Abstract
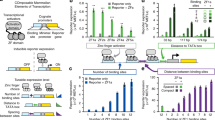
An important goal of synthetic biology is the rational design and predictable implementation of synthetic gene circuits using standardized and interchangeable parts. However, engineering of complex circuits in mammalian cells is currently limited by the availability of well-characterized and orthogonal transcriptional repressors. Here, we introduce a library of 26 reversible transcription activator–like effector repressors (TALERs) that bind newly designed hybrid promoters and exert transcriptional repression through steric hindrance of key transcriptional initiation elements. We demonstrate that using the input-output transfer curves of our TALERs enables accurate prediction of the behavior of modularly assembled TALER cascade and switch circuits. We also show that TALER switches using feedback regulation exhibit improved accuracy for microRNA-based HeLa cancer cell classification versus HEK293 cells. Our TALER library is a valuable toolkit for modular engineering of synthetic circuits, enabling programmable manipulation of mammalian cells and helping elucidate design principles of coupled transcriptional and microRNA-mediated post-transcriptional regulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Basu, S., Mehreja, R., Thiberge, S., Chen, M.T. & Weiss, R. Spatiotemporal control of gene expression with pulse-generating networks. Proc. Natl. Acad. Sci. USA 101, 6355–6360 (2004).
Tigges, M., Marquez-Lago, T.T., Stelling, J. & Fussenegger, M. A tunable synthetic mammalian oscillator. Nature 457, 309–312 (2009).
Prindle, A. et al. A sensing array of radically coupled genetic ′biopixels′. Nature 481, 39–44 (2012).
Gardner, T.S., Cantor, C.R. & Collins, J.J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Kramer, B.P. et al. An engineered epigenetic transgene switch in mammalian cells. Nat. Biotechnol. 22, 867–870 (2004).
Hooshangi, S., Thiberge, S. & Weiss, R. Ultrasensitivity and noise propagation in a synthetic transcriptional cascade. Proc. Natl. Acad. Sci. USA 102, 3581–3586 (2005).
Friedland, A.E. et al. Synthetic gene networks that count. Science 324, 1199–1202 (2009).
Galloway, K.E., Franco, E. & Smolke, C.D. Dynamically reshaping signaling networks to program cell fate via genetic controllers. Science 341, 1235005 (2013).
Basu, S., Gerchman, Y., Collins, C.H., Arnold, F.H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005).
Liu, C. et al. Sequential establishment of stripe patterns in an expanding cell population. Science 334, 238–241 (2011).
Bacchus, W. et al. Synthetic two-way communication between mammalian cells. Nat. Biotechnol. 30, 991–996 (2012).
Bleris, L. et al. Synthetic incoherent feedforward circuits show adaptation to the amount of their genetic template. Mol. Syst. Biol. 7, 519 (2011).
Wei, P. et al. Bacterial virulence proteins as tools to rewire kinase pathways in yeast and immune cells. Nature 488, 384–388 (2012).
Chau, A.H., Walter, J.M., Gerardin, J., Tang, C. & Lim, W.A. Designing synthetic regulatory networks capable of self-organizing cell polarization. Cell 151, 320–332 (2012).
Tamsir, A., Tabor, J.J. & Voigt, C.A. Robust multicellular computing using genetically encoded NOR gates and chemical ′wires′. Nature 469, 212–215 (2011).
Daniel, R., Rubens, J.R., Sarpeshkar, R. & Lu, T.K. Synthetic analog computation in living cells. Nature 497, 619–623 (2013).
Lienert, F. et al. Two- and three-input TALE-based AND logic computation in embryonic stem cells. Nucleic Acids Res. 41, 9967–9975 (2013).
Xie, Z., Wroblewska, L., Prochazka, L., Weiss, R. & Benenson, Y. Multi-input RNAi-based logic circuit for identification of specific cancer cells. Science 333, 1307–1311 (2011).
