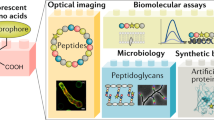
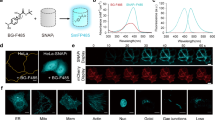
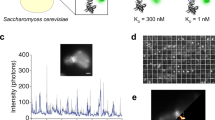
Abstract
The past 20 years have witnessed the advent of numerous technologies to specifically and covalently label proteins in cellulo and in vivo with synthetic probes. These technologies range from self-labeling proteins tags to non-natural amino acids, and the question is no longer how we can specifically label a given protein but rather with what additional functionality we wish to equip it. In addition, progress in fields such as super-resolution microscopy and genome editing have either provided additional motivation to label proteins with advanced synthetic probes or removed some of the difficulties of conducting such experiments. By focusing on two particular applications, live-cell imaging and the generation of reversible protein switches, we outline the opportunities and challenges of the field and how the synergy between synthetic chemistry and protein engineering will make it possible to conduct experiments that are not feasible with conventional approaches.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Yan, Q. & Bruchez, M.P. Advances in chemical labeling of proteins in living cells. Cell Tissue Res. 360, 179–194 (2015).
Jing, C. & Cornish, V.W. Chemical tags for labeling proteins inside living cells. Acc. Chem. Res. 44, 784–792 (2011).
Johnsson, N., George, N. & Johnsson, K. Protein chemistry on the surface of living cells. ChemBioChem 6, 47–52 (2005).
Wang, L. & Schultz, P.G. Expanding the genetic code. Angew. Chem. Int. Ed. Engl. 44, 34–66 (2004).
Davis, L. & Chin, J.W. Designer proteins: applications of genetic code expansion in cell biology. Nat. Rev. Mol. Cell Biol. 13, 168–182 (2012).
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21, 86–89 (2003).
Los, G.V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
Borrmann, A. et al. Genetic encoding of a bicyclo[6.1.0]nonyne-charged amino acid enables fast cellular protein imaging by metal-free ligation. ChemBioChem 13, 2094–2099 (2012).
Lang, K. et al. Genetically encoded norbornene directs site-specific cellular protein labelling via a rapid bioorthogonal reaction. Nat. Chem. 4, 298–304 (2012).
Lang, K. et al. Genetic encoding of bicyclononynes and trans-cyclooctenes for site-specific protein labeling in vitro and in live mammalian cells via rapid fluorogenic Diels–Alder reactions. J. Am. Chem. Soc. 134, 10317–10320 (2012).
Plass, T. et al. Amino acids for Diels-Alder reactions in living cells. Angew. Chem. Int. Ed. Engl. 51, 4166–4170 (2012).
Nikić, I. et al. Minimal tags for rapid dual-color live-cell labeling and super-resolution microscopy. Angew. Chem. Int. Ed. Engl. 53, 2245–2249 (2014).
Sachdeva, A., Wang, K., Elliott, T. & Chin, J.W. Concerted, rapid, quantitative, and site-specific dual labeling of proteins. J. Am. Chem. Soc. 136, 7785–7788 (2014).
Wang, K. et al. Optimized orthogonal translation of unnatural amino acids enables spontaneous protein double-labelling and FRET. Nat. Chem. 6, 393–403 (2014).
Machida, T., Lang, K., Xue, L., Chin, J.W. & Winssinger, N. Site-specific glycoconjugation of protein via bioorthogonal tetrazine cycloaddition with a genetically encoded trans-cyclooctene or bicyclononyne. Bioconjug. Chem. 26, 802–806 (2015).
Uttamapinant, C. et al. Genetic code expansion enables live-cell and super-resolution imaging of site-specifically labeled cellular proteins. J. Am. Chem. Soc. 137, 4602–4605 (2015).
Lukinavičius, G. et al. Selective chemical crosslinking reveals a Cep57-Cep63-Cep152 centrosomal complex. Curr. Biol. 23, 265–270 (2013).
