Abstract
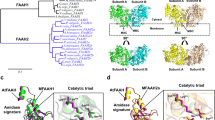
Enzyme classes may contain outlier members that share mechanistic, but not sequence or structural, relatedness with more common representatives. The functional annotation of such exceptional proteins can be challenging. Here, we use activity-based profiling to discover that the poorly characterized multipass transmembrane proteins AIG1 and ADTRP are atypical hydrolytic enzymes that depend on conserved threonine and histidine residues for catalysis. Both AIG1 and ADTRP hydrolyze bioactive fatty acid esters of hydroxy fatty acids (FAHFAs) but not other major classes of lipids. We identify multiple cell-active, covalent inhibitors of AIG1 and show that these agents block FAHFA hydrolysis in mammalian cells. These results indicate that AIG1 and ADTRP are founding members of an evolutionarily conserved class of transmembrane threonine hydrolases involved in bioactive lipid metabolism. More generally, our findings demonstrate how chemical proteomics can excavate potential cases of convergent or parallel protein evolution that defy conventional sequence- and structure-based predictions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Willems, L.I., Overkleeft, H.S. & van Kasteren, S.I. Current developments in activity-based protein profiling. Bioconjug. Chem. 25, 1181–1191 (2014).
Niphakis, M.J. & Cravatt, B.F. Enzyme inhibitor discovery by activity-based protein profiling. Annu. Rev. Biochem. 83, 341–377 (2014).
Berger, A.B., Vitorino, P.M. & Bogyo, M. Activity-based protein profiling: applications to biomarker discovery, in vivo imaging and drug discovery. Am. J. Pharmacogenomics 4, 371–381 (2004).
Liu, Y., Patricelli, M.P. & Cravatt, B.F. Activity-based protein profiling: the serine hydrolases. Proc. Natl. Acad. Sci. USA 96, 14694–14699 (1999).
Simon, G.M. & Cravatt, B.F. Activity-based proteomics of enzyme superfamilies: serine hydrolases as a case study. J. Biol. Chem. 285, 11051–11055 (2010).
Bachovchin, D.A. et al. Superfamily-wide portrait of serine hydrolase inhibition achieved by library-versus-library screening. Proc. Natl. Acad. Sci. USA 107, 20941–20946 (2010).
Jessani, N. et al. A streamlined platform for high-content functional proteomics of primary human specimens. Nat. Methods 2, 691–697 (2005).
Higa, H.H., Diaz, S. & Varki, A. Biochemical and genetic evidence for distinct membrane-bound and cytosolic sialic acid O-acetyl-esterases: serine-active-site enzymes. Biochem. Biophys. Res. Commun. 144, 1099–1108 (1987).
Jessani, N. et al. Class assignment of sequence-unrelated members of enzyme superfamilies by activity-based protein profiling. Angew. Chem. Int. Edn. Engl. 44, 2400–2403 (2005).
Long, J.Z. & Cravatt, B.F. The metabolic serine hydrolases and their functions in mammalian physiology and disease. Chem. Rev. 111, 6022–6063 (2011).
Elias, M. & Tawfik, D.S. Divergence and convergence in enzyme evolution: parallel evolution of paraoxonases from quorum-quenching lactonases. J. Biol. Chem. 287, 11–20 (2012).
Lone, A.M. et al. A substrate-free activity-based protein profiling screen for the discovery of selective PREPL inhibitors. J. Am. Chem. Soc. 133, 11665–11674 (2011).
Kidd, D., Liu, Y. & Cravatt, B.F. Profiling serine hydrolase activities in complex proteomes. Biochemistry 40, 4005–4015 (2001).
Adibekian, A. et al. Click-generated triazole ureas as ultrapotent in vivo–active serine hydrolase inhibitors. Nat. Chem. Biol. 7, 469–478 (2011).
