Abstract
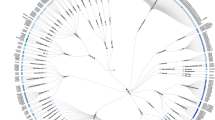
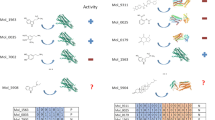
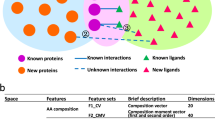
Understanding the pharmacological similarity of G protein–coupled receptors (GPCRs) is paramount for predicting ligand off-target effects, drug repurposing, and ligand discovery for orphan receptors. Phylogenetic relationships do not always correctly capture pharmacological similarity. Previous family-wide attempts to define pharmacological relationships were based on three-dimensional structures and/or known receptor–ligand pairings, both unavailable for orphan GPCRs. Here, we present GPCR–CoINPocket, a novel contact-informed neighboring pocket metric of GPCR binding-site similarity that is informed by patterns of ligand–residue interactions observed in crystallographically characterized GPCRs. GPCR–CoINPocket is applicable to receptors with unknown structure or ligands and accurately captures known pharmacological relationships between GPCRs, even those undetected by phylogeny. When applied to orphan receptor GPR37L1, GPCR–CoINPocket identified its pharmacological neighbors, and transfer of their pharmacology aided in discovery of the first surrogate ligands for this orphan with a 30% success rate. Although primarily designed for GPCRs, the method is easily transferable to other protein families.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
01 March 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41589-021-00746-1
References
Davenport, A.P. et al. International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands. Pharmacol. Rev. 65, 967–986 (2013).
Southern, C. et al. Screening β-arrestin recruitment for the identification of natural ligands for orphan G-protein-coupled receptors. J. Biomol. Screen. 18, 599–609 (2013).
Jenkins, L. et al. Identification of novel species-selective agonists of the G-protein–coupled receptor GPR35 that promote recruitment of β-arrestin-2 and activate Gα13. Biochem. J. 432, 451–459 (2010).
Kroeze, W.K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369 (2015).
Ngo, T. et al. Identifying ligands at orphan GPCRs: current status using structure-based approaches. Br. J. Pharmacol. 173, 2934–2951 (2016).
Fredriksson, R., Lagerström, M.C., Lundin, L.G. & Schiöth, H.B. The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol. Pharmacol. 63, 1256–1272 (2003).
Schiöth, H.B. & Fredriksson, R. The GRAFS classification system of G-protein coupled receptors in comparative perspective. Gen. Comp. Endocrinol. 142, 94–101 (2005).
Guba, W. et al. From astemizole to a novel hit series of small-molecule somatostatin 5 receptor antagonists via GPCR affinity profiling. J. Med. Chem. 50, 6295–6298 (2007).
Martin, R.E., Green, L.G., Guba, W., Kratochwil, N. & Christ, A. Discovery of the first nonpeptidic, small-molecule, highly selective somatostatin receptor subtype 5 antagonists: a chemogenomics approach. J. Med. Chem. 50, 6291–6294 (2007).
Scholten, D.J. et al. Pharmacological modulation of chemokine receptor function. Br. J. Pharmacol. 165, 1617–1643 (2012).
Lin, H., Sassano, M.F., Roth, B.L. & Shoichet, B.K. A pharmacological organization of G protein–coupled receptors. Nat. Methods 10, 140–146 (2013).
van der Horst, E. et al. A novel chemogenomics analysis of G protein-coupled receptors (GPCRs) and their ligands: a potential strategy for receptor de-orphanization. BMC Bioinformatics 11, 316 (2010).
Surgand, J.S., Rodrigo, J., Kellenberger, E. & Rognan, D. A chemogenomic analysis of the transmembrane binding cavity of human G-protein–coupled receptors. Proteins 62, 509–538 (2006).
Jacoby, E. A novel chemogenomics knowledge-based ligand design strategy—application to G protein-coupled receptors. Mol. Inform. 20, 115–123 (2001).
Gloriam, D.E., Foord, S.M., Blaney, F.E. & Garland, S.L. Definition of the G protein–coupled receptor transmembrane bundle binding pocket and calculation of receptor similarities for drug design. J. Med. Chem. 52, 4429–4442 (2009).
Kufareva, I., Ilatovskiy, A.V. & Abagyan, R. Pocketome: an encyclopedia of small-molecule binding sites in 4D. Nucleic Acids Res. 40, D535–D540 (2012).
Venkatakrishnan, A.J. et al. Molecular signatures of G protein-coupled receptors. Nature 494, 185–194 (2013).
