Abstract
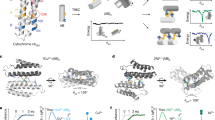
Protein metalloenzymes use various modes for functions for which metal-dependent global conformational change is required in some cases but not in others. In contrast, most ribozymes require a global folding that almost always precedes enzyme reactions. Herein we studied metal-dependent folding and cleavage activity of the 8–17 DNAzyme using single-molecule fluorescence resonance energy transfer. Addition of Zn2+ and Mg2+ induced folding of the DNAzyme into a more compact structure followed by a cleavage reaction, which suggests that the DNAzyme may require metal-dependent global folding for activation. In the presence of Pb2+, however, the cleavage reaction occurred without a precedent folding step, which suggests that the DNAzyme may be prearranged to accept Pb2+ for the activity. Neither ligation reaction of the cleaved substrates nor dynamic changes between folded and unfolded states was observed. These features may contribute to the unusually fast Pb2+-dependent reaction of the DNAzyme. These results suggest that DNAzymes can use all modes of activation that metalloproteins use.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bren, K.L., Pecoraro, V.L. & Gray, H.B. Metalloprotein folding. Inorg. Chem. 43, 7894–7896 (2004).
Kruger, K. et al. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell 31, 147–157 (1982).
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N. & Altman, S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35, 849–857 (1983).
Breaker, R.R. & Joyce, G.F.A. DNA enzyme that cleaves RNA. Chem. Biol. 1, 223–229 (1994).
Sen, D. & Geyer, C.R. DNA enzymes. Curr. Opin. Chem. Biol. 2, 680–687 (1998).
Li, Y. & Breaker, R.R. Deoxyribozymes: new players in the ancient game of biocatalysis. Curr. Opin. Struct. Biol. 9, 315–323 (1999).
Lu, Y. New transition metal-dependent DNAzymes as efficient endonucleases and as selective metal biosensors. Chemistry 8, 4588–4596 (2002).
Santoro, S.W. & Joyce, G.F. A general purpose RNA-cleaving DNA enzyme. Proc. Natl. Acad. Sci. USA 94, 4262–4266 (1997).
Li, J. & Lu, Y. A highly sensitive and selective catalytic DNA biosensor for lead ions. J. Am. Chem. Soc. 122, 10466–10467 (2000).
Liu, J. & Lu, Y. A colorimetric lead biosensor using DNAzyme-directed assembly of gold nanoparticles. J. Am. Chem. Soc. 125, 6642–6643 (2003).
Liu, J. & Lu, Y. Improving fluorescent DNAzyme biosensors by combining inter- and intramolecular quenchers. Anal. Chem. 75, 6666–6672 (2003).
Lu, Y. & Liu, J. Functional DNA nanotechnology: emerging applications of DNAzymes and aptamers. Curr. Opin. Biotechnol. 17, 580–588 (2006).
Lu, Y. & Liu, J. Smart nanomaterials inspired by biology: dynamic assembly of error-free nanomaterials in response to multiple chemical and biological stimuli. Acc. Chem. Res. 40, 315–323 (2007).
Faulhammer, D. & Famulok, M. The Ca2+ ion as a cofactor for a novel RNA-cleaving deoxyribozyme. Angew. Chem. Int. Ed. 35, 2837–2841 (1996).
Li, J., Zheng, W., Kwon, A.H. & Lu, Y. In vitro selection and characterization of a highly efficient Zn(ii)-dependent RNA-cleaving deoxyribozyme. Nucleic Acids Res. 28, 481–488 (2000).
Cruz, R.P.G., Withers, J.B. & Li, Y. Dinucleotide junction cleavage versatility of 8–17 deoxyribozyme. Chem. Biol. 11, 57–67 (2004).
Schlosser, K. & Li, Y. Tracing sequence diversity change of RNA-cleaving deoxyribozymes under increasing selection pressure during in vitro selection. Biochemistry 43, 9695–9707 (2004).
Peracchi, A. Preferential activation of the 8–17 deoxyribozyme by Ca2+ ions. Evidence for the identity of 8–17 with the catalytic domain of the MG5 deoxyribozyme. J. Biol. Chem. 275, 11693–11697 (2000).
Brown, A.K., Li, J., Pavot, C.M.B. & Lu, Y.A. Lead-dependent DNAzyme with a two-step mechanism. Biochemistry 42, 7152–7161 (2003).
Xiao, Y., Rowe, A.A. & Plaxco, K.W. Electrochemical detection of parts-per-billion lead via an electrode-bound DNAzyme assembly. J. Am. Chem. Soc. 129, 262–263 (2007).
