Abstract
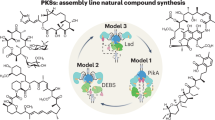
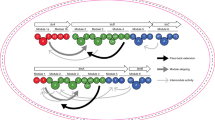
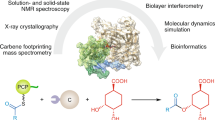
The modular polyketide synthases (PKSs) and nonribosomal peptide synthetases (NRPSs) are among the largest and most complicated enzymes in nature. In these biosynthetic systems, independently folding protein domains, which are organized into units called 'modules', operate in assembly-line fashion to construct polymeric chains and tailor their functionalities. Products of PKSs and NRPSs include a number of blockbuster medicines, and this has motivated researchers to understand how they operate so that they can be modified by genetic engineering. Beginning in the 1990s, structural biology has provided a number of key insights. The emerging picture is one of remarkable dynamics and conformational programming in which the chemical states of individual catalytic domains are communicated to the others, configuring the modules for the next stage in the biosynthesis. This unexpected level of complexity most likely accounts for the low success rate of empirical genetic engineering experiments and suggests ways forward for productive megaenzyme synthetic biology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Cortés, J., Haydock, S.F., Roberts, G.A., Bevitt, D.J. & Leadlay, P.F. An unusually large multifunctional polypeptide in the erythromycin-producing polyketide synthase of Saccharopolyspora erythraea. Nature 348, 176–178 (1990).
Donadio, S., Staver, M.J., McAlpine, J.B., Swanson, S.J. & Katz, L. Modular organization of genes required for complex polyketide biosynthesis. Science 252, 675–679 (1991).
Skarpeid, H.J., Zimmer, T.L. & von Döhren, H. On the domain construction of the multienzyme gramicidin S synthetase 2. Isolation of domains activating valine and leucine. Eur. J. Biochem. 189, 517–522 (1990).
Lawen, A. & Zocher, R. Cyclosporin synthetase. The most complex peptide synthesizing multienzyme polypeptide so far described. J. Biol. Chem. 265, 11355–11360 (1990).
Stinear, T.P. et al. Giant plasmid-encoded polyketide synthases produce the macrolide toxin of Mycobacterium ulcerans. Proc. Natl. Acad. Sci. USA 101, 1345–1349 (2004).
Walsh, C.T. et al. Tailoring enzymes that modify nonribosomal peptides during and after chain elongation on NRPS assembly lines. Curr. Opin. Chem. Biol. 5, 525–534 (2001).
Aparicio, J.F. et al. Organization of the biosynthetic gene cluster for rapamycin in Streptomyces hygroscopicus: analysis of the enzymatic domains in the modular polyketide synthase. Gene 169, 9–16 (1996).
Piel, J. Biosynthesis of polyketides by trans-AT polyketide synthases. Nat. Prod. Rep. 27, 996–1047 (2010).
Nguyen, T. et al. Exploiting the mosaic structure of trans-acyltransferase polyketide synthases for natural product discovery and pathway dissection. Nat. Biotechnol. 26, 225–233 (2008).
Mootz, H.D., Schwarzer, D. & Marahiel, M.A. Ways of assembling complex natural products on modular nonribosomal peptide synthetases. ChemBioChem 3, 490–504 (2002).
Wakil, S.J. Fatty acid synthase, a proficient multifunctional enzyme. Biochemistry 28, 4523–4530 (1989).
Díez, B. et al. The cluster of penicillin biosynthetic genes. Identification and characterization of the pcbAB gene encoding the a-aminoadipyl-cysteinyl-valine synthetase and linkage to the pcbC and penDE genes. J. Biol. Chem. 265, 16358–16365 (1990).
Stachelhaus, T. & Marahiel, M.A. Modular structure of peptide synthetases revealed by dissection of the multifunctional enzyme GrsA. J. Biol. Chem. 270, 6163–6169 (1995).
De Crécy-Lagard, V., Marlière, P. & Saurin, W. Multienzymatic non ribosomal peptide biosynthesis: identification of the functional domains catalysing peptide elongation and epimerisation. C.R. Acad. Sci. III 318, 927–936 (1995).
