Abstract
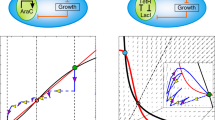
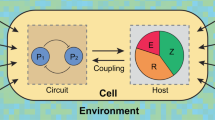
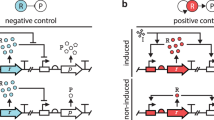
Synthetic gene circuits are often engineered by considering the host cell as an invariable 'chassis'. Circuit activation, however, may modulate host physiology, which in turn can substantially impact circuit behavior. We illustrate this point by a simple circuit consisting of mutant T7 RNA polymerase (T7 RNAP*) that activates its own expression in the bacterium Escherichia coli. Although activation by the T7 RNAP* is noncooperative, the circuit caused bistable gene expression. This counterintuitive observation can be explained by growth retardation caused by circuit activation, which resulted in nonlinear dilution of T7 RNAP* in individual bacteria. Predictions made by models accounting for such effects were verified by further experimental measurements. Our results reveal a new mechanism of generating bistability and underscore the need to account for host physiology modulation when engineering gene circuits.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Chin, J.W. Modular approaches to expanding the functions of living matter. Nat. Chem. Biol. 2, 304–311 (2006).
Marguet, P., Balagadde, F., Tan, C. & You, L. Biology by design: reduction and synthesis of cellular components and behaviour. J. R. Soc. Interface 4, 607–623 (2007).
Voigt, C.A. Genetic parts to program bacteria. Curr. Opin. Biotechnol. 17, 548–557 (2006).
Benner, S.A. & Sismour, A.M. Synthetic biology. Nat. Rev. Genet. 6, 533–543 (2005).
Canton, B., Labno, A. & Endy, D. Refinement and standardization of synthetic biological parts and devices. Nat. Biotechnol. 26, 787–793 (2008).
Andrianantoandro, E., Basu, S., Karig, D.K. & Weiss, R. Synthetic biology: new engineering rules for an emerging discipline. Mol. Syst. Biol. 2, 2006.0028 (2006).
Peretti, S.W. & Bailey, J.E. Simulations of host-plasmid interactions in Escherichia coli: copy number, promoter strength, and ribosome binding site strength effects on metabolic activity and plasmid gene expression. Biotechnol. Bioeng. 29, 316–328 (1987).
Becskei, A., Seraphin, B. & Serrano, L. Positive feedback in eukaryotic gene networks: cell differentiation by graded to binary response conversion. EMBO J. 20, 2528–2535 (2001).
Ajo-Franklin, C.M. et al. Rational design of memory in eukaryotic cells. Genes Dev. 21, 2271–2276 (2007).
Gardner, T.S., Cantor, C.R. & Collins, J.J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Basu, S., Mehreja, R., Thiberge, S., Chen, M.T. & Weiss, R. Spatiotemporal control of gene expression with pulse-generating networks. Proc. Natl. Acad. Sci. USA 101, 6355–6360 (2004).
Elowitz, M.B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Rosenfeld, N., Young, J.W., Alon, U., Swain, P.S. & Elowitz, M.B. Accurate prediction of gene feedback circuit behavior from component properties. Mol. Syst. Biol. 3, 143 (2007).
Balagadde, F.K. et al. A synthetic Escherichia coli predator-prey ecosystem. Mol. Syst. Biol. 4, 187 (2008).
Kramer, B.P. et al. An engineered epigenetic transgene switch in mammalian cells. Nat. Biotechnol. 22, 867–870 (2004).
Anderson, J.C., Voigt, C.A. & Arkin, A.P. Environmental signal integration by a modular AND gate. Mol. Syst. Biol. 3, 133 (2007).
Isaacs, F.J., Hasty, J., Cantor, C.R. & Collins, J.J. Prediction and measurement of an autoregulatory genetic module. Proc. Natl. Acad. Sci. USA 100, 7714–7719 (2003).
Lu, T., Volfson, D., Tsimring, L. & Hasty, J. Cellular growth and division in the Gillespie algorithm. Syst. Biol. (Stevenage) 1, 121–128 (2004).
