Abstract
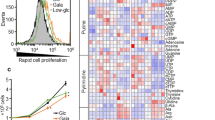
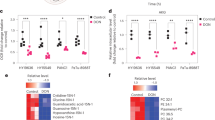
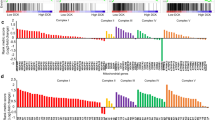
Metabolic reprogramming in tumors represents a potential therapeutic target. Herein we used shRNA depletion and a novel lactate dehydrogenase (LDHA) inhibitor, GNE-140, to probe the role of LDHA in tumor growth in vitro and in vivo. In MIA PaCa-2 human pancreatic cells, LDHA inhibition rapidly affected global metabolism, although cell death only occurred after 2 d of continuous LDHA inhibition. Pancreatic cell lines that utilize oxidative phosphorylation (OXPHOS) rather than glycolysis were inherently resistant to GNE-140, but could be resensitized to GNE-140 with the OXPHOS inhibitor phenformin. Acquired resistance to GNE-140 was driven by activation of the AMPK–mTOR–S6K signaling pathway, which led to increased OXPHOS, and inhibitors targeting this pathway could prevent resistance. Thus, combining an LDHA inhibitor with compounds targeting the mitochondrial or AMPK–S6K signaling axis may not only broaden the clinical utility of LDHA inhibitors beyond glycolytically dependent tumors but also reduce the emergence of resistance to LDHA inhibition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Brand, K.A. & Hermfisse, U. Aerobic glycolysis by proliferating cells: a protective strategy against reactive oxygen species. FASEB J. 11, 388–395 (1997).
Koukourakis, M.I. et al. Lactate dehydrogenase 5 isoenzyme overexpression defines resistance of prostate cancer to radiotherapy. Br. J. Cancer 110, 2217–2223 (2014).
Koukourakis, M.I., Giatromanolaki, A. & Sivridis, E. Lactate dehydrogenase isoenzymes 1 and 5: differential expression by neoplastic and stromal cells in non-small cell lung cancer and other epithelial malignant tumors. Tumour Biol. 24, 199–202 (2003).
Koukourakis, M.I., Giatromanolaki, A., Sivridis, E., Gatter, K.C. & Harris, A.L. Tumour Angiogenesis Research Group. Lactate dehydrogenase 5 expression in operable colorectal cancer: strong association with survival and activated vascular endothelial growth factor pathway—a report of the Tumour Angiogenesis Research Group. J. Clin. Oncol. 24, 4301–4308 (2006).
Girgis, H. et al. Lactate dehydrogenase A is a potential prognostic marker in clear cell renal cell carcinoma. Mol. Cancer 13, 101 (2014).
Shim, H. et al. c-Myc transactivation of LDH-A: implications for tumor metabolism and growth. Proc. Natl. Acad. Sci. USA 94, 6658–6663 (1997).
Semenza, G.L. et al. Hypoxia response elements in the aldolase A, enolase 1, and lactate dehydrogenase A gene promoters contain essential binding sites for hypoxia-inducible factor 1. J. Biol. Chem. 271, 32529–32537 (1996).
Fantin, V.R., St-Pierre, J. & Leder, P. Attenuation of LDH-A expression uncovers a link between glycolysis, mitochondrial physiology, and tumor maintenance. Cancer Cell 9, 425–434 (2006).
Qing, G. et al. Combinatorial regulation of neuroblastoma tumor progression by N-Myc and hypoxia inducible factor HIF-1alpha. Cancer Res. 70, 10351–10361 (2010).
Seth, P. et al. On-target inhibition of tumor fermentative glycolysis as visualized by hyperpolarized pyruvate. Neoplasia 13, 60–71 (2011).
Xie, H. et al. Targeting lactate dehydrogenase-A inhibits tumorigenesis and tumor progression in mouse models of lung cancer and impacts tumor-initiating cells. Cell Metab. 19, 795–809 (2014).
Novoa, W.B., Winer, A.D., Glaid, A.J. & Schwert, G.W. Lactic dehydrogenase. V. Inhibition by oxamate and by oxalate. J. Biol. Chem. 234, 1143–1148 (1959).
Busch, H. & Nair, P.V. Inhibition of lactic acid dehydrogenase by fluoropyruvic acid. J. Biol. Chem. 229, 377–387 (1957).