Doudna, J.A. & Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Lang, K. & Chin, J.W. Bioorthogonal reactions for labeling proteins. ACS Chem. Biol. 9, 16–20 (2014).
Greiss, S. & Chin, J.W. Expanding the genetic code of an animal. J. Am. Chem. Soc. 133, 14196–14199 (2011).
Bianco, A., Townsley, F.M., Greiss, S., Lang, K. & Chin, J.W. Expanding the genetic code of Drosophila melanogaster. Nat. Chem. Biol. 8, 748–750 (2012).
Albizu, L. et al. Time-resolved FRET between GPCR ligands reveals oligomers in native tissues. Nat. Chem. Biol. 6, 587–594 (2010).
Hounsou, C. et al. Time-resolved FRET binding assay to investigate h-oligomer binding properties: proof of concept with dopamine D1/D3 heterodimer. ACS Chem. Biol. 10, 466–474 (2015).
Oueslati, N. et al. Time-resolved FRET strategy to screen GPCR ligand library. in G Protein-Coupled Receptor Screening Assays Vol. 1272 (eds. Prazeres, D.M.F. & Martins, S.A.M.) 23–36 (Springer New York, 2015).
Weissleder, R. A clearer vision for in vivo imaging. Nat. Biotechnol. 19, 316–317 (2001).
Yuan, L., Lin, W., Zheng, K., He, L. & Huang, W. Far-red to near infrared analyte-responsive fluorescent probes based on organic fluorophore platforms for fluorescence imaging. Chem. Soc. Rev. 42, 622–661 (2013).
Frangioni, J.V. In vivo near-infrared fluorescence imaging. Curr. Opin. Chem. Biol. 7, 626–634 (2003).
Hilderbrand, S.A. & Weissleder, R. Near-infrared fluorescence: application to in vivo molecular imaging. Curr. Opin. Chem. Biol. 14, 71–79 (2010).
Lukinavičius, G. et al. A near-infrared fluorophore for live-cell super-resolution microscopy of cellular proteins. Nat. Chem. 5, 132–139 (2013).
Fu, M., Xiao, Y., Qian, X., Zhao, D. & Xu, Y. A design concept of long-wavelength fluorescent analogs of rhodamine dyes: replacement of oxygen with silicon atom. Chem. Commun. (Camb.) 2008, 1780–1782 (2008).
Koide, Y., Urano, Y., Hanaoka, K., Terai, T. & Nagano, T. Evolution of group 14 rhodamines as platforms for near-infrared fluorescence probes utilizing photoinduced electron transfer. ACS Chem. Biol. 6, 600–608 (2011).
Koide, Y., Urano, Y., Hanaoka, K., Terai, T. & Nagano, T. Development of an Si-rhodamine-based far-red to near-infrared fluorescence probe selective for hypochlorous acid and its applications for biological imaging. J. Am. Chem. Soc. 133, 5680–5682 (2011).
Egawa, T. et al. Development of a far-red to near-infrared fluorescence probe for calcium ion and its application to multicolor neuronal imaging. J. Am. Chem. Soc. 133, 14157–14159 (2011).
Yang, G. et al. Genetic targeting of chemical indicators in vivo. Nat. Methods 12, 137–139 (2015).
Grimm, J.B. et al. A general method to improve fluorophores for live-cell and single-molecule microscopy. Nat. Methods 12, 244–250 (2015).
Erdmann, R.S. et al. Super-resolution imaging of the Golgi in live cells with a bioorthogonal ceramide probe. Angew. Chem. Int. Ed. Engl. 53, 10242–10246 (2014).
Lukinavičius, G. et al. SiR–Hoechst is a far-red DNA stain for live-cell nanoscopy. Nat. Commun. 6, 8497 (2015).
Jack, T. et al. Characterizing new fluorescent tools for studying 5–HT3 receptor pharmacology. Neuropharmacology 90, 63–73 (2015).