St-Pierre, F. & Endy, D. Determination of cell fate selection during phage lambda infection. Proc. Natl. Acad. Sci. USA 105, 20705–20710 (2008).
You, L., Suthers, P.F. & Yin, J. Effects of Escherichia coli physiology on growth of phage T7 in vivo and in silico. J. Bacteriol. 184, 1888–1894 (2002).
Santillan, M., Mackey, M.C. & Zeron, E.S. Origin of bistability in the lac Operon. Biophys. J. 92, 3830–3842 (2007).
Dreisigmeyer, D.W., Stajic, J., Nemenman, I., Hlavacek, W.S. & Wall, M.E. Determinants of bistability in induction of the Escherichia coli lac operon. IET Syst. Biol. 2, 293–303 (2008).
Savageau, M.A. Design principles for elementary gene circuits: elements, methods, and examples. Chaos 11, 142–159 (2001).
Neves, S.R. et al. Cell shape and negative links in regulatory motifs together control spatial information flow in signaling networks. Cell 133, 666–680 (2008).
Tan, C., Song, H., Niemi, J. & You, L. A synthetic biology challenge: making cells compute. Mol. Biosyst. 3, 343–353 (2007).
Gesteland, R.F. & Atkins, J.F. Recoding: dynamic reprogramming of translation. Annu. Rev. Biochem. 65, 741–768 (1996).
Kramer, B.P. & Fussenegger, M. Hysteresis in a synthetic mammalian gene network. Proc. Natl. Acad. Sci. USA 102, 9517–9522 (2005).
Lim, H.N. & van Oudenaarden, A. A multistep epigenetic switch enables the stable inheritance of DNA methylation states. Nat. Genet. 39, 269–275 (2007).
Acar, M., Becskei, A. & van Oudenaarden, A. Enhancement of cellular memory by reducing stochastic transitions. Nature 435, 228–232 (2005).
Yao, G., Lee, T.J., Mori, S., Nevins, J.R. & You, L. A bistable Rb-E2F switch underlies the restriction point. Nat. Cell Biol. 10, 476–482 (2008).
Gordon, A.J. et al. Transcriptional infidelity promotes heritable phenotypic change in a bistable gene network. PLoS Biol. 7, e44 (2009).
Ferrell, J.E. Jr. Self-perpetuating states in signal transduction: positive feedback, double-negative feedback and bistability. Curr. Opin. Cell Biol. 14, 140–148 (2002).
Noireaux, V., Bar-Ziv, R. & Libchaber, A. Principles of cell-free genetic circuit assembly. Proc. Natl. Acad. Sci. USA 100, 12672–12677 (2003).
Martin, C.T. & Coleman, J.E. Kinetic analysis of T7 RNA polymerase-promoter interactions with small synthetic promoters. Biochemistry 26, 2690–2696 (1987).
Jia, Y., Kumar, A. & Patel, S.S. Equilibrium and stopped-flow kinetic studies of interaction between T7 RNA polymerase and its promoters measured by protein and 2-aminopurine fluorescence changes. J. Biol. Chem. 271, 30451–30458 (1996).
Davanloo, P., Rosenberg, A.H., Dunn, J.J. & Studier, F.W. Cloning and expression of the gene for bacteriophage T7 RNA polymerase. Proc. Natl. Acad. Sci. USA 81, 2035–2039 (1984).
Yin, Y.W. & Steitz, T.A. Structural basis for the transition from initiation to elongation transcription in T7 RNA polymerase. Science 298, 1387–1395 (2002).
Monod, J. The growth of bacterial cultures. Annu. Rev. Microbiol. 3, 371–394 (1949).
Dubendorff, J.W. & Studier, F.W. Creation of a T7 autogene. Cloning and expression of the gene for bacteriophage T7 RNA polymerase under control of its cognate promoter. J. Mol. Biol. 219, 61–68 (1991).
Elowitz, M.B., Levine, A.J., Siggia, E.D. & Swain, P.S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002).
