Abstract
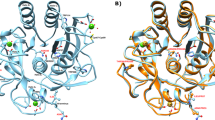
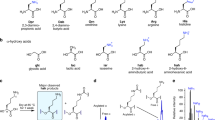
Miniproteins simplify the protein-folding problem, allowing the dissection of forces that stabilize protein structures. Here we describe PPα-Tyr, a designed peptide comprising an α-helix buttressed by a polyproline II helix. PPα-Tyr is water soluble and monomeric, and it unfolds cooperatively with a midpoint unfolding temperature (TM) of 39 °C. NMR structures of PPα-Tyr reveal proline residues docked between tyrosine side chains, as designed. The stability of PPα is sensitive to modifications in the aromatic residues: replacing tyrosine with phenylalanine, i.e., changing three solvent-exposed hydroxyl groups to protons, reduces the TM to 20 °C. We attribute this result to the loss of CH–π interactions between the aromatic and proline rings, which we probe by substituting the aromatic residues with nonproteinogenic side chains. In analyses of natural protein structures, we find a preference for proline–tyrosine interactions over other proline-containing pairs, and observe abundant CH–π interactions in biologically important complexes between proline-rich ligands and SH3 and similar domains.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Nick Pace, C., Scholtz, J.M. & Grimsley, G.R. Forces stabilizing proteins. FEBS Lett. 588, 2177–2184 (2014).
Dougherty, D.A. Cation-π interactions in chemistry and biology: a new view of benzene, Phe, Tyr, and Trp. Science 271, 163–168 (1996).
Brandl, M., Weiss, M.S., Jabs, A., Sühnel, J. & Hilgenfeld, R. C-H...π-interactions in proteins. J. Mol. Biol. 307, 357–377 (2001).
Bartlett, G.J., Choudhary, A., Raines, R.T. & Woolfson, D.N. n→π* interactions in proteins. Nat. Chem. Biol. 6, 615–620 (2010).
Yesselman, J.D., Horowitz, S., Brooks, C.L., III & Trievel, R.C. Frequent side chain methyl carbon-oxygen hydrogen bonding in proteins revealed by computational and stereochemical analysis of neutron structures. Proteins 83, 403–410 (2015).
Bhattacharya, A., Tejero, R. & Montelione, G.T. Evaluating protein structures determined by structural genomics consortia. Proteins 66, 778–795 (2007).
Bartlett, G.J., Newberry, R.W., VanVeller, B., Raines, R.T. & Woolfson, D.N. Interplay of hydrogen bonds and n→π* interactions in proteins. J. Am. Chem. Soc. 135, 18682–18688 (2013).
Zondlo, N.J. & Schepartz, A. Highly specific DNA recognition by a designed iniature protein. J. Am. Chem. Soc. 121, 6938–6939 (1999).
Gellman, S.H. & Woolfson, D.N. Mini-proteins Trp the light fantastic. Nat. Struct. Biol. 9, 408–410 (2002).
Neidigh, J.W., Fesinmeyer, R.M. & Andersen, N.H. Designing a 20-residue protein. Nat. Struct. Biol. 9, 425–430 (2002).
Craven, T.W., Cho, M.-K., Traaseth, N.J., Bonneau, R. & Kirshenbaum, K. A miniature protein stabilized by a cation-π interaction network. J. Am. Chem. Soc. 138, 1543–1550 (2016).
Woolfson, D.N. The design of coiled-coil structures and assemblies. Adv. Protein Chem. 70, 79–112 (2005).
Golemi-Kotra, D. et al. High affinity, paralog-specific recognition of the Mena EVH1 domain by a miniature protein. J. Am. Chem. Soc. 126, 4–5 (2004).
Daly, N.L. & Craik, D.J. Bioactive cystine knot proteins. Curr. Opin. Chem. Biol. 15, 362–368 (2011).
Bhardwaj, G. et al. Accurate de novo design of hyperstable constrained peptides. Nature 538, 329–335 (2016).
Pabo, C.O., Peisach, E. & Grant, R.A. Design and selection of novel Cys2His2 zinc finger proteins. Annu. Rev. Biochem. 70, 313–340 (2001).
Gifford, J.L., Walsh, M.P. & Vogel, H.J. Structures and metal-ion-binding properties of the Ca2+-binding helix-loop-helix EF-hand motifs. Biochem. J. 405, 199–221 (2007).
McKnight, C.J., Matsudaira, P.T. & Kim, P.S. NMR structure of the 35-residue villin headpiece subdomain. Nat. Struct. Biol. 4, 180–184 (1997).
Cochran, A.G., Skelton, N.J. & Starovasnik, M.A. Tryptophan zippers: stable, monomeric β -hairpins. Proc. Natl. Acad. Sci. USA 98, 5578–5583 (2001).
