NADH oxidase from the protozoan parasite Giardia duodenalis. Eur. J. Biochem. 241, 155–161 (1996).
Lountos, G.T. et al. The crystal structure of NAD(P)H oxidase from Lactobacillus sanfranciscensis: insights into the conversion of O2 into two water molecules by the flavoenzyme. Biochemistry 45, 9648–9659 (2006).
Scrutton, N.S., Berry, A. & Perham, R.N. Redesign of the coenzyme specificity of a dehydrogenase by protein engineering. Nature 343, 38–43 (1990).
Hanukoglu, I. & Gutfinger, T. cDNA sequence of adrenodoxin reductase: identification of NADP-binding sites in oxidoreductases. Eur. J. Biochem. 180, 479–484 (1989).
Wallen, J.R., Paige, C., Mallett, T.C., Karplus, P.A. & Claiborne, A. Pyridine nucleotide complexes with Bacillus anthracis coenzyme A-disulfide reductase: a structural analysis of dual NAD(P)H specificity. Biochemistry 47, 5182–5193 (2008).
Krauth-Siegel, R.L., Arscott, L.D., Schönleben-Janas, A., Schirmer, R.H. & Williams, C.H. Jr. Role of active site tyrosine residues in catalysis by human glutathione reductase. Biochemistry 37, 13968–13977 (1998).
Berry, A., Scrutton, N.S. & Perham, R.N. Switching kinetic mechanism and putative proton donor by directed mutagenesis of glutathione reductase. Biochemistry 28, 1264–1269 (1989).
Tischler, M.E., Friedrichs, D., Coll, K. & Williamson, J.R. Pyridine nucleotide distributions and enzyme mass action ratios in hepatocytes from fed and starved rats. Arch. Biochem. Biophys. 184, 222–236 (1977).
Eng, J., Lynch, R.M. & Balaban, R.S. Nicotinamide adenine dinucleotide fluorescence spectroscopy and imaging of isolated cardiac myocytes. Biophys. J. 55, 621–630 (1989).
Morais, R., Gregoire, M., Jeannotte, L. & Gravel, D. Chick embryo cells rendered respiration-deficient by chloramphenicol and ethidium bromide are auxotrophic for pyrimidines. Biochem. Biophys. Res. Commun. 94, 71–77 (1980).
King, M.P. & Attardi, G. Human cells lacking mtDNA: repopulation with exogenous mitochondria by complementation. Science 246, 500–503 (1989).
Perham, R.N., Scrutton, N.S. & Berry, A. New enzymes for old: redesigning the coenzyme and substrate specificities of glutathione reductase. BioEssays 13, 515–525 (1991).
Petschacher, B. et al. Cofactor specificity engineering of Streptococcus mutans NADH oxidase 2 for NAD(P)+ regeneration in biocatalytic oxidations. Comput. Struct. Biotechnol. J. 9, e201402005 (2014).
Rosell, A. et al. Complete reversal of coenzyme specificity by concerted mutation of three consecutive residues in alcohol dehydrogenase. J. Biol. Chem. 278, 40573–40580 (2003).
Chen, R., Greer, A. & Dean, A.M. Redesigning secondary structure to invert coenzyme specificity in isopropylmalate dehydrogenase. Proc. Natl. Acad. Sci. USA 93, 12171–12176 (1996).
Chen, R., Greer, A. & Dean, A.M. A highly active decarboxylating dehydrogenase with rationally inverted coenzyme specificity. Proc. Natl. Acad. Sci. USA 92, 11666–11670 (1995).
Wallen, J.R. et al. Structural analysis of Streptococcus pyogenes NADH oxidase: conformational dynamics involved in formation of the C(4a)-peroxyflavin intermediate. Biochemistry 54, 6815–6829 (2015).
Pai, E.F. & Schulz, G.E. The catalytic mechanism of glutathione reductase as derived from X-ray diffraction analyses of reaction intermediates. J. Biol. Chem. 258, 1752–1757 (1983).
Karplus, P.A. & Schulz, G.E. Substrate binding and catalysis by glutathione reductase as derived from refined enzyme: substrate crystal structures at 2 Å resolution. J. Mol. Biol. 210, 163–180 (1989).
Krebs, H.A. & Veech, R.L. Equilibrium relations between pyridine nucleotides and adenine nucleotides and their roles in the regulation of metabolic processes. Adv. Enzyme Regul. 7, 397–413 (1969).
Petcu, L.G. & Plaut, G.W. NADP-specific isocitrate dehydrogenase in regulation of urea synthesis in rat hepatocytes. Biochem. J. 190, 581–592 (1980).
Burch, H.B., Bradley, M.E. & Lowry, O.H. The measurement of triphosphopyridine nucleotide and reduced triphosphopyridine nucleotide and the role of hemoglobin in producing erroneous triphosphopyridine nucleotide values. J. Biol. Chem. 242, 4546–4554 (1967).
Tabor, H., Tabor, C.W. & Hafner, E.W. Convenient method for detecting 14CO2 in multiple samples: application to rapid screening for mutants. J. Bacteriol. 128, 485–486 (1976).
Carpenter, A.E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Kamentsky, L. et al. Improved structure, function and compatibility for CellProfiler: modular high-throughput image analysis software. Bioinformatics 27, 1179–1180 (2011).
Acknowledgements
We thank D. Trono (Swiss Federal Institute of Technology Lausanne) for providing reagents. This work was supported by grants R01GM099683 and R35GM122455 from the National Institutes of Health and by the Harvard Ludwig Center. This research used resources of the Advanced Light Source, which is a DOE Office of Science User Facility under contract no. DE-AC02-05CH11231. Metabolomics analysis was performed at the Metabolomics Core Facility at the University of Utah, which is supported by 1 S10 OD016232-01, 1 S10 OD021505-01 and 1 U54 DK110858-01. V.K.M. is supported as an Investigator of the Howard Hughes Medical Institute from the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
V.C., D.V.T., Z.G. and V.K.M. designed the study and interpreted data. V.C. and Z.G. performed the biochemical and structure-related experiments. D.V.T., V.C. and H.S. performed cell-based experiments. V.C., D.V.T., Z.G. and V.K.M. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
V.C., D.T., Z.G. and V.K.M. are listed as inventors on a patent application filed by Massachusetts General Hospital on the LbNOX and TPNOX technology.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1 and 2 and Supplementary Figures 1–6 (PDF 2213 kb)
Supplementary Data Set 1
GC–MS data (XLSX 74 kb)
Rights and permissions
About this article
Cite this article
Cracan, V., Titov, D., Shen, H. et al. A genetically encoded tool for manipulation of NADP+/NADPH in living cells. Nat Chem Biol 13, 1088–1095 (2017). https://doi.org/10.1038/nchembio.2454
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2454
This article is cited by
-
SLC7A11 expression level dictates differential responses to oxidative stress in cancer cells
Nature Communications (2023)
-
A genetically encoded tool to increase cellular NADH/NAD+ ratio in living cells
Nature Chemical Biology (2023)
-
Reporting NADPH fluxes
Nature Chemical Biology (2023)
-
Cytosolic and mitochondrial NADPH fluxes are independently regulated
Nature Chemical Biology (2023)
-
NERNST: a genetically-encoded ratiometric non-destructive sensing tool to estimate NADP(H) redox status in bacterial, plant and animal systems
Nature Communications (2023)