Ozbudak, E.M., Thattai, M., Kurtser, I., Grossman, A.D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nat. Genet. 31, 69–73 (2002).
Dekel, E. & Alon, U. Optimality and evolutionary tuning of the expression level of a protein. Nature 436, 588–592 (2005).
Neidhardt, F.C. (ed.). Escherichia coli and Salmonella: Cellular and Molecular Biology (American Society Microbiology, Washington DC, 1996).
Klumpp, S. & Hwa, T. Growth-rate-dependent partitioning of RNA polymerases in bacteria. Proc. Natl. Acad. Sci. USA 105, 20245–20250 (2008).
Liang, S.T., Xu, Y.C., Dennis, P. & Bremer, H. mRNA composition and control of bacterial gene expression. J. Bacteriol. 182, 3037–3044 (2000).
Haseltine, E.L. & Arnold, F.H. Implications of rewiring bacterial quorum sensing. Appl. Environ. Microbiol. 74, 437–445 (2008).
Pruss, B.M., Markovic, D. & Matsumura, P. The Escherichia coli flagellar transcriptional activator flhD regulates cell division through induction of the acid response gene cadA. J. Bacteriol. 179, 3818–3821 (1997).
Hancock, V. & Klemm, P. Global gene expression profiling of asymptomatic bacteriuria Escherichia coli during biofilm growth in human urine. Infect. Immun. 75, 966–976 (2007).
Selinummi, J., Seppala, J., Yli-Harja, O. & Puhakka, J.A. Software for quantification of labeled bacteria from digital microscope images by automated image analysis. Biotechniques 39, 859–863 (2005).
Strogatz, S.H. Nonlinear Dynamics and Chaos: With Applications to Physics, Biology, Chemistry and Engineering (Perseus Books Group, New York, 2001).
Acknowledgements
We thank M. Salehi, G. Yao, J. Wong, H. Song, T.J. Lee, Q. Wang, J. Niemi, I. Molineux, M. Wall and W. Studier for discussions or comments; M. Cook for assistance with flow cytometry; W. Thompson, E. Soderblom and L. Dubois for assistance with mass spectrometry; M. Elowitz (California Institute of Technology), R. Weiss (Princeton University) and Y. Yokobayashi (University of California, Davis) for plasmids and bacterial strains; and T. Hwa for discussions and for sharing unpublished results. This work was partially supported by the US National Science Foundation (BES-0625213), the US National Institutes of Health (1P50GM081883), a DuPont Young Professorship (L.Y.), a David and Lucile Packard Fellowship (L.Y.) and a Medtronic Fellowship (C.T.).
Author information
Authors and Affiliations
Contributions
C.T. conceived research, designed and performed both modeling and experimental analyses, interpreted results and wrote the manuscript. P.M. purified and analyzed T7 RNAP* and assisted in manuscript revisions. L.Y. conceived research, assisted in research design and data interpretation and wrote the manuscript. All authors approved the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 1919 kb)
Supplementary Video 1
A time lapse movie corresponding to colony 1 in Fig. 1b, for 220 minutes of growth. (MOV 393 kb)
Supplementary Video 2
A time lapse movie of another colony, for 335 minutes of growth under the same condition as in Supplementary Video 1. (MOV 221 kb)
Rights and permissions
About this article
Cite this article
Tan, C., Marguet, P. & You, L. Emergent bistability by a growth-modulating positive feedback circuit. Nat Chem Biol 5, 842–848 (2009). https://doi.org/10.1038/nchembio.218
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.218
This article is cited by
-
Unbalanced response to growth variations reshapes the cell fate decision landscape
Nature Chemical Biology (2023)
-
Resource-aware construct design in mammalian cells
Nature Communications (2023)
-
A plasmid system with tunable copy number
Nature Communications (2022)
-
Drug-dependent growth curve reshaping reveals mechanisms of antifungal resistance in Saccharomyces cerevisiae
Communications Biology (2022)
-
A call for caution in analysing mammalian co-transfection experiments and implications of resource competition in data misinterpretation
Nature Communications (2021)