Kim, E., Yang, K.S., Giedt, R.J. & Weissleder, R. Red Si–rhodamine drug conjugates enable imaging in GFP cells. Chem. Commun. (Camb.) 50, 4504–4507 (2014).
Lukinavičius, G. et al. Fluorogenic probes for live-cell imaging of the cytoskeleton. Nat. Methods 11, 731–733 (2014).
Wysocki, L.M. & Lavis, L.D. Advances in the chemistry of small molecule fluorescent probes. Curr. Opin. Chem. Biol. 15, 752–759 (2011).
Lavis, L.D. & Raines, R.T. Bright building blocks for chemical biology. ACS Chem. Biol. 9, 855–866 (2014).
Li, X., Gao, X., Shi, W. & Ma, H. Design strategies for water-soluble small molecular chromogenic and fluorogenic probes. Chem. Rev. 114, 590–659 (2014).
Liu, T.-K. et al. A rapid SNAP-tag fluorogenic probe based on an environment-sensitive fluorophore for no-wash live cell imaging. ACS Chem. Biol. 9, 2359–2365 (2014).
Komatsu, T. et al. Real-time measurements of protein dynamics using fluorescence activation-coupled protein labeling method. J. Am. Chem. Soc. 133, 6745–6751 (2011).
Hell, S.W. Nanoscopy with focused light (Nobel Lecture). Angew. Chem. Int. Ed. Engl. 54, 8054–8066 (2015).
Uno, S.-N. et al. A spontaneously blinking fluorophore based on intramolecular spirocyclization for live-cell super-resolution imaging. Nat. Chem. 6, 681–689 (2014).
Deschout, H. et al. Precisely and accurately localizing single emitters in fluorescence microscopy. Nat. Methods 11, 253–266 (2014).
Fornasiero, E.F. & Opazo, F. Super-resolution imaging for cell biologists. BioEssays 37, 436–451 (2015).
Tsukiji, S. & Hamachi, I. Ligand-directed tosyl chemistry for in situ native protein labeling and engineering in living systems: from basic properties to applications. Curr. Opin. Chem. Biol. 21, 136–143 (2014).
Takaoka, Y., Ojida, A. & Hamachi, I. Protein organic chemistry and applications for labeling and engineering in live-cell systems. Angew. Chem. Int. Ed. Engl. 52, 4088–4106 (2013).
Hayashi, T. & Hamachi, I. Traceless affinity labeling of endogenous proteins for functional analysis in living cells. Acc. Chem. Res. 45, 1460–1469 (2012).
Miki, T. et al. LDAI-based chemical labeling of intact membrane proteins and its pulse-chase analysis under live cell conditions. Chem. Biol. 21, 1013–1022 (2014).
Krishnamurthy, V.M., Semetey, V., Bracher, P.J., Shen, N. & Whitesides, G.M. Dependence of effective molarity on linker length for an intramolecular protein−ligand system. J. Am. Chem. Soc. 129, 1312–1320 (2007).
Zhou, H.-X. Quantitative relation between intermolecular and intramolecular binding of Pro-rich peptides to SH3 domains. Biophys. J. 91, 3170–3181 (2006).
Zheng, Q. et al. Ultra-stable organic fluorophores for single-molecule research. Chem. Soc. Rev. 43, 1044–1056 (2014).
Erazo-Oliveras, A. et al. Protein delivery into live cells by incubation with an endosomolytic agent. Nat. Methods 11, 861–867 (2014).
Stein, V. & Alexandrov, K. Synthetic protein switches: design principles and applications. Trends Biotechnol. 33, 101–110 (2015).
Szymański, W., Beierle, J.M., Kistemaker, H.A.V., Velema, W.A. & Feringa, B.L. Reversible photocontrol of biological systems by the incorporation of molecular photoswitches. Chem. Rev. 113, 6114–6178 (2013).
Banghart, M., Borges, K., Isacoff, E., Trauner, D. & Kramer, R.H. Light-activated ion channels for remote control of neuronal firing. Nat. Neurosci. 7, 1381–1386 (2004).
