Abstract
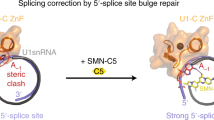
Spinal muscular atrophy (SMA), which results from the loss of expression of the survival of motor neuron-1 (SMN1) gene, represents the most common genetic cause of pediatric mortality. A duplicate copy (SMN2) is inefficiently spliced, producing a truncated and unstable protein. We describe herein a potent, orally active, small-molecule enhancer of SMN2 splicing that elevates full-length SMN protein and extends survival in a severe SMA mouse model. We demonstrate that the molecular mechanism of action is via stabilization of the transient double-strand RNA structure formed by the SMN2 pre-mRNA and U1 small nuclear ribonucleic protein (snRNP) complex. The binding affinity of U1 snRNP to the 5′ splice site is increased in a sequence-selective manner, discrete from constitutive recognition. This new mechanism demonstrates the feasibility of small molecule–mediated, sequence-selective splice modulation and the potential for leveraging this strategy in other splicing diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
15 July 2015
The authors forgot to include a description of the measurement of SMN protein levels in tissue samples in the Online Methods. This description has been included in the HTML and PDF versions of the article.
11 February 2016
In the version of this article originally published online, the schematic for the construct in Figure 4a was incorrect. A corrected figure has been provided in the HTML and PDF versions of the article.
References
Sugarman, E.A. et al. Pan-ethnic carrier screening and prenatal diagnosis for spinal muscular atrophy: clinical laboratory analysis of >72,400 specimens. Eur. J. Hum. Genet. 20, 27–32 (2012).
Lunn, M.R. & Wang, C.H. Spinal muscular atrophy. Lancet 371, 2120–2133 (2008).
Zhou, J., Zheng, X. & Shen, H. Targeting RNA-splicing for SMA treatment. Mol. Cells 33, 223–228 (2012).
Kolb, S.J. & Kissel, J.T. Spinal muscular atrophy: a timely review. Arch. Neurol. 68, 979–984 (2011).
Lefebvre, S. et al. Identification and characterization of a spinal muscular atrophy-determining gene. Cell 80, 155–165 (1995).
Monani, U.R. A single nucleotide difference that alters splicing patterns distinguishes the SMA gene SMN1 from the copy gene SMN2. Hum. Mol. Genet. 8, 1177–1183 (1999).
Gavrilov, D.K., Shi, X., Das, K., Gilliam, T.C. & Wang, C.H. Differential SMN2 expression associated with SMA severity. Nat. Genet. 20, 230–231 (1998).
Lorson, C.L., Hahnen, E., Androphy, E.J. & Wirth, B. A single nucleotide in the SMN gene regulates splicing and is responsible for spinal muscular atrophy. Proc. Natl. Acad. Sci. USA 96, 6307–6311 (1999).
Le, T.T. et al. SMNΔ7, the major product of the centromeric survival motor neuron (SMN2) gene, extends survival in mice with spinal muscular atrophy and associates with full-length SMN. Hum. Mol. Genet. 14, 845–857 (2005).
Vitali, T. et al. Detection of the survival motor neuron (SMN) genes by FISH: further evidence for a role for SMN2 in the modulation of disease severity in SMA patients. Hum. Mol. Genet. 8, 2525–2532 (1999).
Monani, U.R. Spinal muscular atrophy: a deficiency in a ubiquitous protein; a motor neuron-specific disease. Neuron 48, 885–896 (2005).
Sumner, C.J. Molecular mechanisms of spinal muscular atrophy. J. Child Neurol. 22, 979–989 (2007).
McAndrew, P.E. et al. Identification of proximal spinal muscular atrophy carriers and patients by analysis of SMNT and SMNC gene copy number. Am. J. Hum. Genet. 60, 1411–1422 (1997).
Matera, A.G. & Wang, Z. A day in the life of the spliceosome. Nat. Rev. Mol. Cell Biol. 15, 108–121 (2014).
Wang, E.T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Wang, Z. & Burge, C.B. Splicing regulation: from a parts list of regulatory elements to an integrated splicing code. RNA 14, 802–813 (2008).
Xiong, H.Y. et al. The human splicing code reveals new insights into the genetic determinants of disease. Science 347, 1254806 (2014).
Hua, Y., Vickers, T.A., Baker, B.F., Bennett, C.F. & Krainer, A.R. Enhancement of SMN2 exon 7 inclusion by antisense oligonucleotides targeting the exon. PLoS Biol. 5, e73 (2007).
Hua, Y., Vickers, T.A., Okunola, H.L., Bennett, C.F. & Krainer, A.R. Antisense masking of an hnRNP A1/A2 intronic splicing silencer corrects SMN2 splicing in transgenic mice. Am. J. Hum. Genet. 82, 834–848 (2008).
