Abstract
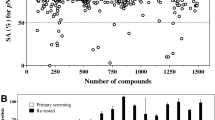
In bacteria, disulfide bonds confer stability on many proteins exported to the cell envelope or beyond. These proteins include numerous bacterial virulence factors, and thus bacterial enzymes that promote disulfide bond formation represent targets for compounds inhibiting bacterial virulence. Here, we describe a new target- and cell-based screening methodology for identifying compounds that inhibit the disulfide bond–forming enzymes Escherichia coli DsbB (EcDsbB) or Mycobacterium tuberculosis VKOR (MtbVKOR), which can replace EcDsbB, although the two are not homologs. Initial screening of 51,487 compounds yielded six specifically inhibiting EcDsbB. These compounds share a structural motif and do not inhibit MtbVKOR. A medicinal chemistry approach led us to select related compounds, some of which are much more effective DsbB inhibitors than those found in the screen. These compounds inhibit purified DsbB and prevent anaerobic growth of E. coli. Furthermore, these compounds inhibit all but one of the DsbBs of nine other Gram-negative pathogenic bacteria tested.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
19 February 2015
In the version of this article initially published online, the organism Mycobacterium tuberculosis was incorrectly named Mycoplasma tuberculosis. The error has been corrected for the print, PDF and HTML versions of this article.
References
Heras, B. et al. DSB proteins and bacterial pathogenicity. Nat. Rev. Microbiol. 7, 215–225 (2009).
Depuydt, M., Messens, J. & Collet, J.F. How proteins form disulfide bonds. Antioxid. Redox Signal. 15, 49–66 (2011).
Kadokura, H. & Beckwith, J. Mechanisms of oxidative protein folding in the bacterial cell envelope. Antioxid. Redox Signal. 13, 1231–1246 (2010).
Bardwell, J.C., McGovern, K. & Beckwith, J. Identification of a protein required for disulfide bond formation in vivo. Cell 67, 581–589 (1991).
Dutton, R.J., Boyd, D., Berkmen, M. & Beckwith, J. Bacterial species exhibit diversity in their mechanisms and capacity for protein disulfide bond formation. Proc. Natl. Acad. Sci. USA 105, 11933–11938 (2008).
Li, W. et al. Structure of a bacterial homologue of vitamin K epoxide reductase. Nature 463, 507–512 (2010).
Wang, X., Dutton, R.J., Beckwith, J. & Boyd, D. Membrane topology and mutational analysis of Mycobacterium tuberculosis VKOR, a protein involved in disulfide bond formation and a homologue of human vitamin K epoxide reductase. Antioxid. Redox Signal. 14, 1413–1420 (2011).
Premkumar, L. et al. Rv2969c, essential for optimal growth in Mycobacterium tuberculosis, is a DsbA-like enzyme that interacts with VKOR-derived peptides and has atypical features of DsbA-like disulfide oxidases. Acta Crystallogr. D Biol. Crystallogr. 69, 1981–1994 (2013).
Dutton, R.J. et al. Inhibition of bacterial disulfide bond formation by the anticoagulant warfarin. Proc. Natl. Acad. Sci. USA 107, 297–301 (2010).
Sassetti, C.M., Boyd, D.H. & Rubin, E.J. Genes required for mycobacterial growth defined by high density mutagenesis. Mol. Microbiol. 48, 77–84 (2003).
Froshauer, S., Green, G.N., Boyd, D., McGovern, K. & Beckwith, J. Genetic analysis of the membrane insertion and topology of MalF, a cytoplasmic membrane protein of Escherichia coli. J. Mol. Biol. 200, 501–511 (1988).
Tian, H., Boyd, D. & Beckwith, J. A mutant hunt for defects in membrane protein assembly yields mutations affecting the bacterial signal recognition particle and Sec machinery. Proc. Natl. Acad. Sci. USA 97, 4730–4735 (2000).
Tian, H. & Beckwith, J. Genetic screen yields mutations in genes encoding all known components of the Escherichia coli signal recognition particle pathway. J. Bacteriol. 184, 111–118 (2002).
Goldman, R.C. & Laughon, B.E. Discovery and validation of new antitubercular compounds as potential drug leads and probes. Tuberculosis (Edinb.) 89, 331–333 (2009).
Regeimbal, J. et al. Disulfide bond formation involves a quinhydrone-type charge-transfer complex. Proc. Natl. Acad. Sci. USA 100, 13779–13784 (2003).
Inaba, K. et al. DsbB elicits a red-shift of bound ubiquinone during the catalysis of DsbA oxidation. J. Biol. Chem. 279, 6761–6768 (2004).