Gorostiza, P. et al. Mechanisms of photoswitch conjugation and light activation of an ionotropic glutamate receptor. Proc. Natl. Acad. Sci. USA 104, 10865–10870 (2007).
Numano, R. et al. Nanosculpting reversed wavelength sensitivity into a photoswitchable iGluR. Proc. Natl. Acad. Sci. USA 106, 6814–6819 (2009).
Volgraf, M. et al. Allosteric control of an ionotropic glutamate receptor with an optical switch. Nat. Chem. Biol. 2, 47–52 (2006).
Caporale, N. et al. LiGluR restores visual responses in rodent models of inherited blindness. Mol. Ther. 19, 1212–1219 (2011).
Levitz, J. et al. Optical control of metabotropic glutamate receptors. Nat. Neurosci. 16, 507–516 (2013).
Wyart, C. et al. Optogenetic dissection of a behavioural module in the vertebrate spinal cord. Nature 461, 407–410 (2009).
Tsai, Y.-H., Essig, S., James, J.R., Lang, K. & Chin, J.W. Selective, rapid and optically switchable regulation of protein function in live mammalian cells. Nat. Chem. 7, 554–561 (2015).
Broichhagen, J., Frank, J.A. & Trauner, D. A roadmap to success in photopharmacology. Acc. Chem. Res. 48, 1947–1960 (2015).
Masharina, A., Reymond, L., Maurel, D., Umezawa, K. & Johnsson, K. A Fluorescent sensor for GABA and synthetic GABA B receptor ligands. J. Am. Chem. Soc. 134, 19026–19034 (2012).
Brun, M.A. et al. Semisynthesis of fluorescent metabolite sensors on cell surfaces. J. Am. Chem. Soc. 133, 16235–16242 (2011).
Brun, M.A. et al. A semisynthetic fluorescent sensor protein for glutamate. J. Am. Chem. Soc. 134, 7676–7678 (2012).
Schena, A. & Johnsson, K. Sensing acetylcholine and anticholinesterase compounds. Angew. Chem. Int. Ed. Engl. 53, 1302–1305 (2014).
Griss, R. et al. Bioluminescent sensor proteins for point-of-care therapeutic drug monitoring. Nat. Chem. Biol. 10, 598–603 (2014).
Schena, A., Griss, R. & Johnsson, K. Modulating protein activity using tethered ligands with mutually exclusive binding sites. Nat. Commun. 6, 7830 (2015).
Acknowledgements
The Johnsson laboratory acknowledges support from the Swiss National Science Foundation and the NCCR Chemical Biology. The authors are grateful to P. Heppenstall (EMBL Monterotondo) and G. Lukinavicius (EPFL) for providing images. I.A.K. acknowledges funding through an EMBO Long-Term Fellowship (ALTF 302-2015) co-funded by Marie Curie Action (LTFCOFUND2013, GA-2013-609409).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
K.J. has filed and licensed out patent applications on different protein labeling technologies and fluorophores that are mentioned in this article.
Rights and permissions
About this article
Cite this article
Xue, L., Karpenko, I., Hiblot, J. et al. Imaging and manipulating proteins in live cells through covalent labeling. Nat Chem Biol 11, 917–923 (2015). https://doi.org/10.1038/nchembio.1959
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1959
This article is cited by
-
Location-agnostic site-specific protein bioconjugation via Baylis Hillman adducts
Nature Communications (2024)
-
Light-induced Pd catalyst enables C(sp2)–C(sp2) cross-electrophile coupling bypassing the demand for transmetalation
Nature Catalysis (2024)
-
A general method for the development of multicolor biosensors with large dynamic ranges
Nature Chemical Biology (2023)
-
Design of a palette of SNAP-tag mimics of fluorescent proteins and their use as cell reporters
Cell Discovery (2023)
-
A general design of caging-group-free photoactivatable fluorophores for live-cell nanoscopy
Nature Chemistry (2022)