Seo, J., Kim, J. & Kim, M. Cloning of androgen-inducible gene 1 (AIG1) from human dermal papilla cells. Mol. Cells 11, 35–40 (2001).
Wu, G., Sun, M., Zhang, W. & Huo, K. AIG1 is a novel Pirh2-interacting protein that activates the NFAT signaling pathway. Front. Biosci. 3, 834–842 (2011).
Lupu, C., Zhu, H., Popescu, N.I., Wren, J.D. & Lupu, F. Novel protein ADTRP regulates TFPI expression and function in human endothelial cells in normal conditions and in response to androgen. Blood 118, 4463–4471 (2011).
Weerapana, E. et al. Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature 468, 790–795 (2010).
Dodson, G. & Wlodawer, A. Catalytic triads and their relatives. Trends Biochem. Sci. 23, 347–352 (1998).
Ekici, O.D., Paetzel, M. & Dalbey, R.E. Unconventional serine proteases: variations on the catalytic Ser/His/Asp triad configuration. Protein Sci. 17, 2023–2037 (2008).
Yore, M.M. et al. Discovery of a class of endogenous mammalian lipids with anti-diabetic and anti-inflammatory effects. Cell 159, 318–332 (2014).
Cognetta, A.B. III et al. Selective N-hydroxyhydantoin carbamate inhibitors of mammalian serine hydrolases. Chem. Biol. 22, 928–937 (2015).
Kamat, S.S. et al. Immunomodulatory lysophosphatidylserines are regulated by ABHD16A and ABHD12 interplay. Nat. Chem. Biol. 11, 164–171 (2015).
Hoover, H.S., Blankman, J.L., Niessen, S. & Cravatt, B.F. Selectivity of inhibitors of endocannabinoid biosynthesis evaluated by activity-based protein profiling. Bioorg. Med. Chem. Lett. 18, 5838–5841 (2008).
Nomura, D.K. et al. Monoacylglycerol lipase exerts dual control over endocannabinoid and fatty acid pathways to support prostate cancer. Chem. Biol. 18, 846–856 (2011).
Brown, M.S., Ye, J., Rawson, R.B. & Goldstein, J.L. Regulated intramembrane proteolysis: a control mechanism conserved from bacteria to humans. Cell 100, 391–398 (2000).
Wolfe, M.S. Intramembrane-cleaving proteases. J. Biol. Chem. 284, 13969–13973 (2009).
Urban, S., Lee, J.R. & Freeman, M. Drosophila rhomboid-1 defines a family of putative intramembrane serine proteases. Cell 107, 173–182 (2001).
Strisovsky, K. Why cells need intramembrane proteases—a mechanistic perspective. FEBS J. 10.1111/febs.13638 (2015).
Sherratt, A.R., Blais, D.R., Ghasriani, H., Pezacki, J.P. & Goto, N.K. Activity-based protein profiling of the Escherichia coli GlpG rhomboid protein delineates the catalytic core. Biochemistry 51, 7794–7803 (2012).
Vosyka, O. et al. Activity-based probes for rhomboid proteases discovered in a mass spectrometry-based assay. Proc. Natl. Acad. Sci. USA 110, 2472–2477 (2013).
Nguyen, M.T., Van Kersavond, T. & Verhelst, S.H. Chemical tools for the study of intramembrane proteases. ACS Chem. Biol. 10, 2423–2434 (2015).
Rath, A., Glibowicka, M., Nadeau, V.G., Chen, G. & Deber, C.M. Detergent binding explains anomalous SDS-PAGE migration of membrane proteins. Proc. Natl. Acad. Sci. USA 106, 1760–1765 (2009).
Gerlt, J.A. et al. The enzyme function initiative. Biochemistry 50, 9950–9962 (2011).
Adibekian, A. et al. Confirming target engagement for reversible inhibitors in vivo by kinetically tuned activity-based probes. J. Am. Chem. Soc. 134, 10345–10348 (2012).
Söding, J., Biegert, A. & Lupas, A.N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005).
