Abstract
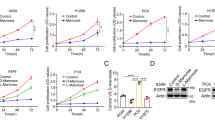
Asparagine (N)-linked glycosylation is a protein modification critical for glycoprotein folding, stability, and cellular localization. To identify small molecules that inhibit new targets in this biosynthetic pathway, we initiated a cell-based high-throughput screen and lead-compound-optimization campaign that delivered a cell-permeable inhibitor, NGI-1. NGI-1 targets oligosaccharyltransferase (OST), a hetero-oligomeric enzyme that exists in multiple isoforms and transfers oligosaccharides to recipient proteins. In non-small-cell lung cancer cells, NGI-1 blocks cell-surface localization and signaling of the epidermal growth factor receptor (EGFR) glycoprotein, but selectively arrests proliferation in only those cell lines that are dependent on EGFR (or fibroblast growth factor, FGFR) for survival. In these cell lines, OST inhibition causes cell-cycle arrest accompanied by induction of p21, autofluorescence, and cell morphology changes, all hallmarks of senescence. These results identify OST inhibition as a potential therapeutic approach for treating receptor-tyrosine-kinase-dependent tumors and provides a chemical probe for reversibly regulating N-linked glycosylation in mammalian cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Zielinska, D.F., Gnad, F., Wiśniewski, J.R. & Mann, M. Precision mapping of an in vivo N-glycoproteome reveals rigid topological and sequence constraints. Cell 141, 897–907 (2010).
Freeze, H.H. Genetic defects in the human glycome. Nat. Rev. Genet. 7, 537–551 (2006).
Lehrman, M.A. Teaching dolichol-linked oligosaccharides more tricks with alternatives to metabolic radiolabeling. Glycobiology 17, 75R–85R (2007).
Takatsuki, A., Arima, K. & Tamura, G. Tunicamycin, a new antibiotic. I. Isolation and characterization of tunicamycin. J. Antibiot. (Tokyo) 24, 215–223 (1971).
Helenius, J. et al. Translocation of lipid-linked oligosaccharides across the ER membrane requires Rft1 protein. Nature 415, 447–450 (2002).
Frank, C.G., Sanyal, S., Rush, J.S., Waechter, C.J. & Menon, A.K. Does Rft1 flip an N-glycan lipid precursor? Nature 454, E3–E4 (2008).
Jelk, J. et al. Glycoprotein biosynthesis in a eukaryote lacking the membrane protein Rft1. J. Biol. Chem. 288, 20616–20623 (2013).
Aebi, M. N-linked protein glycosylation in the ER. Biochim. Biophys. Acta 1833, 2430–2437 (2013).
Kelleher, D.J. & Gilmore, R. An evolving view of the eukaryotic oligosaccharyltransferase. Glycobiology 16, 47R–62R (2006).
Pfeffer, S. et al. Structure of the mammalian oligosaccharyl-transferase complex in the native ER protein translocon. Nat. Commun. 5, 3072 (2014).
Reiss, G., te Heesen, S., Gilmore, R., Zufferey, R. & Aebi, M. A specific screen for oligosaccharyltransferase mutations identifies the 9 kDa OST5 protein required for optimal activity in vivo and in vitro. EMBO J. 16, 1164–1172 (1997).
Zufferey, R. et al. STT3, a highly conserved protein required for yeast oligosaccharyl transferase activity in vivo. EMBO J. 14, 4949–4960 (1995).
Schwarz, F. & Aebi, M. Mechanisms and principles of N-linked protein glycosylation. Curr. Opin. Struct. Biol. 21, 576–582 (2011).
Ruiz-Canada, C., Kelleher, D.J. & Gilmore, R. Cotranslational and posttranslational N-glycosylation of polypeptides by distinct mammalian OST isoforms. Cell 136, 272–283 (2009).
Shrimal, S. & Gilmore, R. Glycosylation of closely spaced acceptor sites in human glycoproteins. J. Cell Sci. 126, 5513–5523 (2013).
Cherepanova, N.A., Shrimal, S. & Gilmore, R. Oxidoreductase activity is necessary for N-glycosylation of cysteine-proximal acceptor sites in glycoproteins. J. Cell Biol. 206, 525–539 (2014).