Caffrey, P., Bevitt, D.J., Staunton, J. & Leadlay, P.F. Identification of DEBS 1, DEBS 2 and DEBS 3, the multienzyme polypeptides of the erythromycin-producing polyketide synthase from Saccharopolyspora erythraea. FEBS Lett. 304, 225–228 (1992).
Staunton, J. et al. Evidence for a double-helical structure for modular polyketide synthases. Nat. Struct. Biol. 3, 188–192 (1996).
Gevers, W., Kleinkauf, H. & Lipmann, F. Peptidyl transfers in gramicidin S biosynthesis from enzyme-bound thioester intermediates. Proc. Natl. Acad. Sci. USA 63, 1335–1342 (1969).
Sieber, S.A. et al. Evidence for a monomeric structure of nonribosomal peptide synthetases. Chem. Biol. 9, 997–1008 (2002).
Hillson, N.J. & Walsh, C.T. Dimeric structure of the six-domain VibF subunit of vibriobactin synthetase: mutant domain activity regain and ultracentrifugation studies. Biochemistry 42, 766–775 (2003).
Lu, H., Tsai, S.-C., Khosla, C. & Cane, D.E. Expression, site-directed mutagenesis, and steady state kinetic analysis of the terminal thioesterase domain of the methymycin/picromycin polyketide synthase. Biochemistry 41, 12590–12597 (2002).
Gehret, J.J. et al. Terminal alkene formation by the thioesterase of curacin A biosynthesis: structure of a decarboxylating thioesterase. J. Biol. Chem. 286, 14445–14454 (2011).
Tsai, S.C. et al. Crystal structure of the macrocycle-forming thioesterase domain of the erythromycin polyketide synthase: versatility from a unique substrate channel. Proc. Natl. Acad. Sci. USA 98, 14808–14813 (2001).
Tsai, S.-C., Lu, H., Cane, D.E., Khosla, C. & Stroud, R.M. Insights into channel architecture and substrate specificity from crystal structures of two macrocycle-forming thioesterases of modular polyketide synthases. Biochemistry 41, 12598–12606 (2002).
Akey, D.L. et al. Structural basis for macrolactonization by the pikromycin thioesterase. Nat. Chem. Biol. 2, 537–542 (2006). This paper and the next were used to develop a model for how thioesterases promote macrocyclization of their linear polyketide substrates.
Giraldes, J.W. et al. Structural and mechanistic insights into polyketide macrolactonization from polyketide-based affinity labels. Nat. Chem. Biol. 2, 531–536 (2006).
Scaglione, J.B. et al. Biochemical and structural characterization of the tautomycetin thioesterase: analysis of a stereoselective polyketide hydrolase. Angew. Chem. Int. Edn Engl. 49, 5726–5730 (2010).
Tang, Y., Kim, C.-Y., Mathews, I.I., Cane, D.E. & Khosla, C. The 2.7-Å crystal structure of a 194-kDa homodimeric fragment of the 6-deoxyerythronolide B synthase. Proc. Natl. Acad. Sci. USA 103, 11124–11129 (2006).
Tang, Y., Chen, A.Y., Kim, C.-Y., Cane, D.E. & Khosla, C. Structural and mechanistic analysis of protein interactions in module 3 of the 6-deoxyerythronolide B synthase. Chem. Biol. 14, 931–943 (2007).
Keatinge-Clay, A.T. & Stroud, R.M. The structure of a ketoreductase determines the organization of the β-carbon processing enzymes of modular polyketide synthases. Structure 14, 737–748 (2006).
Keatinge-Clay, A.T. A tylosin ketoreductase reveals how chirality is determined in polyketides. Chem. Biol. 14, 898–908 (2007). This paper reported sequence motifs in ketoreductase domains that allows their assignment into six functional classes.
Zheng, J., Taylor, C.A., Piasecki, S.K. & Keatinge-Clay, A.T. Structural and functional analysis of A-type ketoreductases from the amphotericin modular polyketide synthase. Structure 18, 913–922 (2010).