Isberg, V. et al. Generic GPCR residue numbers - aligning topology maps while minding the gaps. Trends Pharmacol. Sci. 36, 22–31 (2015).
Nandigam, R.K., Kim, S., Singh, J. & Chuaqui, C. Position specific interaction dependent scoring technique for virtual screening based on weighted protein–ligand interaction fingerprint profiles. J. Chem. Inf. Model. 49, 1185–1192 (2009).
Marcou, G. & Rognan, D. Optimizing fragment and scaffold docking by use of molecular interaction fingerprints. J. Chem. Inf. Model. 47, 195–207 (2007).
Sokal, R.R. & Michener, C.D. A statistical method for evaluating systematic relationships. Univ. Kans. Sci. Bull. 38, 1409–1438 (1958).
Keiser, M.J. et al. Predicting new molecular targets for known drugs. Nature 462, 175–181 (2009).
Modi, M.E. et al. Melanocortin receptor agonists facilitate oxytocin-dependent partner preference formation in the prairie vole. Neuropsychopharmacology 40, 1856–1865 (2015).
Bento, A.P. et al. The ChEMBL bioactivity database: an update. Nucleic Acids Res. 42, D1083–D1090 (2014).
Kufareva, I., Katritch, V., Stevens, R.C. & Abagyan, R. Advances in GPCR modeling evaluated by the GPCR Dock 2013 assessment: meeting new challenges. Structure 22, 1120–1139 (2014).
Kufareva, I., Rueda, M., Katritch, V., Stevens, R.C. & Abagyan, R. Status of GPCR modeling and docking as reflected by community-wide GPCR Dock 2010 assessment. Structure 19, 1108–1126 (2011).
Thelen, M. & Thelen, S. CXCR7, CXCR4 and CXCL12: an eccentric trio? J. Neuroimmunol. 198, 9–13 (2008).
Min, K.D. et al. Identification of genes related to heart failure using global gene expression profiling of human failing myocardium. Biochem. Biophys. Res. Commun. 393, 55–60 (2010).
Marazziti, D. et al. Precocious cerebellum development and improved motor functions in mice lacking the astrocyte cilium-, patched 1-associated Gpr37l1 receptor. Proc. Natl. Acad. Sci. USA 110, 16486–16491 (2013).
Smith, N.J. Drug discovery opportunities at the endothelin B receptor-related orphan G protein–coupled receptors, GPR37 and GPR37L1. Front. Pharmacol. 6, 275 (2015).
Leng, N., Gu, G., Simerly, R.B. & Spindel, E.R. Molecular cloning and characterization of two putative G protein-coupled receptors which are highly expressed in the central nervous system. Brain Res. Mol. Brain Res. 69, 73–83 (1999).
Valdenaire, O. et al. A new family of orphan G protein-coupled receptors predominantly expressed in the brain. FEBS Lett. 424, 193–196 (1998).
Shihoya, W. et al. Activation mechanism of endothelin ETB receptor by endothelin-1. Nature 537, 363–368 (2016).
Coleman, J.L. et al. Metalloprotease cleavage of the N terminus of the orphan G protein-coupled receptor GPR37L1 reduces its constitutive activity. Sci. Signal. 9, ra36 (2016).
Ngo, T., Coleman, J.L. & Smith, N.J. Using constitutive activity to define appropriate high-throughput screening assays for orphan G protein-coupled receptors. Methods Mol. Biol. 1272, 91–106 (2015).
Sassano, M.F., Doak, A.K., Roth, B.L. & Shoichet, B.K. Colloidal aggregation causes inhibition of G protein-coupled receptors. J. Med. Chem. 56, 2406–2414 (2013).
Rezgaoui, M. et al. The neuropeptide head activator is a high-affinity ligand for the orphan G-protein-coupled receptor GPR37. J. Cell Sci. 119, 542–549 (2006).
Civelli, O. et al. G protein-coupled receptor deorphanizations. Annu. Rev. Pharmacol. Toxicol. 53, 127–146 (2013).
Isberg, V. et al. Computer-aided discovery of aromatic L-α-amino acids as agonists of the orphan G protein-coupled receptor GPR139. J. Chem. Inf. Model. 54, 1553–1557 (2014).
Liu, C. et al. GPR139, an orphan receptor highly enriched in the habenula and septum, is activated by the essential amino acids L-tryptophan and L-phenylalanine. Mol. Pharmacol. 88, 911–925 (2015).