Billiard, J. et al. Quinoline 3-sulfonamides inhibit lactate dehydrogenase A and reverse aerobic glycolysis in cancer cells. Cancer Metab. 1, 19 (2013).
Dragovich, P.S. et al. Identification of substituted 2-thio-6-oxo-1,6-dihydropyrimidines as inhibitors of human lactate dehydrogenase. Bioorg. Med. Chem. Lett. 23, 3186–3194 (2013).
Fauber, B.P. et al. Identification of 2-amino-5-aryl-pyrazines as inhibitors of human lactate dehydrogenase. Bioorg. Med. Chem. Lett. 23, 5533–5539 (2013).
Granchi, C. et al. N-Hydroxyindole-based inhibitors of lactate dehydrogenase against cancer cell proliferation. Eur. J. Med. Chem. 46, 5398–5407 (2011).
Le, A. et al. Inhibition of lactate dehydrogenase A induces oxidative stress and inhibits tumor progression. Proc. Natl. Acad. Sci. USA 107, 2037–2042 (2010).
Ward, R.A. et al. Design and synthesis of novel lactate dehydrogenase A inhibitors by fragment-based lead generation. J. Med. Chem. 55, 3285–3306 (2012).
Vanderporten, E. et al. Label-free high-throughput assays to screen and characterize novel lactate dehydrogenase inhibitors. Anal. Biochem. 441, 115–122 (2013).
Dragovich, P.S. et al. Identification of substituted 3-hydroxy-2-mercaptocyclohex-2-enones as potent inhibitors of human lactate dehydrogenase. Bioorg. Med. Chem. Lett. 24, 3764–3771 (2014).
Fauber, B.P. et al. Identification of 3,6-disubstituted dihydropyrones as inhibitors of human lactate dehydrogenase. Bioorg. Med. Chem. Lett. 24, 5683–5687 (2014).
Aithal, H.N., Walsh-Reitz, M.M. & Toback, F.G. Regulation of glyceraldehyde-3-phosphate dehydrogenase by a cytosolic protein. Am. J. Physiol. 249, C111–C116 (1985).
Copeland, L. & Zammit, A. Kinetic properties of NAD-dependent glyceraldehyde-3-phosphate dehydrogenase from the host fraction of soybean root nodules. Arch. Biochem. Biophys. 312, 107–113 (1994).
Owen, M.R., Doran, E. & Halestrap, A.P. Evidence that metformin exerts its anti-diabetic effects through inhibition of complex 1 of the mitochondrial respiratory chain. Biochem. J. 348, 607–614 (2000).
Shitara, Y. et al. Role of organic cation/carnitine transporter 1 in uptake of phenformin and inhibitory effect on complex I respiration in mitochondria. Toxicol. Sci. 132, 32–42 (2013).
Borisy, A.A. et al. Systematic discovery of multicomponent therapeutics. Proc. Natl. Acad. Sci. USA 100, 7977–7982 (2003).
Hardie, D.G. AMPK: positive and negative regulation, and its role in whole-body energy homeostasis. Curr. Opin. Cell Biol. 33, 1–7 (2015).
Shirwany, N.A. & Zou, M.H. AMPK: a cellular metabolic and redox sensor. A minireview. Front. Biosci. (Landmark Ed.) 19, 447–474 (2014).
Inoki, K., Zhu, T. & Guan, K.L. TSC2 mediates cellular energy response to control cell growth and survival. Cell 115, 577–590 (2003).
Gwinn, D.M. et al. AMPK phosphorylation of raptor mediates a metabolic checkpoint. Mol. Cell 30, 214–226 (2008).
Pearce, L.R. et al. Characterization of PF-4708671, a novel and highly specific inhibitor of p70 ribosomal S6 kinase (S6K1). Biochem. J. 431, 245–255 (2010).
Ma, X.M. & Blenis, J. Molecular mechanisms of mTOR-mediated translational control. Nat. Rev. Mol. Cell Biol. 10, 307–318 (2009).
Magnuson, B., Ekim, B. & Fingar, D.C. Regulation and function of ribosomal protein S6 kinase (S6K) within mTOR signalling networks. Biochem. J. 441, 1–21 (2012).
Feng, Z. & Levine, A.J. The regulation of energy metabolism and the IGF-1/mTOR pathways by the p53 protein. Trends Cell Biol. 20, 427–434 (2010).
Viale, A. et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 514, 628–632 (2014).