Zheng, J. & Keatinge-Clay, A.T. Structural and functional analysis of C2-type ketoreductases from modular polyketide synthases. J. Mol. Biol. 410, 105–117 (2011).
Zheng, J., Gay, D.C., Demeler, B., White, M.A. & Keatinge-Clay, A.T. Divergence of multimodular polyketide synthases revealed by a didomain structure. Nat. Chem. Biol. 8, 615–621 (2012).
Zheng, J., Piasecki, S.K. & Keatinge-Clay, A.T. Structural studies of an A2-type modular polyketide synthase ketoreductase reveal features controlling α-substituent stereochemistry. ACS Chem. Biol. 8, 1964–1971 (2013).
Bonnett, S.A. et al. Structural and stereochemical analysis of a modular polyketide synthase ketoreductase domain required for the generation of a cis-alkene. Chem. Biol. 20, 772–783 (2013). This paper suggests a new, unified model for stereocontrol by A- and B-type KR domains.
Keatinge-Clay, A. Crystal structure of the erythromycin polyketide synthase dehydratase. J. Mol. Biol. 384, 941–953 (2008).
Akey, D.L. et al. Crystal structures of dehydratase domains from the curacin polyketide biosynthetic pathway. Structure 18, 94–105 (2010).
Alekseyev, V.Y., Liu, C.W., Cane, D.E., Puglisi, J.D. & Khosla, C. Solution structure and proposed domain domain recognition interface of an acyl carrier protein domain from a modular polyketide synthase. Protein Sci. 16, 2093–2107 (2007).
Tran, L., Broadhurst, R.W., Tosin, M., Cavalli, A. & Weissman, K.J. Insights into protein-protein and enzyme-substrate interactions in modular polyketide synthases. Chem. Biol. 17, 705–716 (2010). This paper provides the only evidence to date for an ACP-partner interaction in cis -AT PKSs that occurs in the absence of formation of a specific domain-domain complex.
Busche, A. et al. Characterization of molecular interactions between ACP and halogenase domains in the curacin A polyketide synthase. ACS Chem. Biol. 7, 378–386 (2012).
Zheng, J., Fage, C.D., Demeler, B., Hoffman, D.W. & Keatinge-Clay, A.T. The missing linker: a dimerization motif located within polyketide synthase modules. ACS Chem. Biol. 8, 1263–1270 (2013).
Broadhurst, R.W., Nietlispach, D., Wheatcroft, M.P., Leadlay, P.F. & Weissman, K.J. The structure of docking domains in modular polyketide synthases. Chem. Biol. 10, 723–731 (2003).
Buchholz, T.J. et al. Structural basis for binding specificity between subclasses of modular polyketide synthase docking domains. ACS Chem. Biol. 4, 41–52 (2009).
Whicher, J.R. et al. Cyanobacterial polyketide synthase docking domains: a tool for engineering natural product biosynthesis. Chem. Biol. 20, 1340–1351 (2013).
Xu, W., Qiao, K. & Tang, Y. Structural analysis of protein-protein interactions in type I polyketide synthases. Crit. Rev. Biochem. Mol. Biol. 48, 98–122 (2013).
Smith, S., Witkowski, A. & Joshi, A.K. Structural and functional organization of the animal fatty acid synthase. Prog. Lipid Res. 42, 289–317 (2003).
Marsden, A.F. et al. Stereospecific acyl transfers on the erythromycin-producing polyketide synthase. Science 263, 378–380 (1994).
Del Vecchio, F. et al. Active-site residue, domain and module swaps in modular polyketide synthases. J. Ind. Microbiol. Biotechnol. 30, 489–494 (2003).
Weissman, K.J. et al. The molecular basis of Celmer's rules: the stereochemistry of the condensation step in chain extension on the erythromycin polyketide synthase. Biochemistry 36, 13849–13855 (1997).
Holzbaur, I.E. et al. Molecular basis of Celmer's rules: role of the ketosynthase domain in epimerisation and demonstration that ketoreductase domains can have altered product specificity with unnatural substrates. Chem. Biol. 8, 329–340 (2001).