Clegg, R.M. Fluorescence resonance energy transfer and nucleic acids. Methods Enzymol. 211, 353–388 (1992).
Lilley, D.M.J. & Wilson, T.J. Fluorescence resonance energy transfer as a structural tool for nucleic acids. Curr. Opin. Chem. Biol. 4, 507–517 (2000).
Bassi, G.S., Murchie, A.I.H., Walter, F., Clegg, R.M. & Lilley, D.M.J. Ion-induced folding of the hammerhead ribozyme: a fluorescence resonance energy transfer study. EMBO J. 16, 7481–7489 (1997).
Walter, N.G., Burke, J.M. & Millar, D.P. Stability of hairpin ribozyme tertiary structure is governed by the interdomain junction. Nat. Struct. Biol. 6, 544–549 (1999).
Fang, X.-W., Pan, T. & Sosnick, T.R. Mg2+-dependent folding of a large ribozyme without kinetic traps. Nat. Struct. Biol. 6, 1091–1095 (1999).
Wilson, T.J. & Lilley, D.M.J. Metal ion binding and the folding of the hairpin ribozyme. RNA 8, 587–600 (2002).
Kim, H.-K. et al. Metal-dependent global folding and activity of the 8–17 DNAzyme studied by fluorescence resonance energy transfer. J. Am. Chem. Soc. 129, 6896–6902 (2007).
Nahas, M.K. et al. Observation of internal cleavage and ligation reactions of a ribozyme. Nat. Struct. Mol. Biol. 11, 1107–1113 (2004).
Ha, T. et al. Probing the interaction between two single molecules: fluorescence resonance energy transfer between a single donor and a single acceptor. Proc. Natl. Acad. Sci. USA 93, 6264–6268 (1996).
Myong, S., Stevens, B.C. & Ha, T. Bridging conformational dynamics and function using single-molecule spectroscopy. Structure 14, 633–643 (2006).
Zhuang, X. et al. Correlating structural dynamics and function in single ribozyme molecules. Science 296, 1473–1476 (2002).
Tan, E. et al. A four-way junction accelerates hairpin ribozyme folding via a discrete intermediate. Proc. Natl. Acad. Sci. USA 100, 9308–9313 (2003).
Bokinsky, G. et al. Single-molecule transition-state analysis of RNA folding. Proc. Natl. Acad. Sci. USA 100, 9302–9307 (2003).
Zhuang, X. et al. A single-molecule study of RNA catalysis and folding. Science 288, 2048–2051 (2000).
Roychowdhury-Saha, M. & Burke, D.H. Distinct reaction pathway promoted by non-divalent-metal cations in a tertiary stabilized hammerhead ribozyme. RNA 13, 841–848 (2007).
Hampel, K.J. & Tinsley, M.M. Evidence for preorganization of the glmS ribozyme ligand binding pocket. Biochemistry 45, 7861–7871 (2006).
Cochrane, J.C., Lipchock, S.V. & Strobel, S.A. Structural investigation of the GlmS ribozyme bound to its catalytic cofactor. Chem. Biol. 14, 97–105 (2007).
Hohng, S. et al. Conformational flexibility of four-way junctions in RNA. J. Mol. Biol. 336, 69–79 (2004).
Rasnik, I., McKinney, S.A. & Ha, T. Nonblinking and long-lasting single-molecule fluorescence imaging. Nat. Methods 3, 891–893 (2006).
Acknowledgements
We thank C. Joo and R. Roy for generous help with performing experiments and data analysis, and J.H. Lee for help with drawing the graphical abstract. This material is based on work supported by the US Department of Energy (DEFG02–01–ER63179), the US National Institutes of Health (GM065367) and the US National Science Foundation (CTS–0120978 and DMI- 0328162). T.H. is a Howard Hughes Medical Institute Investigator.
Author information
Authors and Affiliations
Contributions
H.-K.K. designed and carried out all the experiments and wrote the manuscript; I.R. designed and carried out smFRET experiments; J.L. designed smFRET experiments; T.H. designed smFRET experiments and wrote the manuscript; Y.L. designed all the experiments and wrote the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 275 kb)
Rights and permissions
About this article
Cite this article
Kim, HK., Rasnik, I., Liu, J. et al. Dissecting metal ion–dependent folding and catalysis of a single DNAzyme. Nat Chem Biol 3, 763–768 (2007). https://doi.org/10.1038/nchembio.2007.45
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2007.45
This article is cited by
-
Crystal structure of an RNA-cleaving DNAzyme
Nature Communications (2017)
-
A label-free and portable graphene FET aptasensor for children blood lead detection
Scientific Reports (2016)
-
Understanding DNA-based catalysis one molecule at a time
Nature Chemical Biology (2007)