Naryshkin, N.A. et al. SMN2 splicing modifiers improve motor function and longevity in mice with spinal muscular atrophy. Science 345, 688–693 (2014).
Roca, X., Krainer, A.R. & Eperon, I.C. Pick one, but be quick: 5′ splice sites and the problems of too many choices. Genes Dev. 27, 129–144 (2013).
Lewandowska, M.A. The missing puzzle piece: splicing mutations. Int. J. Clin. Exp. Pathol. 6, 2675–2682 (2013).
Lara-Pezzi, E., Gómez-Salinero, J., Gatto, A. & García-Pavía, P. The alternative heart: impact of alternative splicing in heart disease. J. Cardiovasc. Transl. Res. 6, 945–955 (2013).
Wang, J., Zhang, J., Li, K., Zhao, W. & Cui, Q. SpliceDisease database: linking RNA splicing and disease. Nucleic Acids Res. 40, D1055–D1059 (2012).
Osborne, M. et al. Characterization of behavioral and neuromuscular junction phenotypes in a novel allelic series of SMA mouse models. Hum. Mol. Genet. 21, 4431–4447 (2012).
Burnett, B.G. et al. Regulation of SMN protein stability. Mol. Cell. Biol. 29, 1107–1115 (2009).
El-Khodor, B.F. et al. Identification of a battery of tests for drug candidate evaluation in the SMNΔ7 neonate model of spinal muscular atrophy. Exp. Neurol. 212, 29–43 (2008).
Le, T.T. et al. Temporal requirement for high SMN expression in SMA mice. Hum. Mol. Genet. 20, 3578–3591 (2011).
Kariya, S. et al. Requirement of enhanced Survival Motoneuron protein imposed during neuromuscular junction maturation. J. Clin. Invest. 124, 785–800 (2014).
Richard, H. et al. Prediction of alternative isoforms from exon expression levels in RNA-Seq experiments. Nucleic Acids Res. 38, e112 (2010).
Goina, E., Skoko, N. & Pagani, F. Binding of DAZAP1 and hnRNPA1/A2 to an exonic splicing silencer in a natural BRCA1 exon 18 mutant. Mol. Cell. Biol. 28, 3850–3860 (2008).
Singh, N.N., Singh, R.N. & Androphy, E.J. Modulating role of RNA structure in alternative splicing of a critical exon in the spinal muscular atrophy genes. Nucleic Acids Res. 35, 371–389 (2007).
Crooks, G.E., Hon, G., Chandonia, J.-M. & Brenner, S.E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Bailey, T.L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Akerman, M., David-Eden, H., Pinter, R.Y. & Mandel-Gutfreund, Y. A computational approach for genome-wide mapping of splicing factor binding sites. Genome Biol. 10, R30 (2009).
Yeo, G. & Burge, C.B. Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J. Comput. Biol. 11, 377–394 (2004).
Salcius, M. et al. SEC-TID: a label-free method for small-molecule target identification. J. Biomol. Screen. 19, 917–927 (2014).
Kondo, Y., Oubridge, C., van Roon, A.-M.M. & Nagai, K. Crystal structure of human U1 snRNP, a small nuclear ribonucleoprotein particle, reveals the mechanism of 5′ splice site recognition. Elife 4, e04986 (2015).
Rigo, F. et al. Pharmacology of a central nervous system delivered 2′-O-methoxyethyl-modified survival of motor neuron splicing oligonucleotide in mice and nonhuman primates. J. Pharmacol. Exp. Ther. 350, 46–55 (2014).
Kotake, Y. et al. Splicing factor SF3b as a target of the antitumor natural product pladienolide. Nat. Chem. Biol. 3, 570–575 (2007).
Zhang, M.L., Lorson, C.L., Androphy, E.J. & Zhou, J. An in vivo reporter system for measuring increased inclusion of exon 7 in SMN2 mRNA: potential therapy of SMA. Gene Ther. 8, 1532–1538 (2001).
Cheung, A.K. et al. 1,4-disubstituted pyridazine analogs there of and methods for treating SMN-deficiency–related conditions. United States Patent 8,729,263 (2014).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Lacoste, A., Berenshteyn, F. & Brivanlou, A.H. An efficient and reversible transposable system for gene delivery and lineage-specific differentiation in human embryonic stem cells. Cell Stem Cell 5, 332–342 (2009).
Chambers, S.M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Pruitt, K.D., Tatusova, T. & Maglott, D.R. NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 35, D61–D65 (2007).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Krainer, A.R., Maniatis, T., Ruskin, B. & Green, M.R. Normal and mutant human β-globin pre-mRNAs are faithfully and efficiently spliced in vitro. Cell 36, 993–1005 (1984).