Garg, A., Xie, X., Keatinge-Clay, A., Khosla, C. & Cane, D.E. Elucidation of the cryptic epimerase activity of redox-inactive ketoreductase domains from modular polyketide synthases by tandem equilibrium isotope exchange. J. Am. Chem. Soc. 136, 10190–10193 (2014).
Annaval, T., Paris, C., Leadlay, P.F., Jacob, C. & Weissman, K.J. Evaluating ketoreductase exchanges as a means of rationally altering polyketide stereochemistry. ChemBioChem 16, 1357–1364 (2015).
Caffrey, P. Conserved amino acid residues correlating with ketoreductase stereospecificity in modular polyketide synthases. ChemBioChem 4, 654–657 (2003).
Reid, R. et al. A model of structure and catalysis for ketoreductase domains in modular polyketide synthases. Biochemistry 42, 72–79 (2003).
Aggarwal, R., Caffrey, P., Leadlay, P., Smith, C. & Staunton, J. The thioesterase of the erythromycin-producing polyketide synthase: mechanistic studies in vitro to investigate its mode of action and substrate specificity. J. Chem. Soc. Chem. Commun. 1519–1520 (1995).
Cortés, J. et al. Repositioning of a domain in a modular polyketide synthase to promote specific chain cleavage. Science 268, 1487–1489 (1995).
Wong, F.T., Jin, X., Mathews, I.I., Cane, D.E. & Khosla, C. Structure and mechanism of the trans-acting acyltransferase from the disorazole synthase. Biochemistry 50, 6539–6548 (2011).
Bretschneider, T. et al. Vinylogous chain branching catalysed by a dedicated polyketide synthase module. Nature 502, 124–128 (2013).
Gay, D.C. et al. A close look at a ketosynthase from a trans-acyltransferase modular polyketide synthase. Structure 22, 444–451 (2014). This paper reports the crystal structure of a trans -AT PKS ketosynthase domain in the presence of native substrate, enabling identification of putative specificity determinants.
Haines, A.S. et al. A conserved motif flags acyl carrier proteins for β-branching in polyketide synthesis. Nat. Chem. Biol. 9, 685–692 (2013).
Davison, J. et al. Insights into the function of trans-acyl transferase polyketide synthases from the SAXS structure of a complete module. Chem. Sci. 5, 3081–3095 (2014). This paper describes the characterization by SAXS of the first structure of an intact module from a trans -AT PKS.
Piasecki, S.K., Zheng, J., Axelrod, A.J., Detelich, E.M. & Keatinge-Clay, A.T. Structural and functional studies of a trans-acyltransferase polyketide assembly line enzyme that catalyzes stereoselective α- and β-ketoreduction. Proteins 82, 2067–2077 (2014).
Gay, D.C., Spear, P.J. & Keatinge-Clay, A.T. A double-hotdog with a new trick: structure and mechanism of the trans-acyltransferase polyketide synthase enoyl-isomerase. ACS Chem. Biol. 9, 2374–2381 (2014).
Keating, T.A., Marshall, C.G., Walsh, C.T. & Keating, A.E. The structure of VibH represents nonribosomal peptide synthetase condensation, cyclization and epimerization domains. Nat. Struct. Biol. 9, 522–526 (2002).
Bloudoff, K., Rodionov, D. & Schmeing, T.M. Crystal structures of the first condensation domain of CDA synthetase suggest conformational changes during the synthetic cycle of nonribosomal peptide synthetases. J. Mol. Biol. 425, 3137–3150 (2013).
Conti, E., Stachelhaus, T., Marahiel, M.A. & Brick, P. Structural basis for the activation of phenylalanine in the non-ribosomal biosynthesis of gramicidin S. EMBO J. 16, 4174–4183 (1997). This structure formed the basis for a nonribosomal 'code' that allows prediction of A domain specificity in newly discovered gene clusters.
May, J.J., Kessler, N., Marahiel, M.A. & Stubbs, M.T. Crystal structure of DhbE, an archetype for aryl acid activating domains of modular nonribosomal peptide synthetases. Proc. Natl. Acad. Sci. USA 99, 12120–12125 (2002).