Inaba, K., Takahashi, Y.H., Ito, K. & Hayashi, S. Critical role of a thiolate-quinone charge transfer complex and its adduct form in de novo disulfide bond generation by DsbB. Proc. Natl. Acad. Sci. USA 103, 287–292 (2006).
Grimshaw, J.P. et al. DsbL and DsbI form a specific dithiol oxidase system for periplasmic arylsulfate sulfotransferase in uropathogenic Escherichia coli. J. Mol. Biol. 380, 667–680 (2008).
Lin, D., Kim, B. & Slauch, J.M. DsbL and DsbI contribute to periplasmic disulfide bond formation in Salmonella enterica serovar Typhimurium. Microbiology 155, 4014–4024 (2009).
Harvey, H., Habash, M., Aidoo, F. & Burrows, L.L. Single-residue changes in the C-terminal disulfide-bonded loop of the Pseudomonas aeruginosa type IV pilin influence pilus assembly and twitching motility. J. Bacteriol. 191, 6513–6524 (2009).
Ha, U.H., Wang, Y. & Jin, S. DsbA of Pseudomonas aeruginosa is essential for multiple virulence factors. Infect. Immun. 71, 1590–1595 (2003).
Kim, S.H., Park, S.Y., Heo, Y.J. & Cho, Y.H. Drosophila melanogaster-based screening for multihost virulence factors of Pseudomonas aeruginosa PA14 and identification of a virulence-attenuating factor, HudA. Infect. Immun. 76, 4152–4162 (2008).
Arts, I.S. et al. Dissecting the machinery that introduces disulfide bonds in Pseudomonas aeruginosa. MBio 4, e00912–e00913 (2013).
Hatahet, F., Boyd, D. & Beckwith, J. Disulfide bond formation in prokaryotes: History, diversity and design. Biochim. Biophys. Acta 1844, 1402–1414 (2014).
Hatahet, F. & Ruddock, L.W. Topological plasticity of enzymes involved in disulfide bond formation allows catalysis in either the periplasm or the cytoplasm. J. Mol. Biol. 425, 3268–3276 (2013).
Li, T. et al. Identification of the gene for vitamin K epoxide reductase. Nature 427, 541–544 (2004).
Westhofen, P. et al. Human vitamin K 2,3-epoxide reductase complex subunit 1-like 1 (VKORC1L1) mediates vitamin K-dependent intracellular antioxidant function. J. Biol. Chem. 286, 15085–15094 (2011).
Früh, V. et al. Application of fragment-based drug discovery to membrane proteins: identification of ligands of the integral membrane enzyme DsbB. Chem. Biol. 17, 881–891 (2010).
Clatworthy, A.E., Pierson, E. & Hung, D.T. Targeting virulence: a new paradigm for antimicrobial therapy. Nat. Chem. Biol. 3, 541–548 (2007).
O'Loughlin, C.T. et al. A quorum-sensing inhibitor blocks Pseudomonas aeruginosa virulence and biofilm formation. Proc. Natl. Acad. Sci. USA 110, 17981–17986 (2013).
Hassett, D.J. et al. Pseudomonas aeruginosa hypoxic or anaerobic biofilm infections within cystic fibrosis airways. Trends Microbiol. 17, 130–138 (2009).
Kolpen, M. et al. Nitrous oxide production in sputum from cystic fibrosis patients with chronic Pseudomonas aeruginosa lung infection. PLoS ONE 9, e84353 (2014).
Singh, J., Petter, R.C., Baillie, T.A. & Whitty, A. The resurgence of covalent drugs. Nat. Rev. Drug Discov. 10, 307–317 (2011).
Mah, R., Thomas, J.R. & Shafer, C.M. Drug discovery considerations in the development of covalent inhibitors. Bioorg. Med. Chem. Lett. 24, 33–39 (2014).
Boyd, D., Weiss, D.S., Chen, J.C. & Beckwith, J. Towards single-copy gene expression systems making gene cloning physiologically relevant: lambda InCh, a simple Escherichia coli plasmid-chromosome shuttle system. J. Bacteriol. 182, 842–847 (2000).
Haldimann, A. & Wanner, B.L. Conditional-replication, integration, excision, and retrieval plasmid-host systems for gene structure-function studies of bacteria. J. Bacteriol. 183, 6384–6393 (2001).
Larsen, M.H., Biermann, K. & Jacobs, W.R. Jr. Laboratory maintenance of Mycobacterium tuberculosis. Curr. Protoc. Microbiol. Chapter 10, Unit 10A.1 (2007).