Barua, B. et al. The Trp-cage: optimizing the stability of a globular miniprotein. Protein Eng. Des. Sel. 21, 171–185 (2008).
Baker, E.G. et al. Local and macroscopic electrostatic interactions in single α-helices. Nat. Chem. Biol. 11, 221–228 (2015).
Crick, F.H.C. The packing of α-Helices: simple coiled-coils. Acta Crystallogr. 6, 689–697 (1953).
Walshaw, J. & Woolfson, D.N. Socket: a program for identifying and analysing coiled-coil motifs within protein structures. J. Mol. Biol. 307, 1427–1450 (2001).
Woolfson, D.N. et al. De novo protein design: how do we expand into the universe of possible protein structures? Curr. Opin. Struct. Biol. 33, 16–26 (2015).
Plevin, M.J., Bryce, D.L. & Boisbouvier, J. Direct detection of CH–π interactions in proteins. Nat. Chem. 2, 466–471 (2010).
Nishio, M., Umezawa, Y., Fantini, J., Weiss, M.S. & Chakrabarti, P. CH–π hydrogen bonds in biological macromolecules. Phys. Chem. Chem. Phys. 16, 12648–12683 (2014).
Blundell, T.L., Pitts, J.E., Tickle, I.J., Wood, S.P. & Wu, C.-W. X-ray analysis (1. 4-A resolution) of avian pancreatic polypeptide: Small globular protein hormone. Proc. Natl. Acad. Sci. USA 78, 4175–4179 (1981).
Larson, M.R. et al. Elongated fibrillar structure of a streptococcal adhesin assembled by the high-affinity association of α- and PPII-helices. Proc. Natl. Acad. Sci. USA 107, 5983–5988 (2010).
Noelken, M.E., Chang, P.J. & Kimmel, J.R. Conformation and association of pancreatic polypeptide from three species. Biochemistry 19, 1838–1843 (1980).
Fiser, A., Do, R.K.G. & Šali, A. Modeling of loops in protein structures. Protein Sci. 9, 1753–1773 (2000).
Hu, X., Wang, H., Ke, H. & Kuhlman, B. High-resolution design of a protein loop. Proc. Natl. Acad. Sci. USA 104, 17668–17673 (2007).
Hudson, K.L. et al. Carbohydrate-aromatic interactions in proteins. J. Am. Chem. Soc. 137, 15152–15160 (2015).
Saha, R.P., Bhattacharyya, R. & Chakrabarti, P. Interaction geometry involving planar groups in protein–protein interfaces. Proteins 67, 84–97 (2007).
Wheeler, S.E. & Houk, K.N. Through-space effects of substituents dominate molecular electrostatic potentials of substituted arenes. J. Chem. Theory Comput. 5, 2301–2312 (2009).
Carver, F.J., Hunter, C.A. & Seward, E.M. Struacture–activity relationship for quantifying aromatic interactions. Chem. Commun. (Camb.) 1998, 775–776 (1998).
Zondlo, N.J. Aromatic-proline interactions: electronically tunable CH–π interactions. Acc. Chem. Res. 46, 1039–1049 (2013).
Bloom, J.W.G., Raju, R.K. & Wheeler, S.E. Physical nature of substituent effects in XH/π interactions. J. Chem. Theory Comput. 8, 3167–3174 (2012).
Tsuzuki, S., Honda, K., Uchimaru, T., Mikami, M. & Tanabe, K. The magnitude of the CH/π interaction between benzene and some model hydrocarbons. J. Am. Chem. Soc. 122, 3746–3753 (2000).
Kay, B.K., Williamson, M.P. & Sudol, M. The importance of being proline: the interaction of proline-rich motifs in signaling proteins with their cognate domains. FASEB J. 14, 231–241 (2000).
Kay, B.K. SH3 domains come of age. FEBS Lett. 586, 2606–2608 (2012).
Ball, L.J., Kühne, R., Schneider-Mergener, J. & Oschkinat, H. Recognition of proline-rich motifs by protein–protein-interaction domains. Angew. Chem. Int. Ed. Engl. 44, 2852–2869 (2005).
Parsons, Z.D., Bland, J.M., Mullins, E.A. & Eichman, B.F. A catalytic role for C-H/π interactions in base excision repair by Bacillus cereus DNA glycosylase AlkD. J. Am. Chem. Soc. 138, 11485–11488 (2016).
Zhang, Y., Malamakal, R.M. & Chenoweth, D.M. Aza-Glycine induces collagen hyperstability. J. Am. Chem. Soc. 137, 12422–12425 (2015).
Arnold, U. & Raines, R.T. Replacing a single atom accelerates the folding of a protein and increases its thermostability. Org. Biomol. Chem. 14, 6780–6785 (2016).
