Abstract
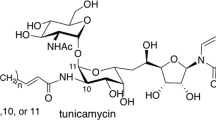
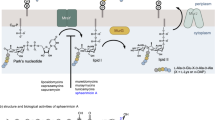
The rapid increase of antibiotic resistance has created an urgent need to develop novel antimicrobial agents. Here we describe the crystal structure of the promising bacterial target phospho-N-acetylmuramoyl–pentapeptide translocase (MraY) in complex with the nucleoside antibiotic tunicamycin. The structure not only reveals the mode of action of several related natural-product antibiotics but also gives an indication on the binding mode of the MraY UDP–MurNAc–pentapeptide and undecaprenyl-phosphate substrates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Boyle, D.S. & Donachie, W.D. J. Bacteriol. 180, 6429–6432 (1998).
Anderson, J.S., Matsuhashi, M., Haskin, M.A. & Strominger, J.L. Proc. Natl. Acad. Sci. USA 53, 881–889 (1965).
Struve, W.G. & Neuhaus, F.C. Biochem. Biophys. Res. Commun. 18, 6–12 (1965).
Lehrman, M.A. Glycobiology 4, 768–771 (1994).
Lehrman, M.A. Glycobiology 1, 553–562 (1991).
Anderson, M.S., Eveland, S.S. & Price, N.P. FEMS Microbiol. Lett. 191, 169–175 (2000).
Heifetz, A. & Elbein, A.D. J. Biol. Chem. 252, 3057–3063 (1977).
Dini, C. Curr. Top. Med. Chem. 5, 1221–1236 (2005).
Tkacz, J.S. & Lampen, O. Biochem. Biophys. Res. Commun. 65, 248–257 (1975).
Tamura, G., Sasaki, T., Matsuhashi, M., Takatsuki, A. & Yamasaki, M. Agric. Biol. Chem. 40, 447–449 (1976).
Schwarz, R.T. & Datema, R. Trends Biochem. Sci. 5, 65–67 (1980).
Brandish, P.E. et al. Antimicrob. Agents Chemother. 40, 1640–1644 (1996).
Keller, R.K., Adair, W.L. Jr. & Ness, G.C. J. Biol. Chem. 254, 9966–9969 (1979).
Chung, B.C. et al. Science 341, 1012–1016 (2013).
Chung, B.C. et al. Nature 533, 557–560 (2016).
Shapiro, A.B., Jahić, H., Gao, N., Hajec, L. & Rivin, O. J. Biomol. Screen. 17, 662–672 (2012).
Al-Dabbagh, B. et al. Biochemistry 47, 8919–8928 (2008).
Al-Dabbagh, B. et al. Biochimie 127, 249–257 (2016).
Suami, T., Sasai, H., Matsuno, K. & Suzuki, N. Carbohydr. Res. 143, 85–96 (1985).
Myers, A.G., Gin, D.Y. & Rogers, D.H. J. Am. Chem. Soc. 116, 4697–4718 (1994).
Li, J. & Yu, B. Angew. Chem. Int. Edn Engl. 54, 6618–6621 (2015).
Wyszynski, F.J. et al. Nat. Chem. 4, 539–546 (2012).
Backmark, A.E. et al. Protein Sci. 22, 1124–1132 (2013).
Studier, F.W. Protein Expr. Purif. 41, 207–234 (2005).
Bowler, M.W. et al. Acta Crystallogr. D Biol. Crystallogr. 66, 855–864 (2010).
Kabsch, W. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
McCoy, A.J. et al. J. Appl. Crystallogr. 40, 658–674 (2007).
Cowtan, K., Zhang, K.Y.J. & Main, P. International Tables for Crystallography, Crystallography of Biological Macromolecules (Kluwer Academic Publishers, 2001).
Cowtan, K. Acta Crystallogr. D Biol. Crystallogr. 66, 470–478 (2010).
Cowtan, K. Acta Crystallogr. D Biol. Crystallogr. 64, 83–89 (2008).
de La Fortelle, E. & Bricogne, G. Methods Enzymol. 276, 472–494 (1997).
Emsley, P. & Cowtan, K. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Gasteiger, J., Rudolph, C. & Sadowski, J. Tetrahedron Comp. Method. 3, 537–547 (1990).
Acknowledgements
We thank A. Shapiro and J. Bernström for assistance with activity measurements, beamline scientists at ESRF (Grenoble, France) and Diamond Light Source (Didcot, UK) for assistance during data collection, and members of Global Phasing Ltd. for help with STARANISO. We are also grateful to I. Moraes at the Membrane Protein Laboratory, Diamond for providing beamtime and assistance. This work was supported by the European Union under the programme FP7-PEOPLE-2011-ITN NanoMem, project number 317079 (M.E.) and the European Union's Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie agreement No 637295, X-probe (M.E.).
Author information
Authors and Affiliations
Contributions
A.S. and M.E. designed the study, J.K.H. established purification and crystallized the protein, J.H. collected and analyzed the activity data, J.K.H. and P.J. collected the crystallographic data, P.J. solved and built the initial model, P.J. and G.B. refined the structure, H.C. performed the docking studies, J.K.H., G.B., M.E. and P.J. prepared the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
J.H., H.C., A.S., M.E. and P.J. are employees of AstraZeneca.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–8 and Supplementary Tables 1 and 2. (PDF 2748 kb)
Rights and permissions
About this article
Cite this article
Hakulinen, J., Hering, J., Brändén, G. et al. MraY–antibiotic complex reveals details of tunicamycin mode of action. Nat Chem Biol 13, 265–267 (2017). https://doi.org/10.1038/nchembio.2270
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2270
This article is cited by
-
The tunicamycin derivative TunR2 exhibits potent antibiotic properties with low toxicity in an in vivo Mycobacterium marinum-zebrafish TB infection model
The Journal of Antibiotics (2024)
-
Synthesis of lipid-linked precursors of the bacterial cell wall is governed by a feedback control mechanism in Pseudomonas aeruginosa
Nature Microbiology (2024)
-
The application of Aptamer in biomarker discovery
Biomarker Research (2023)
-
Unrealized targets in the discovery of antibiotics for Gram-negative bacterial infections
Nature Reviews Drug Discovery (2023)
-
Peptidoglycan biosynthesis is driven by lipid transfer along enzyme-substrate affinity gradients
Nature Communications (2022)