Roboti, P. & High, S. The oligosaccharyltransferase subunits OST48, DAD1 and KCP2 function as ubiquitous and selective modulators of mammalian N-glycosylation. J. Cell Sci. 125, 3474–3484 (2012).
Blomen, V.A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
Freeze, H.H. & Aebi, M. Altered glycan structures: the molecular basis of congenital disorders of glycosylation. Curr. Opin. Struct. Biol. 15, 490–498 (2005).
Contessa, J.N. et al. Molecular imaging of N-linked glycosylation suggests glycan biosynthesis is a novel target for cancer therapy. Clin. Cancer Res. 16, 3205–3214 (2010).
Dennis, J.W., Nabi, I.R. & Demetriou, M. Metabolism, cell surface organization, and disease. Cell 139, 1229–1241 (2009).
Croci, D.O. et al. Glycosylation-dependent lectin-receptor interactions preserve angiogenesis in anti-VEGF refractory tumors. Cell 156, 744–758 (2014).
Weinstein, I.B. & Joe, A. Oncogene addiction. Cancer Res. 68, 3077–3080, discussion 3080 (2008).
Kellenberger, C., Hendrickson, T.L. & Imperiali, B. Structural and functional analysis of peptidyl oligosaccharyl transferase inhibitors. Biochemistry 36, 12554–12559 (1997).
Weerapana, E. & Imperiali, B. Peptides to peptidomimetics: towards the design and synthesis of bioavailable inhibitors of oligosaccharyl transferase. Org. Biomol. Chem. 1, 93–99 (2003).
Bennett, D.C. et al. High-throughput screening identifies aclacinomycin as a radiosensitizer of EGFR-mutant non-small cell lung cancer. Transl. Oncol. 6, 382–391 (2013).
Baell, J.B. & Holloway, G.A. New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays. J. Med. Chem. 53, 2719–2740 (2010).
Auld, D.S. et al. Molecular basis for the high-affinity binding and stabilization of firefly luciferase by PTC124. Proc. Natl. Acad. Sci. USA 107, 4878–4883 (2010).
Maeda, Y., Tomita, S., Watanabe, R., Ohishi, K. & Kinoshita, T. DPM2 regulates biosynthesis of dolichol phosphate-mannose in mammalian cells: correct subcellular localization and stabilization of DPM1, and binding of dolichol phosphate. EMBO J. 17, 4920–4929 (1998).
Anand, M. et al. Requirement of the Lec35 gene for all known classes of monosaccharide-P-dolichol-dependent glycosyltransferase reactions in mammals. Mol. Biol. Cell 12, 487–501 (2001).
Mohorko, E. et al. Structural basis of substrate specificity of human oligosaccharyl transferase subunit N33/Tusc3 and its role in regulating protein N-glycosylation. Structure 22, 590–601 (2014).
Kelleher, D.J., Karaoglu, D. & Gilmore, R. Large-scale isolation of dolichol-linked oligosaccharides with homogeneous oligosaccharide structures: determination of steady-state dolichol-linked oligosaccharide compositions. Glycobiology 11, 321–333 (2001).
Shrimal, S., Trueman, S.F. & Gilmore, R. Extreme C-terminal sites are posttranslocationally glycosylated by the STT3B isoform of the OST. J. Cell Biol. 201, 81–95 (2013).
Jafari, R. et al. The cellular thermal shift assay for evaluating drug target interactions in cells. Nat. Protoc. 9, 2100–2122 (2014).
Paez, J.G. et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 304, 1497–1500 (2004).
Pao, W. et al. EGF receptor gene mutations are common in lung cancers from “never smokers” and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc. Natl. Acad. Sci. USA 101, 13306–13311 (2004).
Sordella, R., Bell, D.W., Haber, D.A. & Settleman, J. Gefitinib-sensitizing EGFR mutations in lung cancer activate anti-apoptotic pathways. Science 305, 1163–1167 (2004).
Weiss, J. et al. Frequent and focal FGFR1 amplification associates with therapeutically tractable FGFR1 dependency in squamous cell lung cancer. Sci. Transl. Med. 2, 62ra93 (2010).