Birsoy, K. et al. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell 162, 540–551 (2015).
Chen, J. et al. Piperidine-dione derivatives. World Intellectual Property Organization patent WO 2015/140133 A1 (2015).
Reitman, Z.J. et al. Profiling the effects of isocitrate dehydrogenase 1 and 2 mutations on the cellular metabolome. Proc. Natl. Acad. Sci. USA 108, 3270–3275 (2011).
Gray, D.C. et al. pHUSH: a single vector system for conditional gene expression. BMC Biotechnol. 7, 61 (2007).
Acknowledgements
We would like to thank the Discovery Chemistry Small Molecule analytical group for their support; the in vivo cell culture and dosing groups for their support of in vivo studies; R. Bourgon for bioinformatic assistance; the Genentech gCell group (R. Neve, M. Yu and M. Liang-Chu) for cell line banking and maintenance; and members of Genentech gCSI group (E. Lin, B. Lam, Y. Yu and J. Tan) for assistance on cell-based drug screens; S. Sriraman for assistance with the 18FDG-uptake assays; and A. Bruce for assistance with generating figures.
Author information
Authors and Affiliations
Contributions
A.B. contributed in the generation of, and characterized, resistant cell lines and to manuscript writing; H.E.P. and K.R. designed and directed compound generation and characterization of lead molecules and assisted in generating co-crystal structure; A.H. contributed through the generation and characterization shLDHA cell lines and by performing metabolic profiling experiments; D.P. characterized the induction of ROS, performed cell line assays and characterized lactate levels in cell lines; S.L. designed and assisted in compound generation and characterization of lead molecules; M.K. and M.G. performed all MS-based assays; R.H. designed and carried out in vivo studies; C.D.N. performed cell cycle analysis of cells; R.P. assisted in understanding the mechanism of resistance; M.M. and R.F. assisted with knockdown construct design and characterization; S. Ma, L.S. and J.P. designed and carried out pharmacokinetic studies in mice; A.Z. and T.L. designed and assisted in compound generation and characterization of lead molecules; Y.L. and Z.C. obtained a co-crystal structure; B.W. designed molecules and assisted in obtaining a co-crystal structure; I.Y., S.S. and S. Malek designed and performed biochemical and selectivity assays; L.C. designed and generated shLDHA cells; A.V. and S.W. performed cellular glucose uptake assays; A.D. assisted in analysis of metabolite profiling in cells; M.B. performed characterization of lead molecule in cells; C.E. directed protein production and generation of a co-crystal structure; P.K.J. directed the design and characterization of shLDHA cells; G.H. contributed intellectually to the design of all metabolic assays and helped draft the manuscript; D.S. directed in vivo studies and helped draft the manuscript; M.E. directed the generation and characterization of resistant clones, as well as cell line screening and in vivo studies; T.O'B. directed the characterization of the role of LDHA in cells and helped design and direct metabolic and cell cycle and in vivo studies; M.E. and T.O'B. drafted and revised the manuscript for intellectual content and final approval.
Corresponding authors
Ethics declarations
Competing interests
All authors were full-time employees of either Genentech or WuXi at the time these studies were conducted.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–3 and Supplementary Figures 1–9. (PDF 15669 kb)
Supplementary Dataset 1
Complete list of the fold change of all metabolites detected in MIA PaCa-2 cells following treatment with 2 μM GNE-140. (XLSX 61 kb)
Rights and permissions
About this article
Cite this article
Boudreau, A., Purkey, H., Hitz, A. et al. Metabolic plasticity underpins innate and acquired resistance to LDHA inhibition. Nat Chem Biol 12, 779–786 (2016). https://doi.org/10.1038/nchembio.2143
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2143
This article is cited by
-
Transcriptional regulation and post-translational modifications in the glycolytic pathway for targeted cancer therapy
Acta Pharmacologica Sinica (2024)
-
The potential role of reprogrammed glucose metabolism: an emerging actionable codependent target in thyroid cancer
Journal of Translational Medicine (2023)
-
The mitochondrial NADH shuttle system is a targetable vulnerability for Group 3 medulloblastoma in a hypoxic microenvironment
Cell Death & Disease (2023)
-
Lactate-dependent transcriptional regulation controls mammalian eye morphogenesis
Nature Communications (2023)
-
Tumor lactic acid: a potential target for cancer therapy
Archives of Pharmacal Research (2023)