Abstract
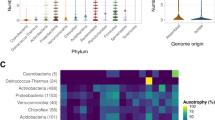
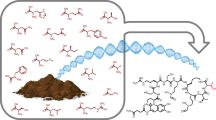
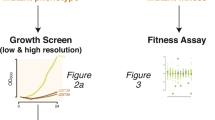
Only 25% of bacterial membrane transporters have functional annotation owing to the difficulty of experimental study and of accurate prediction of their function. Here we report a sequence-independent method for high-throughput mining of novel transporters. The method is based on ligand-responsive biosensor systems that enable selective growth of cells only if they encode a ligand-specific importer. We developed such a synthetic selection system for thiamine pyrophosphate and mined soil and gut metagenomes for thiamine-uptake functions. We identified several members of a novel class of thiamine transporters, PnuT, which is widely distributed across multiple bacterial phyla. We demonstrate that with modular replacement of the biosensor, we could expand our method to xanthine and identify xanthine permeases from gut and soil metagenomes. Our results demonstrate how synthetic-biology approaches can effectively be deployed to functionally mine metagenomes and elucidate sequence–function relationships of small-molecule transport systems in bacteria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kell, D.B., Swainston, N., Pir, P. & Oliver, S.G. Membrane transporter engineering in industrial biotechnology and whole cell biocatalysis. Trends Biotechnol. 33, 237–246 (2015).
Walsh, C. Molecular mechanisms that confer antibacterial drug resistance. Nature 406, 775–781 (2000).
Herbert, M. et al. Nicotinamide ribosyl uptake mutants in Haemophilus influenzae. Infect. Immun. 71, 5398–5401 (2003).
Degnan, P.H., Barry, N.A., Mok, K.C., Taga, M.E. & Goodman, A.L. Human gut microbes use multiple transporters to distinguish vitamin B12 analogs and compete in the gut. Cell Host Microbe 15, 47–57 (2014).
Arai, M., Okumura, K., Satake, M. & Shimizu, T. Proteome-wide functional classification and identification of prokaryotic transmembrane proteins by transmembrane topology similarity comparison. Protein Sci. 13, 2170–2183 (2004).
Ren, Q., Chen, K. & Paulsen, I.T. TransportDB: a comprehensive database resource for cytoplasmic membrane transport systems and outer membrane channels. Nucleic Acids Res. 35, D274–D279 (2007).
Ren, Q. & Paulsen, I.T. Large-scale comparative genomic analyses of cytoplasmic membrane transport systems in prokaryotes. J. Mol. Microbiol. Biotechnol. 12, 165–179 (2007).
Michener, J.K. & Smolke, C.D. High-throughput enzyme evolution in Saccharomyces cerevisiae using a synthetic RNA switch. Metab. Eng. 14, 306–316 (2012).
van Sint Fiet, S., van Beilen, J.B. & Witholt, B. Selection of biocatalysts for chemical synthesis. Proc. Natl. Acad. Sci. USA 103, 1693–1698 (2006).
Yang, J. et al. Synthetic RNA devices to expedite the evolution of metabolite-producing microbes. Nat. Commun. 4, 1413 (2013).
Raman, S., Rogers, J.K., Taylor, N.D. & Church, G.M. Evolution-guided optimization of biosynthetic pathways. Proc. Natl. Acad. Sci. USA 111, 17803–17808 (2014).
Binder, S. et al. A high-throughput approach to identify genomic variants of bacterial metabolite producers at the single-cell level. Genome Biol. 13, R40 (2012).
Siedler, S. et al. SoxR as a single-cell biosensor for NADPH-consuming enzymes in Escherichia coli. ACS Synth. Biol. 3, 41–47 (2014).
Novichkov, P.S. et al. RegPrecise 3.0—a resource for genome-scale exploration of transcriptional regulation in bacteria. BMC Genomics 14, 745 (2013).
Wittmann, A. & Suess, B. Engineered riboswitches: Expanding researchers' toolbox with synthetic RNA regulators. FEBS Lett. 586, 2076–2083 (2012).
Muranaka, N., Sharma, V., Nomura, Y. & Yokobayashi, Y. An efficient platform for genetic selection and screening of gene switches in Escherichia coli. Nucleic Acids Res. 37, e39 (2009).
Winkler, W., Nahvi, A. & Breaker, R.R. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature 419, 952–956 (2002).
Rodionov, D.A. et al. A novel class of modular transporters for vitamins in prokaryotes. J. Bacteriol. 191, 42–51 (2009).
Webb, E., Claas, K. & Downs, D. thiBPQ encodes an ABC transporter required for transport of thiamine and thiamine pyrophosphate in Salmonella typhimurium. J. Biol. Chem. 273, 8946–8950 (1998).