Osman, K.T., Du, L., He, Y. & Luo, Y. Crystal structure of Bacillus cereus D-alanyl carrier protein ligase (DltA) in complex with ATP. J. Mol. Biol. 388, 345–355 (2009).
Drake, E.J., Duckworth, B.P., Neres, J., Aldrich, C.C. & Gulick, A.M. Biochemical and structural characterization of bisubstrate inhibitors of BasE, the self-standing nonribosomal peptide synthetase adenylate-forming enzyme of acinetobactin synthesis. Biochemistry 49, 9292–9305 (2010).
Weber, T., Baumgartner, R., Renner, C., Marahiel, M.A. & Holak, T.A. Solution structure of PCP, a prototype for the peptidyl carrier domains of modular peptide synthetases. Structure 8, 407–418 (2000).
Koglin, A. et al. Conformational switches modulate protein interactions in peptide antibiotic synthetases. Science 312, 273–276 (2006).
Lohman, J.R. et al. The crystal structure of BlmI as a model for nonribosomal peptide synthetase peptidyl carrier proteins. Proteins 82, 1210–1218 (2014).
Haslinger, K., Redfield, C. & Cryle, M.J. Structure of the terminal PCP domain of the non-ribosomal peptide synthetase in teicoplanin biosynthesis. Proteins 83, 711–721 (2015).
Bruner, S.D. et al. Structural basis for the cyclization of the lipopeptide antibiotic surfactin by the thioesterase domain SrfTE. Structure 10, 301–310 (2002).
Samel, S.A., Wagner, B., Marahiel, M.A. & Essen, L.-O. The thioesterase domain of the fengycin biosynthesis cluster: a structural base for the macrocyclization of a non-ribosomal lipopeptide. J. Mol. Biol. 359, 876–889 (2006).
Samel, S.A., Czodrowski, P. & Essen, L.-O. Structure of the epimerization domain of tyrocidine synthetase A. Acta Crystallogr. D Biol. Crystallogr. 70, 1442–1452 (2014).
Koglin, A. et al. Structural basis for the selectivity of the external thioesterase of the surfactin synthetase. Nature 454, 907–911 (2008).
Tufar, P. et al. Crystal structure of a PCP/Sfp complex reveals the structural basis for carrier protein posttranslational modification. Chem. Biol. 21, 552–562 (2014).
Samel, S.A., Schoenafinger, G., Knappe, T.A., Marahiel, M.A. & Essen, L.-O. Structural and functional insights into a peptide bond–forming bidomain from a nonribosomal peptide synthetase. Structure 15, 781–792 (2007).
Liu, Y., Zheng, T. & Bruner, S.D. Structural basis for phosphopantetheinyl carrier domain interactions in the terminal module of nonribosomal peptide synthetases. Chem. Biol. 18, 1482–1488 (2011).
Mitchell, C.A., Shi, C., Aldrich, C.C. & Gulick, A.M. Structure of PA1221, a nonribosomal peptide synthetase containing adenylation and peptidyl carrier protein domains. Biochemistry 51, 3252–3263 (2012).
Sundlov, J.A., Shi, C., Wilson, D.J., Aldrich, C.C. & Gulick, A.M. Structural and functional investigation of the intermolecular interaction between NRPS adenylation and carrier protein domains. Chem. Biol. 19, 188–198 (2012).
Tan, X.F. et al. Structure of the adenylation-peptidyl carrier protein didomain of the Microcystis aeruginosa microcystin synthetase McyG. Acta Crystallogr. D Biol. Crystallogr. 71, 873–881 (2015).
Tanovic, A., Samel, S.A., Essen, L.-O. & Marahiel, M.A. Crystal structure of the termination module of a nonribosomal peptide synthetase. Science 321, 659–663 (2008). This paper describes the first X-ray structure of an intact NRPS termination module.
Gulick, A.M. Conformational dynamics in the acyl-CoA synthetases, adenylation domains of non-ribosomal peptide synthetases, and firefly luciferase. ACS Chem. Biol. 4, 811–827 (2009).
Strieker, M., Tanović, A. & Marahiel, M.A. Nonribosomal peptide synthetases: structures and dynamics. Curr. Opin. Struct. Biol. 20, 234–240 (2010).