Patricelli, M.P., Giang, D.K., Stamp, L.M. & Burbaum, J.J. Direct visualization of serine hydrolase activities in complex proteomes using fluorescent active site-directed probes. Proteomics 1, 1067–1071 (2001).
Tully, S.E. & Cravatt, B.F. Activity-based probes that target functional subclasses of phospholipases in proteomes. J. Am. Chem. Soc. 132, 3264–3265 (2010).
Hsu, K.L. et al. DAGLβ inhibition perturbs a lipid network involved in macrophage inflammatory responses. Nat. Chem. Biol. 8, 999–1007 (2012).
Washburn, M.P., Wolters, D. & Yates, J.R. III. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247 (2001).
Acknowledgements
We are grateful to K. Masuda and L. Bar-Peled for discussions on and assistance with cloning; M. Dix, J. Wu and K. Lum for discussions and technical expertise in designing and analyzing proteomics experiments; J. Teijaro for providing T cells; G. Simon for assistance with the HHpred analysis; and C. Walsh and M. Niphakis for numerous helpful discussions. This work was supported by the US National Institutes of Health (DA033760, DK909810), The Leona M. and Harry B. Helmsley Charitable Trust (grant no. 2012-PG-MED002 to A.S.), National Cancer Institute Cancer Center Support grant P30 (CA014195 MASS core, A.S.), Dr. Frederick Paulsen Chair/Ferring Pharmaceuticals (A.S.), a Hewitt Foundation for Medical Research Fellowship (to W.H.P.), a Chapman Charitable Trust Fellowship (to M.J.K.), UCSD Medical Scientist Training Program funding (T32 GM007198 to M.J.K.) and an Irving S. Sigal postdoctoral fellowship from the American Chemical Society (to S.S.K.).
Author information
Authors and Affiliations
Contributions
W.H.P., M.J.K., A.S. and B.F.C. conceived the project. W.H.P., M.J.K., S.S.K., A.S. and B.F.C. designed experiments. W.H.P. performed the molecular biology and proteomics experiments. W.H.P., A.B.C., J.J.H. and A.S. synthesized compounds. M.J.K., S.S.K. and W.H.P. performed substrate assays and biochemical experiments. W.H.P., M.J.K., S.S.K., E.S., B.B.K., A.S. and B.F.C. analyzed and interpreted the data. W.H.P. and B.F.C. wrote the paper. M.J.K., S.S.K. and A.S. edited the paper.
Corresponding authors
Ethics declarations
Competing interests
B.F.C. is a founder and advisor to Abide Therapeutics, a biotechnology company interested in developing serine hydrolase inhibitors and therapeutics.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–18, Supplementary Table 1 and 2 captions, and Supplementary Tables 3 and 4. (PDF 3492 kb)
Supplementary Note
Synthetic Procedures. (PDF 270 kb)
Supplementary Table 1
Complete ABPP-SILAC data sets. (XLSX 5746 kb)
Supplementary Table 2
HHpred search results for AIG1/ADTRP alignment. (XLSX 20 kb)
Rights and permissions
About this article
Cite this article
Parsons, W., Kolar, M., Kamat, S. et al. AIG1 and ADTRP are atypical integral membrane hydrolases that degrade bioactive FAHFAs. Nat Chem Biol 12, 367–372 (2016). https://doi.org/10.1038/nchembio.2051
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2051
This article is cited by
-
ATGL is a biosynthetic enzyme for fatty acid esters of hydroxy fatty acids
Nature (2022)
-
Adtrp regulates thermogenic activity of adipose tissue via mediating the secretion of S100b
Cellular and Molecular Life Sciences (2022)
-
Origin and recent expansion of an endogenous gammaretroviral lineage in domestic and wild canids
Retrovirology (2019)
-
Metabolites as regulators of insulin sensitivity and metabolism
Nature Reviews Molecular Cell Biology (2018)