Cobos, E.S. et al. A miniprotein scaffold used to assemble the polyproline II binding epitope recognized by SH3 domains. J. Mol. Biol. 342, 355–365 (2004).
Velankar, S. et al. PDBe: Protein Data Bank in Europe. Nucleic Acids Res. 38, D308–D317 (2010).
Wang, G. & Dunbrack, R.L. Jr. PISCES: recent improvements to a PDB sequence culling server. Nucleic Acids Res. 33, W94–W98 (2005).
Singh, J. & Thornton, J.M. Atlas of Protein Side-Chain Interactions Vol. 1 (Oxford University Press, 1992).
Finn, R.D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Fox, N.K., Brenner, S.E. & Chandonia, J.M. SCOPe: Structural Classification of Proteins--extended, integrating SCOP and ASTRAL data and classification of new structures. Nucleic Acids Res. 42, D304–D309 (2014).
Word, J.M., Lovell, S.C., Richardson, J.S. & Richardson, D.C. Asparagine and glutamine: using hydrogen atom contacts in the choice of side chain amide orientation. J. Mol. Biol. 285, 1735–1747 (1999).
Kuipers, B.J.H. & Gruppen, H. Prediction of molar extinction coefficients of proteins and peptides using UV absorption of the constituent amino acids at 214 nm to enable quantitative reverse phase high-performance liquid chromatography–mass spectrometry analysis. J. Agric. Food Chem. 55, 5445–5451 (2007).
Marky, L.A. & Breslauer, K.J. Calculating thermodynamic data for transitions of any molecularity from equilibrium melting curves. Biopolymers 26, 1601–1620 (1987).
LiCata, V.J. & Liu, C.-C. in Methods in Enzymology: Biothermodynamics, Part C (Eds. Johnson, M.L., Holt, J.M., & Ackers, G.K.) Vol. 488, 219–238 (Academic Press, 2011).
Vranken, W.F. et al. The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59, 687–696 (2005).
Rieping, W. et al. ARIA2: automated NOE assignment and data integration in NMR structure calculation. Bioinformatics 23, 381–382 (2007).
Brünger, A.T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Cheung, M.S., Maguire, M.L., Stevens, T.J. & Broadhurst, R.W. DANGLE: A Bayesian inferential method for predicting protein backbone dihedral angles and secondary structure. J. Magn. Reson. 202, 223–233 (2010).
Touw, W.G. et al. A series of PDB-related databanks for everyday needs. Nucleic Acids Res. 43, D364–D368 (2015).
Mansiaux, Y., Joseph, A.P., Gelly, J.C. & de Brevern, A.G. Assignment of PolyProline II conformation and analysis of sequence–structure relationship. PLoS One 6, e18401 (2011).
Acknowledgements
E.G.B. and D.N.W. are supported by a Biotechnology and Biological Sciences Research Council (BBSRC)/ERASynBio grant (BB/M005615/1); K.L.H., G.J.B., J.W.H. and D.N.W. are supported by the ERC (340764); D.N.W. is a Royal Society Wolfson Research Merit Award holder (WM140008); and K.L.P.G. is supported by the Engineering and Physical Sciences Research Council (EPSRC)-funded Bristol Chemical Synthesis Centre for Doctoral Training (EP/G036764/1). We thank BrisSynBio for access to the BBSRC/EPSRC-funded 700 MHz NMR spectrometer (BB/L01386X/1), S. Whittaker and the Henry Wellcome Building NMR Facility at the University of Birmingham for access to the Wellcome Trust–funded 900 MHz spectrometer (099185/Z/12/Z), and R. Alder and members of the Woolfson group for helpful discussions.
Author information
Authors and Affiliations
Contributions
E.G.B. and D.N.W. designed the research. E.G.B. made the synthetic peptides and performed the CD spectroscopy and AUC experiments. C.W. and M.P.C. collected the NMR data. C.W., K.L.P.G. and E.G.B. analyzed the NMR data, and C.W. solved the NMR structures. K.L.H., G.J.B., D.N.W. conducted the bioinformatics. E.G.B. and R.B.S. carried out the molecular-dynamics studies. J.W.H. and E.G.B. performed the van't Hoff analyses. E.G.B. and D.N.W. wrote the paper. All authors reviewed and contributed to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–5 and Supplementary Figures 1–10 (PDF 5216 kb)
Rights and permissions
About this article
Cite this article
Baker, E., Williams, C., Hudson, K. et al. Engineering protein stability with atomic precision in a monomeric miniprotein. Nat Chem Biol 13, 764–770 (2017). https://doi.org/10.1038/nchembio.2380
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2380
This article is cited by
-
Hyperthermostable cube-shaped assembly in water
Communications Chemistry (2018)