Ausländer, S., Auslander, D., Muller, M., Wieland, M. & Fussenegger, M. Programmable single-cell mammalian biocomputers. Nature 487, 123–127 (2012).
Bonnet, J., Yin, P., Ortiz, M.E., Subsoontorn, P. & Endy, D. Amplifying genetic logic gates. Science 340, 599–603 (2013).
Gaber, R. et al. Designable DNA-binding domains enable construction of logic circuits in mammalian cells. Nat. Chem. Biol. 10, 203–208 (2014).
Martin, V.J., Pitera, D.J., Withers, S.T., Newman, J.D. & Keasling, J.D. Engineering a mevalonate pathway in Escherichia coli for production of terpenoids. Nat. Biotechnol. 21, 796–802 (2003).
Lohmueller, J.J., Armel, T.Z. & Silver, P.A. A tunable zinc finger–based framework for Boolean logic computation in mammalian cells. Nucleic Acids Res. 40, 5180–5187 (2012).
Khalil, A.S. et al. A synthetic biology framework for programming eukaryotic transcription functions. Cell 150, 647–658 (2012).
Cong, L., Zhou, R., Kuo, Y.C., Cunniff, M. & Zhang, F. Comprehensive interrogation of natural TALE DNA-binding modules and transcriptional repressor domains. Nat. Commun. 3, 968 (2012).
Garg, A., Lohmueller, J.J., Silver, P.A. & Armel, T.Z. Engineering synthetic TAL effectors with orthogonal target sites. Nucleic Acids Res. 40, 7584–7595 (2012).
Li, Y., Moore, R., Guinn, M. & Bleris, L. Transcription activator-like effector hybrids for conditional control and rewiring of chromosomal transgene expression. Sci. Rep. 2, 897 (2012).
Qi, L.S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Gilbert, L.A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Nan, X. et al. Transcriptional repression by the methyl-CpG–binding protein MeCP2 involves a histone deacetylase complex. Nature 393, 386–389 (1998).
Rinaudo, K. et al. A universal RNAi-based logic evaluator that operates in mammalian cells. Nat. Biotechnol. 25, 795–801 (2007).
Zhang, F. et al. Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nat. Biotechnol. 29, 149–153 (2011).
Cermak, T. et al. Efficient design and assembly of custom TALEN and other TAL effector–based constructs for DNA targeting. Nucleic Acids Res. 39, e82 (2011).
Müller, J., Oehler, S. & Muller-Hill, B. Repression of lac promoter as a function of distance, phase and quality of an auxiliary lac operator. J. Mol. Biol. 257, 21–29 (1996).
Guye, P., Li, Y., Wroblewska, L., Duportet, X. & Weiss, R. Rapid, modular and reliable construction of complex mammalian gene circuits. Nucleic Acids Res. 41, e156 (2013).
Balázsi, G., van Oudenaarden, A. & Collins, J.J. Cellular decision making and biological noise: from microbes to mammals. Cell 144, 910–925 (2011).
Wang, G., Zhu, X., Hood, L. & Ao, P. From Phage lambda to human cancer: endogenous molecular-cellular network hypothesis. Quantitative Biology 1, 32–49 (2013).
Szymczak, A.L. et al. Correction of multi-gene deficiency in vivo using a single ′self-cleaving′ 2A peptide-based retroviral vector. Nat. Biotechnol. 22, 589–594 (2004).
Filonov, G.S. et al. Bright and stable near-infrared fluorescent protein for in vivo imaging. Nat. Biotechnol. 29, 757–761 (2011).
Landgraf, P. et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129, 1401–1414 (2007).
Boch, J. et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326, 1509–1512 (2009).
Liang, J., Chao, R., Abil, Z., Bao, Z. & Zhao, H. FairyTALE: a high-throughput TAL effector synthesis platform. ACS Synth. Biol. 3, 67–73 (2014).