Jeanguenin, L. et al. Comparative genomics and functional analysis of the NiaP family uncover nicotinate transporters from bacteria, plants, and mammals. Funct. Integr. Genomics 12, 25–34 (2012).
Rodionova, I.A. et al. Genomic distribution of B-vitamin auxotrophy and uptake transporters in environmental bacteria from the Chloroflexi phylum. Environ. Microbiol. Rep. 7, 204–210 (2015).
LeBlanc, J.G. et al. Bacteria as vitamin suppliers to their host: a gut microbiota perspective. Curr. Opin. Biotechnol. 24, 160–168 (2013).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Aziz, R.K. et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9, 75 (2008).
Sommer, M.O., Dantas, G. & Church, G.M. Functional characterization of the antibiotic resistance reservoir in the human microflora. Science 325, 1128–1131 (2009).
Forsberg, K.J. et al. The shared antibiotic resistome of soil bacteria and human pathogens. Science 337, 1107–1111 (2012).
Sauer, E., Merdanovic, M., Mortimer, A.P., Bringmann, G. & Reidl, J. PnuC and the utilization of the nicotinamide riboside analog 3-aminopyridine in Haemophilus influenzae. Antimicrob. Agents Chemother. 48, 4532–4541 (2004).
Jaehme, M., Guskov, A. & Slotboom, D.J. Crystal structure of the vitamin B3 transporter PnuC, a full-length SWEET homolog. Nat. Struct. Mol. Biol. 21, 1013–1015 (2014).
Grose, J.H. et al. Assimilation of nicotinamide mononucleotide requires periplasmic AphA phosphatase in Salmonella enterica. J. Bacteriol. 187, 4521–4530 (2005).
Vogl, C. et al. Characterization of riboflavin (vitamin B 2) transport proteins from Bacillus subtilis and Corynebacterium glutamicum. J. Bacteriol. 189, 7367–7375 (2007).
Hemberger, S. et al. RibM from Streptomyces davawensis is a riboflavin/roseoflavin transporter and may be useful for the optimization of riboflavin production strains. BMC Biotechnol. 11, 119 (2011).
Rodionov, D.A., Vitreschak, A.G., Mironov, A.A. & Gelfand, M.S. Comparative genomics of thiamin biosynthesis in procaryotes. New genes and regulatory mechanisms. J. Biol. Chem. 277, 48949–48959 (2002).
Gelfand, M.S. & Rodionov, D.A. Comparative genomics and functional annotation of bacterial transporters. Phys. Life Rev. 5, 22–49 (2008).
Jaehme, M. & Slotboom, D.J. Structure, function, evolution, and application of bacterial Pnu-type vitamin transporters. Biol. Chem. 396, 955–966 (2015).
Abreu-Goodger, C. & Merino, E. RibEx: a web server for locating riboswitches and other conserved bacterial regulatory elements. Nucleic Acids Res. 33, W690–W692 (2005).
Schauer, K., Rodionov, D.A. & de Reuse, H. New substrates for TonB-dependent transport: do we only see the 'tip of the iceberg'? Trends Biochem. Sci. 33, 330–338 (2008).
Methé, B.A. et al. A framework for human microbiome research. Nature 486, 215–221 (2012).
Dehal, P.S. et al. MicrobesOnline: an integrated portal for comparative and functional genomics. Nucleic Acids Res. 38, D396–D400 (2010).
Nedenskov, P. Nutritional requirements for growth of Helicobacter pylori. Appl. Environ. Microbiol. 60, 3450–3453 (1994).
Lynch, S.A. & Gallivan, J.P. A flow cytometry-based screen for synthetic riboswitches. Nucleic Acids Res. 37, 184–192 (2009).
Jenison, R., Gill, S., Pardi, A. & Polisky, B. High-resolution molecular discrimination by RNA. Science 263, 1425–1429 (1994).
Karatza, P. & Frillingos, S. Cloning and functional characterization of two bacterial members of the NAT/NCS2 family in Escherichia coli. Mol. Membr. Biol. 22, 251–261 (2005).
Mahr, R. & Frunzke, J. Transcription factor-based biosensors in biotechnology: current state and future prospects. Appl. Microbiol. Biotechnol. 100, 79–90 (2016).
Taylor, N.D. et al. Engineering an allosteric transcription factor to respond to new ligands. Nat. Methods 13, 177–183 (2016).
Goler, J.A., Carothers, J.M. & Keasling, J.D. Dual-selection for evolution of in vivo functional aptazymes as riboswitch parts. Methods Mol. Biol. 1111, 221–235 (2014).