Krainer, A.R. Pre-mRNA splicing by complementation with purified human U1, U2, U4/U6 and U5 snRNPs. Nucleic Acids Res. 16, 9415–9429 (1988).
Kastner, B. & Lührmann, R. Purification of U small nuclear ribonucleoprotein particles. Methods Mol. Biol. 118, 289–298 (1999).
Frostell-Karlsson, A. et al. Biosensor analysis of the interaction between immobilized human serum albumin and drug compounds for prediction of human serum albumin binding levels. J. Med. Chem. 43, 1986–1992 (2000).
Parisien, M. & Major, F. The MC-Fold and MC-Sym pipeline infers RNA structure from sequence data. Nature 452, 51–55 (2008).
Rueda, M., Totrov, M. & Abagyan, R. ALiBERO: evolving a team of complementary pocket conformations rather than a single leader. J. Chem. Inf. Model. 52, 2705–2714 (2012).
Abagyan, R. & Totrov, M. Biased probability Monte Carlo conformational searches and electrostatic calculations for peptides and proteins. J. Mol. Biol. 235, 983–1002 (1994).
Acknowledgements
The authors wish to acknowledge members of the Novartis Institutes for BioMedical Research (NIBR) Leadership Spinal Muscular Atrophy Advisory Board (J. Hastewell, J. Bell, K. Briner, P. Bouchard, E. Beckman and G. Kwei), NIBR Project Management (D. Silva and C. Gauthier) and NIBR Translational Medicine (R. Roubenoff) for their advice and contributions to the drug discovery efforts; J.R. Kerrigan, D. Glass, P. Manos, F. Harbinski, C. Mickanin, R.E.J. Beckwith, R. Sun, W. Broom, S.J. Luchanksy, L. Murphy, M. Schirle, J. Duca, R. Chopra and K. Clark for their contributions to experimental efforts and insights on the manuscript; S.J. Burden (NYU School of Medicine) for his gift of SMNΔ7 mouse myoblasts; K. Mineev and A.S. Arseniev for their assistance with NMR peak assignments; A. Abrams for his artwork in the schematic diagram; the SMA Foundation (K. Chen, D. Kobayashi, S. Paushkin and L. Eng) for their advice and contributions to the drug discovery efforts; Psychogenics (S. Ramboz and K. Cirillo); and PharmOptima (D. Decker, R. Poorman and P. Zaworski) for their contributions to in vivo studies.
Author information
Authors and Affiliations
Contributions
C.S., T.M.S. and R.S. performed the high-throughput screen. A.K.C., L.S., L.G.H. and N.A.D. performed the chemical synthesis. C.S., M.V.H., Y.S., C. Blaustein, F.B., A.L. and R.S. performed cell-based structure-activity experiments. M.V.H., Y.S., R.S., M.J., L.D., C. Bullock, M.M., W.F.D. and R.S. performed in vivo experiments. J.P., C.G.K., M.B., N.A.R., X.S., M.H., S.S., L.M., G.R. and R.S. performed the RNAseq experiments. J.P. and C.S. performed the chimera experiments. S.E.S., M.S. and J.R.T. performed the biochemistry and biophysics experiments. X.Z. and M.J.J.B. performed the NMR experiments. D.N.C. performed the computational modeling. L.G.H. and N.A.D. supervised the medicinal chemistry experiments. M.J., B.S.T., W.F.D. and R.S. provided intellectual input to the in vivo mouse biology experiments. B.S.T., J.A.P., D.C., M.C.F. and R.S. provided intellectual input to the overall drug discovery studies. G.A.M., J.A.P., V.E.M. and J.A.T. provided intellectual input to the mechanism of action studies. N.A.D. and R.S. directed the drug discovery experiments. J.P. and S.E.S. directed the mechanism-of-action experiments. J.P., S.E.S., J.A.T., N.A.D. and R.S. prepared the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1 and 2 and Supplementary Figures 1–16. (PDF 4533 kb)
Rights and permissions
About this article
Cite this article
Palacino, J., Swalley, S., Song, C. et al. SMN2 splice modulators enhance U1–pre-mRNA association and rescue SMA mice. Nat Chem Biol 11, 511–517 (2015). https://doi.org/10.1038/nchembio.1837
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1837
This article is cited by
-
SMN deficiency perturbs monoamine neurotransmitter metabolism in spinal muscular atrophy
Communications Biology (2023)
-
Human disease models in drug development
Nature Reviews Bioengineering (2023)
-
mRNA 3’UTR lengthening by alternative polyadenylation attenuates inflammatory responses and correlates with virulence of Influenza A virus
Nature Communications (2023)
-
Molecular basis of RNA-binding and autoregulation by the cancer-associated splicing factor RBM39
Nature Communications (2023)
-
The pyridazine heterocycle in molecular recognition and drug discovery
Medicinal Chemistry Research (2023)