Huang, X.P. et al. Allosteric ligands for the pharmacologically dark receptors GPR68 and GPR65. Nature 527, 477–483 (2015).
Tagat, J.R. et al. Piperazine-based CCR5 antagonists as HIV-1 inhibitors. I: 2(S)-methyl piperazine as a key pharmacophore element. Bioorg. Med. Chem. Lett. 11, 2143–2146 (2001).
Alexander, S.P. et al. The concise guide to PHARMACOLOGY 2015/16: G protein–coupled receptors. Br. J. Pharmacol. 172, 5744–5869 (2015).
Kufareva, I. & Abagyan, R. in Homology Modelling: Methods and Protocols Vol. 857 (eds Orry, A.J.W. & Abagyan, R.) Ch. 10 (Humana Press, 2012).
McRobb, F.M., Sahagún, V., Kufareva, I. & Abagyan, R. In silico analysis of the conservation of human toxicity and endocrine disruption targets in aquatic species. Environ. Sci. Technol. 48, 1964–1972 (2014).
Gonnet, G.H., Cohen, M.A. & Benner, S.A. Exhaustive matching of the entire protein sequence database. Science 256, 1443–1445 (1992).
Abagyan, R., Totrov, M. & Kuznetsov, D. ICM—A new method for protein modeling and design: applications to docking and structure prediction from the distorted native conformation. J. Comput. Chem. 15, 488–506 (1994).
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Abagyan, R., Chen, W. & Kufareva, I. in Computational Approaches to Nuclear Receptors (eds. Cozzini, P. & Kellogg, G.E.) 84–109 (The Royal Society of Chemistry, 2012).
Acknowledgements
This work was supported in part by an Endeavour Research Fellowship from the Australian Government to T.N., an NHMRC & NHF CJ Martin Fellowship (N.J.S.), NHMRC Program Grants 573732 and 1074386 (R.M.G.), and Australian Postgraduate Awards (T.N. & J.L.J.C.). A.G.S. is supported by NHMRC Early Career Fellowship 1090408. R.A., I.K. and A.V.I. are supported by the NIH Grant R01 GM071872. I.K. is supported by R01 AI118985, R01 GM117424, R21 AI121918 and R21 AI122211. N.J.S. thanks the Mostyn Family Foundation for philanthropic support.
Author information
Authors and Affiliations
Contributions
T.N., F.M.M., R.A., I.K. and N.J.S. conceived the study; T.N. and I.K. wrote the code for the weighted alignment scores with assistance from A.V.I. and R.A.; A.V.I. generated the ChEMBL data set, which was analyzed by T.N., A.V.I. and I.K.; A.G.S. performed dynamic light scattering experiments and analyzed the data; T.N. and N.J.S. designed and performed reporter gene assay experiments; R.M.G. and J.L.J.C. provided key reagents and ideas and R.P.R. provided the initial transmembrane alignments of all class A GPCRs; N.J.S. provided overall supervision of the project; T.N., I.K. & N.J.S. wrote the manuscript. All authors have provided input and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Figures 1–7. (PDF 4543 kb)
Supplementary Dataset 1
Pairwise comparison of GPCR–CoINPocket scores between Class A GPCRs. (XLSX 23959 kb)
Supplementary Dataset 2
List of class A G protein–coupled receptors entries in the Pocketome. (XLSX 26281 kb)
Supplementary Dataset 3
Raw pairwise contact strength profiled binding site similarities between class A GPCRs. (XLSX 10 kb)
About this article
Cite this article
Ngo, T., Ilatovskiy, A., Stewart, A. et al. RETRACTED ARTICLE: Orphan receptor ligand discovery by pickpocketing pharmacological neighbors. Nat Chem Biol 13, 235–242 (2017). https://doi.org/10.1038/nchembio.2266
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2266
This article is cited by
-
Building the Chordata Olfactory Receptor Database using more than 400,000 receptors annotated by Genome2OR
Science China Life Sciences (2022)
-
Benchmarking GPCR homology model template selection in combination with de novo loop generation
Journal of Computer-Aided Molecular Design (2020)
-
Molecular pharmacology of metabotropic receptors targeted by neuropsychiatric drugs
Nature Structural & Molecular Biology (2019)
-
Computational advances in combating colloidal aggregation in drug discovery
Nature Chemistry (2019)
-
Novel pharmacological maps of protein lysine methyltransferases: key for target deorphanization
Journal of Cheminformatics (2018)