Chng, S.S. et al. Overexpression of the rhodanese PspE, a single cysteine-containing protein, restores disulphide bond formation to an Escherichia coli strain lacking DsbA. Mol. Microbiol. 85, 996–1006 (2012).
Kadokura, H., Bader, M., Tian, H., Bardwell, J.C. & Beckwith, J. Roles of a conserved arginine residue of DsbB in linking protein disulfide-bond-formation pathway to the respiratory chain of Escherichia coli. Proc. Natl. Acad. Sci. USA 97, 10884–10889 (2000).
Weiss, D.S., Chen, J.C., Ghigo, J.M., Boyd, D. & Beckwith, J. Localization of FtsI (PBP3) to the septal ring requires its membrane anchor, the Z ring, FtsA, FtsQ, and FtsL. J. Bacteriol. 181, 508–520 (1999).
Thibodeau, S.A., Fang, R. & Joung, J.K. High-throughput beta-galactosidase assay for bacterial cell-based reporter systems. Biotechniques 36, 410–415 (2004).
Semmler, A.B., Whitchurch, C.B. & Mattick, J.S. A re-examination of twitching motility in Pseudomonas aeruginosa. Microbiology 145, 2863–2873 (1999).
Acknowledgements
We thank the ICCB-Longwood screening facility for access to their compound libraries, equipment and, screening supplies, and the staff, including S. Chiang, K. Lee, S. Rudnicki and D. Flood, for helpful advice and data handling. We thank R. Goldman of the US National Institute of Allergy and Infectious disease for the collection of M. tuberculosis growth inhibitors. We thank R. Tomaino of the Taplin Mass Spectrometry Core Facility at Harvard Medical School for his assistance in sample analysis. This work was supported by US National Institute of General Medical Sciences grants GMO41883 (to J.B. and D.B.), Harvard Catalyst Pilot Grant Harvard #149734 (to J.B.), US National Institutes of Health (NIH) Grant 5-UL1RR02568-01 (to J.B.), Blavatnik Biomedical Accelerator at Harvard University (to J.B.), NIH Grant 3P01-A1AI074805-04S1 Subaward R01638 (to E.J.R.), NIH Grant PO1-HL087203 (to B.F.), NIH Grant RO1-HL092125 (to B.C.F.) and New England Regional Center of Excellence for Biodefense and Emerging Infectious Diseases (NERCE-BEID) Grant U54-AI057159. C.L. was partially supported by a Consejo Nacional de Ciencia y Tecnología (CONACYT) postdoctoral fellowship. J.L.B. was supported by T32-HL07917 Grant (to B.F.). B.M.M. was supported by a Ruth L. Kirschstein National Research Service Award. M.E. was supported by a New England BioLabs Grant. A.M. was supported by a Harvard School of Public Health Yerby postdoctoral fellowship. J.B. is an American Cancer Society Professor.
Author information
Authors and Affiliations
Contributions
M.E. developed the agar assay. J.L.B., M.E., R.D., H.A., N.K. and D.B. performed the HTS. J.L.B. performed cherry-pick tests. J.L.B. and C.L. performed inhibitor retests. C.L. performed substructure analysis, in vivo DsbA and DsbB inhibition and other Gram-negative bacteria assays. F.H. performed enzyme kinetics and in vitro analysis. B.M.M. performed anaerobic and M. smegmatis growth assays. L.B., B.C.F. and B.F. performed in vitro mice VKOR assays. A.M., S.M. and E.J.R. performed M. tuberculosis growth assays. C.L., J.L.B., F.H., B.M.M., D.B. and J.B. analyzed and discussed the data. C.L. and J.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–5 and Supplementary Tables 1–5. (PDF 11836 kb)
Rights and permissions
About this article
Cite this article
Landeta, C., Blazyk, J., Hatahet, F. et al. Compounds targeting disulfide bond forming enzyme DsbB of Gram-negative bacteria. Nat Chem Biol 11, 292–298 (2015). https://doi.org/10.1038/nchembio.1752
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1752
This article is cited by
-
A high-throughput cell-based assay pipeline for the preclinical development of bacterial DsbA inhibitors as antivirulence therapeutics
Scientific Reports (2021)
-
Disulfide bond formation in prokaryotes
Nature Microbiology (2018)
-
Polysulfurating reagent design for unsymmetrical polysulfide construction
Nature Communications (2018)
-
Sulfur–Sulfur Bond Construction
Topics in Current Chemistry (2018)
-
Bacterial thiol oxidoreductases — from basic research to new antibacterial strategies
Applied Microbiology and Biotechnology (2017)