Schneider, A. et al. Successful prenatal mannose treatment for congenital disorder of glycosylation-Ia in mice. Nat. Med. 18, 71–73 (2011).
Leung, E.L. et al. SRC promotes survival and invasion of lung cancers with epidermal growth factor receptor abnormalities and is a potential candidate for molecular-targeted therapy. Mol. Cancer Res. 7, 923–932 (2009).
Shimamura, T., Lowell, A.M., Engelman, J.A. & Shapiro, G.I. Epidermal growth factor receptors harboring kinase domain mutations associate with the heat shock protein 90 chaperone and are destabilized following exposure to geldanamycins. Cancer Res. 65, 6401–6408 (2005).
Kobayashi, S. et al. Transcriptional profiling identifies cyclin D1 as a critical downstream effector of mutant epidermal growth factor receptor signaling. Cancer Res. 66, 11389–11398 (2006).
Arrowsmith, C.H. et al. The promise and peril of chemical probes. Nat. Chem. Biol. 11, 536–541 (2015).
Gao, N. & Lehrman, M.A. Analyses of dolichol pyrophosphate-linked oligosaccharides in cell cultures and tissues by fluorophore-assisted carbohydrate electrophoresis. Glycobiology 12, 353–360 (2002).
Gao, N., Shang, J. & Lehrman, M.A. Analysis of glycosylation in CDG-Ia fibroblasts by fluorophore-assisted carbohydrate electrophoresis: implications for extracellular glucose and intracellular mannose 6-phosphate. J. Biol. Chem. 280, 17901–17909 (2005).
Acknowledgements
This work was funded by US National Institutes of Health (NIH) R03DA033178 and R01CA172391, and in part by a Research Scholar Grant from the American Cancer Society (J.N.C.) and by NIH RO1GM43768 (R.G.) and RO1GM038545 (M.A.L.). Additional funding was provided by the NIH-MLPCN program U54HG005031 (Kansas University) and U54HG005032 (Broad Institute). The mass spectrometry work was supported by NIH grant P41GM103490.
Author information
Authors and Affiliations
Contributions
C.L.-S. designed and performed the cell biology experiments in NSCLC cells and contributed to analysis and interpretation of all presented data. N.R. performed and analyzed the CTSA assays; J.C.C. performed experiments in CHO and Lec cells. J.N.C. designed the study and was involved in all experimental designs, data analysis, and data interpretation. S.S. and R.G. designed and performed in vitro glycosylation studies. C.K. and T.A.L. performed and triaged the HTS; D.P.F. and J.E.G. performed analysis of SAR and synthesis of chemical analogs; N.G. and M.A.L. performed the LLO analysis; P.Z. and L.W. collected and analyzed the MS data. All authors contributed to writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.N.C. and J.E.G. are listed as inventors on a provisional patent application for the analogs reported in this manuscript.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–9 and Supplementary Tables 1–6. (PDF 7267 kb)
Supplementary Note
Synthetic Procedures. (PDF 7757 kb)
Rights and permissions
About this article
Cite this article
Lopez-Sambrooks, C., Shrimal, S., Khodier, C. et al. Oligosaccharyltransferase inhibition induces senescence in RTK-driven tumor cells. Nat Chem Biol 12, 1023–1030 (2016). https://doi.org/10.1038/nchembio.2194
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2194
This article is cited by
-
Anti-PD-L1 antibodies promote cellular proliferation by activating the PD-L1–AXL signal relay in liver cancer cells
Hepatology International (2023)
-
5-AZA-dC induces epigenetic changes associated with modified glycosylation of secreted glycoproteins and increased EMT and migration in chemo-sensitive cancer cells
Clinical Epigenetics (2021)
-
STING enhances cell death through regulation of reactive oxygen species and DNA damage
Nature Communications (2021)
-
Receptor tyrosine kinases and cancer: oncogenic mechanisms and therapeutic approaches
Oncogene (2021)
-
Hexosamine pathway inhibition overcomes pancreatic cancer resistance to gemcitabine through unfolded protein response and EGFR-Akt pathway modulation
Oncogene (2020)