Libis, V., Delépine, B. & Faulon, J.-L. Expanding biosensing abilities through computer-aided design of metabolic pathways. ACS Synth. Biol. http://dx.doi.org/10.1021/acssynbio.5b00225 (2016).
Wagner, S. et al. Consequences of membrane protein overexpression in Escherichia coli. Mol. Cell. Proteomics 6, 1527–1550 (2007).
Genee, H.J. et al. Software-supported USER cloning strategies for site-directed mutagenesis and DNA assembly. ACS Synth. Biol. 4, 342–349 (2015).
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997).
Chaudhuri, R.R. et al. Complete genome sequence and comparative metabolic profiling of the prototypical enteroaggregative Escherichia coli strain 042. PLoS One 5, e8801 (2010).
Schyns, G. et al. Isolation and characterization of new thiamine-deregulated mutants of Bacillus subtilis 187, 8127–8136 (2005).
Bontemps, J. et al. Determination of thiamine and thiamine phosphates in excitable tissues as thiochrome derivatives by reversed-phase high-performance liquid chromatography on octadecyl silica. J. Chromatogr. 307, 283–294 (1984).
Volkmer, B. & Heinemann, M. Condition-dependent cell volume and concentration of Escherichia coli to facilitate data conversion for systems biology modeling. PLoS One 6, e23126 (2011).
van der Helm, E., Geertz-Hansen, H.M., Genee, H.J., Malla, S. & Sommer, M.O.A. deFUME: dynamic exploration of functional metagenomic sequencing data. BMC Res. Notes 8, 328 (2015).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Jaehme, M. & Slotboom, D.J. Diversity of membrane transport proteins for vitamins in bacteria and archaea. Biochim. Biophys. Acta 1850, 565–576 (2015).
Gibson, D.G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Hillson, N.J., Rosengarten, R.D. & Keasling, J.D. j5 DNA assembly design automation software. ACS Synth. Biol. 1, 14–21 (2012).
Acknowledgements
We thank Y. Yokobayashi (University of California, Davis, California, USA) for providing plasmid pLacthiM19tetA-gfpuv, J. Gallivan (Emory University, Atlanta, Georgia, USA) for providing plasmid pSKD314, D. Paiva (Technical University of Denmark, Kongens Lyngby, Denmark) for metagenomic libraries, R. Lavallee for technical HPLC support, and C. Munck for critical reading of the manuscript. This study was funded by the Novo Nordisk Foundation and the European Union Seventh Framework Programme (FP7-KBBE-2013-7 single stage) under grant agreement no. 613745, Promys. H.J.G. acknowledges additional financial support from Novozymes A/S.
Author information
Authors and Affiliations
Contributions
H.J.G. and M.O.A.S. conceived the study. H.J.G. and S.D.P. developed the thiamine functional selection system, and H.J.G. and A.P.B. performed functional metagenomic selections for thiamine uptake. H.J.G., M.K. and L.S.G. developed the HPLC assay for measurement of thiamines. H.J.G. and A.P.B. cloned heterologous genes. H.J.G. and M.T.B. developed the xanthine alkaloid selection system, and H.J.G. performed functional metagenomic selections and analysis. H.J.G. and S.S. performed dose-response characterizations of selected xanthine importers. S.J.H. developed the LC-MS method for xanthine alkaloids and performed measurements of samples prepared by S.S. and H.J.G. H.J.G. wrote the manuscript with contributions from all other authors.
Corresponding author
Ethics declarations
Competing interests
H.J.G. and M.O.A.S. are named on a pending patent application relating to dual genetic selection systems (WO 2014/187829 A1).
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–4 and Supplementary Tables 1–10. (PDF 1926 kb)
Supplementary Table 11
Comparative genomics of thiamine biosynthesis, salvage and transport of 1752 complete bacterial genomes. (XLSX 166 kb)
Rights and permissions
About this article
Cite this article
Genee, H., Bali, A., Petersen, S. et al. Functional mining of transporters using synthetic selections. Nat Chem Biol 12, 1015–1022 (2016). https://doi.org/10.1038/nchembio.2189
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2189
This article is cited by
-
A previously unidentified sugar transporter for engineering of high-yield Streptomyces
Applied Microbiology and Biotechnology (2024)
-
The Y-ome Conundrum: Insights into Uncharacterized Genes and Approaches for Functional Annotation
Molecular and Cellular Biochemistry (2023)
-
Synthetic biology for sustainable food ingredients production: recent trends
Systems Microbiology and Biomanufacturing (2023)
-
Identification Process and Physiological Properties of Transporters of Carboxylic Acids in Escherichia coli
Biotechnology and Bioprocess Engineering (2022)
-
Genetically encoded biosensors for lignocellulose valorization
Biotechnology for Biofuels (2019)