Tsang, J., Zhu, J. & van Oudenaarden, A. MicroRNA-mediated feedback and feedforward loops are recurrent network motifs in mammals. Mol. Cell 26, 753–767 (2007).
Beisel, C.L., Bayer, T.S., Hoff, K.G. & Smolke, C.D. Model-guided design of ligand-regulated RNAi for programmable control of gene expression. Mol. Syst. Biol. 4, 224 (2008).
Xie, Z., Liu, S.J., Bleris, L. & Benenson, Y. Logic integration of mRNA signals by an RNAi-based molecular computer. Nucleic Acids Res. 38, 2692–2701 (2010).
Leisner, M., Bleris, L., Lohmueller, J., Xie, Z. & Benenson, Y. Rationally designed logic integration of regulatory signals in mammalian cells. Nat. Nanotechnol. 5, 666–670 (2010).
Beisel, C.L., Chen, Y.Y., Culler, S.J., Hoff, K.G. & Smolke, C.D. Design of small molecule–responsive microRNAs based on structural requirements for Drosha processing. Nucleic Acids Res. 39, 2981–2994 (2011).
Kashida, S., Inoue, T. & Saito, H. Three-dimensionally designed protein-responsive RNA devices for cell signaling regulation. Nucleic Acids Res. 40, 9369–9378 (2012).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Acknowledgements
We thank members of the Xie lab and the Weiss lab for helpful discussions. We thank H. Huang and T. Wang for technical support. We thank C. Le and F. Zhang (both at Department of Biological Engineering, Massachusetts Institute of Technology) for providing TALE1 and TALE2 as gifts. We thank J. Beal and M.Q. Zhang for insightful discussions. The research is supported by Beijing Natural Science Foundation (5152013 to Z.X.), National Key Basic Research Program of China (2014CB745200 to Z.X.), Junior '1,000 Plan' Program (to Z.X.), Cross-strait Tsinghua University Joint Grant (2012THZ0 to Z.X.) and US National Institutes of Health Grants (5R01CA155320-04, P50GM098792 and 1R01CA173712-01 to R.W.).
Author information
Authors and Affiliations
Contributions
Z.X., R.W. and Y.L. conceived the ideas implemented in this work. Y.L. and Z.X. performed TALER promoter optimization experiments. Y.L. constructed the TALER library and circuits for stable cell lines and performed orthogonality analysis and initial experiments for transfer function curves, TALER switches and stable cell lines. Y.J. performed experiments for transfer function curves, color models, TALER switches and cell classification. H.C. performed initial experiments for TALER switches and cell classification. W.L. performed experiments for stable cell lines and cell classification. Z.L. performed initial tests for TALE transfer function curves. Z.X., Y.L., W.L. and R.W. performed data analysis. Z.X. and R.W. supervised the project. Z.X., Y.L. and R.W. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
Three pending patent applications recently submitted to Sate Intellectual Property Office of P.R. China (application nos. 201410341963.8, 201410342334.7 and 201410342350.6) by the authors are related to TALER circuit engineering described in this manuscript.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–11 and Supplementary Tables 1–4. (PDF 3780 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Jiang, Y., Chen, H. et al. Modular construction of mammalian gene circuits using TALE transcriptional repressors. Nat Chem Biol 11, 207–213 (2015). https://doi.org/10.1038/nchembio.1736
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1736
This article is cited by
-
PERSIST platform provides programmable RNA regulation using CRISPR endoRNases
Nature Communications (2022)
-
Quantitative characterization of recombinase-based digitizer circuits enables predictable amplification of biological signals
Communications Biology (2021)
-
Designer membraneless organelles sequester native factors for control of cell behavior
Nature Chemical Biology (2021)
-
The COMET toolkit for composing customizable genetic programs in mammalian cells
Nature Communications (2020)
-
An endoribonuclease-based feedforward controller for decoupling resource-limited genetic modules in mammalian cells
Nature Communications (2020)