Stachelhaus, T., Mootz, H.D. & Marahiel, M.A. The specificity-conferring code of adenylation domains in nonribosomal peptide synthetases. Chem. Biol. 6, 493–505 (1999).
Challis, G.L., Ravel, J. & Townsend, C.A. Predictive, structure-based model of amino acid recognition by nonribosomal peptide synthetase adenylation domains. Chem. Biol. 7, 211–224 (2000).
Röttig, M. et al. NRPSpredictor2—a web server for predicting NRPS adenylation domain specificity. Nucleic Acids Res. 39, W362–W367 (2011).
Thirlway, J. et al. Introduction of a non-natural amino acid into a nonribosomal peptide antibiotic by modification of adenylation domain specificity. Angew. Chem. Int. Ed. Engl. 51, 7181–7184 (2012).
Frueh, D.P. et al. Dynamic thiolation–thioesterase structure of a non-ribosomal peptide synthetase. Nature 454, 903–906 (2008).
Gehring, A.M., Mori, I. & Walsh, C.T. Reconstitution and characterization of the Escherichia coli enterobactin synthetase from EntB, EntE, and EntF. Biochemistry 37, 2648–2659 (1998).
Smith, J.L., Skiniotis, G. & Sherman, D.H. Architecture of the polyketide synthase module: surprises from electron cryo-microscopy. Curr. Opin. Struct. Biol. 31, 9–19 (2015).
Maier, T., Leibundgut, M. & Ban, N. The crystal structure of a mammalian fatty acid synthase. Science 321, 1315–1322 (2008).
Dutta, S. et al. Structure of a modular polyketide synthase. Nature 510, 512–517 (2014). This paper and the next report the cryo-EM structures of a model cis -AT PKS module at multiple stages of the catalytic cycle.
Whicher, J.R. et al. Structural rearrangements of a polyketide synthase module during its catalytic cycle. Nature 510, 560–564 (2014).
Edwards, A.L., Matsui, T., Weiss, T.M. & Khosla, C. Architectures of whole-module and bimodular proteins from the 6-deoxyerythronolide B synthase. J. Mol. Biol. 426, 2229–2245 (2014).
Rittner, A. & Grininger, M. Modular polyketide synthases (PKSs): a new model fits all? ChemBioChem 15, 2489–2493 (2014).
Weissman, K.J. Uncovering the structures of modular polyketide synthases. Nat. Prod. Rep. 32, 436–453 (2015).
Anselmi, C., Grininger, M., Gipson, P. & Faraldo-Gómez, J.D. Mechanism of substrate shuttling by the acyl-carrier protein within the fatty acid mega-synthase. J. Am. Chem. Soc. 132, 12357–12364 (2010).
Acknowledgements
Work in the author's laboratory is supported by the Agence Nationale de la Recherche (ANR JCJC 2011 PKS-PPIs, K.J.W.), the Centre National de la Recherche Scientifique (CNRS), and the University of Lorraine and the Lorraine Region (BQR grants to K.J.W. and B. Chagot). P.F. Leadlay is gratefully acknowledged for helpful comments and critical reading of the manuscript. T. Annaval and A. Gruez are thanked for help with figure preparation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing financial interests.
Rights and permissions
About this article
Cite this article
Weissman, K. The structural biology of biosynthetic megaenzymes. Nat Chem Biol 11, 660–670 (2015). https://doi.org/10.1038/nchembio.1883
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1883
This article is cited by
-
Enzymology of assembly line synthesis by modular polyketide synthases
Nature Chemical Biology (2023)
-
Chemoenzymatic synthesis of fluorinated polyketides
Nature Chemistry (2022)
-
Structures and function of a tailoring oxidase in complex with a nonribosomal peptide synthetase module
Nature Communications (2022)
-
Engineering the stambomycin modular polyketide synthase yields 37-membered mini-stambomycins
Nature Communications (2022)
-
Engineering the acyltransferase domain of epothilone polyketide synthase to alter the substrate specificity
Microbial Cell Factories (2021)