Abstract
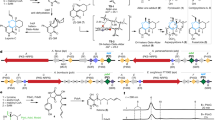
An improved understanding of enzymes' catalytic proficiency and stereoselectivity would further enable applications in chemistry, biocatalysis and industrial biotechnology. We use a chemical probe to dissect individual catalytic steps of enoyl-thioester reductases (Etrs), validating an active site tyrosine as the cryptic proton donor and explaining how it had eluded definitive identification. This information enabled the rational redesign of Etr, yielding mutants that create products with inverted stereochemistry at wild type–like turnover frequency.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Kiss, G., Celebi-Olcum, N., Moretti, R., Baker, D. & Houk, K.N. Angew. Chem. Int. Edn Engl. 52, 5700–5725 (2013).
Baker, D. Protein Sci. 19, 1817–1819 (2010).
Jiang, L. et al. Science 319, 1387–1391 (2008).
Siegel, J.B. et al. Science 329, 309–313 (2010).
Hedlund, J., Jornvall, H. & Persson, B. BMC Bioinformatics 11, 534 (2010).
Chen, Z.J. et al. J. Mol. Biol. 379, 830–844 (2008).
Zheng, J., Gay, D.C., Demeler, B., White, M.A. & Keatinge-Clay, A.T. Nat. Chem. Biol. 8, 615–621 (2012).
Airenne, T.T. et al. J. Mol. Biol. 327, 47–59 (2003).
Kwan, D.H. & Leadlay, P.F. ACS Chem. Biol. 5, 829–838 (2010).
Mesa, J. et al. Chem. Biol. Interact. 10.1016/j.cbi.2015.01.021 (22 January 2015).
Wu, Y.H. et al. Structure 16, 1714–1723 (2008).
Libby, R.D. & Mehl, R.A. Bioorg. Chem. 40, 57–66 (2012).
Rosenthal, R.G. et al. Nat. Chem. Biol. 10, 50–55 (2014).
Hamilton, G.A. in Progress in Bioorganic Chemistry (eds. Kaiser, E.T. and Kezdy, T.J.) 83–157 (Wiley Interscience, New York, 1971).
Bryan, P., Pantoliano, M.W., Quill, S.G., Hsiao, H.Y. & Poulos, T. Proc. Natl. Acad. Sci. USA 83, 3743–3745 (1986).
Fersht, A.R. et al. Nature 314, 235–238 (1985).
Kwan, D.H. et al. Chem. Biol. 15, 1231–1240 (2008).
Toogood, H.S. & Scrutton, N.S. Curr. Opin. Chem. Biol. 19, 107–115 (2014).
Zheng, J., Piasecki, S.K. & Keatinge-Clay, A.T. ACS Chem. Biol. 8, 1964–1971 (2013).
Rafi, S. et al. J. Biol. Chem. 281, 39285–39293 (2006).
Maier, T., Leibundgut, M. & Ban, N. Science 321, 1315–1322 (2008).
Erb, T.J. et al. Proc. Natl. Acad. Sci. USA 104, 10631–10636 (2007).
Erb, T.J., Brecht, V., Fuchs, G., Muller, M. & Alber, B.E. Proc. Natl. Acad. Sci. USA 106, 8871–8876 (2009).
Tabor, S. & Richardson, C.C. Proc. Natl. Acad. Sci. USA 82, 1074–1078 (1985).
Studier, F.W. Protein Expr. Purif. 41, 207–234 (2005).
Dawson, R.M.C. Data for Biochemical Research (Clarendon Press, Oxford, 1986).
Torkko, J.M. et al. J. Biol. Chem. 278, 41213–41220 (2003).
Kabsch, W. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Mccoy, A.J. et al. J. Appl. Crystallogr. 40, 658–674 (2007).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Painter, J. & Merritt, E.A. Acta Crystallogr. D Biol. Crystallogr. 62, 439–450 (2006).
Peyraud, R. et al. Proc. Natl. Acad. Sci. USA 106, 4846–4851 (2009).
Hoops, S. et al. Bioinformatics 22, 3067–3074 (2006).
Acknowledgements
This work was supported by the Swiss National Science Foundation (SNF-Ambizione program PZ00P3_136828/1; granted to T.J.E.).
Author information
Authors and Affiliations
Contributions
R.G.R., B.V. and T.J.E. conceived and designed all experiments, with the exception of the NMR experiments, which were designed together with M.-O.E., and the MS analyses, which were designed together with P.K. and J.A.V. NMR experiments were performed by R.G.R., B.V. and M.-O.E. MS experiments were performed by B.V., R.G.R. and P.K. B.V. prepared enzyme crystals of Etr1p and mutants, N.Q. and G.C. collected the diffraction data and interpreted the results. Enzyme kinetic assays and stopped-flow measurements, as well as purification of the intermediate, were performed by R.G.R. and B.V. R.G.R., B.V. and T.J.E. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–5 and Supplementary Figures 1–8 (PDF 1928 kb)
Rights and permissions
About this article
Cite this article
Rosenthal, R., Vögeli, B., Quade, N. et al. The use of ene adducts to study and engineer enoyl-thioester reductases. Nat Chem Biol 11, 398–400 (2015). https://doi.org/10.1038/nchembio.1794
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1794
This article is cited by
-
An engineered variant of MECR reductase reveals indispensability of long-chain acyl-ACPs for mitochondrial respiration
Nature Communications (2023)
-
Benzylmalonyl-CoA dehydrogenase, an enzyme involved in bacterial auxin degradation
Archives of Microbiology (2021)
-
A conserved threonine prevents self-intoxication of enoyl-thioester reductases
Nature Chemical Biology (2017)
-
Kovalente Zwischenprodukte in NAD(P)H-abhängigen Oxidoreduktasen
BIOspektrum (2016)