Abstract
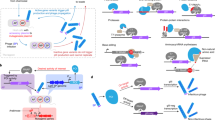
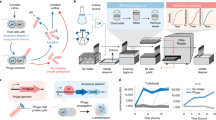
Phage-assisted continuous evolution (PACE) uses a modified filamentous bacteriophage life cycle to substantially accelerate laboratory evolution experiments. In this work, we expand the scope and capabilities of the PACE method with two key advances that enable the evolution of biomolecules with radically altered or highly specific new activities. First, we implemented small molecule–controlled modulation of selection stringency that enables otherwise inaccessible activities to be evolved directly from inactive starting libraries through a period of evolutionary drift. Second, we developed a general negative selection that enables continuous counterselection against undesired activities. We integrated these developments to continuously evolve mutant T7 RNA polymerase enzymes with ∼10,000-fold altered, rather than merely broadened, substrate specificities during a single three-day PACE experiment. The evolved enzymes exhibit specificity for their target substrate that exceeds that of wild-type RNA polymerases for their cognate substrates while maintaining wild type–like levels of activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Esvelt, K.M., Carlson, J.C. & Liu, D.R. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011).
Nelson, F.K., Friedman, S.M. & Smith, G.P. Filamentous phage DNA cloning vectors: a noninfective mutant with a nonpolar deletion in gene III. Virology 108, 338–350 (1981).
Calendar, R. The Bacteriophages 1st edn., 376–455 (Oxford Univ. Press, 2006).
Sergeeva, A., Kolonin, M.G., Molldrem, J.J., Pasqualini, R. & Arap, W. Display technologies: application for the discovery of drug and gene delivery agents. Adv. Drug Deliv. Rev. 58, 1622–1654 (2006).
Yuan, L., Kurek, I., English, J. & Keenan, R. Laboratory-directed protein evolution. Microbiol. Mol. Biol. Rev. 69, 373–392 (2005).
Leconte, A.M. et al. A population-based experimental model for protein evolution: effects of mutation rate and selection stringency on evolutionary outcomes. Biochemistry 52, 1490–1499 (2013).
Dickinson, B.C., Leconte, A.M., Allen, B., Esvelt, K.M. & Liu, D.R. Experimental interrogation of the path dependence and stochasticity of protein evolution using phage-assisted continuous evolution. Proc. Natl. Acad. Sci. USA 110, 9007–9012 (2013).
Doyon, J.B., Pattanayak, V., Meyer, C.B. & Liu, D.R. Directed evolution and substrate specificity profile of homing endonuclease I-SceI. J. Am. Chem. Soc. 128, 2477–2484 (2006).
Tracewell, C.A. & Arnold, F.H. Directed enzyme evolution: climbing fitness peaks one amino acid at a time. Curr. Opin. Chem. Biol. 13, 3–9 (2009).
Boeke, J.D., Model, P. & Zinder, N.D. Effects of bacteriophage f1 gene III protein on the host cell membrane. Mol. Gen. Genet. 186, 185–192 (1982).
Brissette, J.L., Weiner, L., Ripmaster, T.L. & Model, P. Characterization and sequence of the Escherichia coli stress-induced psp operon. J. Mol. Biol. 220, 35–48 (1991).
Brissette, J.L., Russel, M., Weiner, L. & Model, P. Phage shock protein, a stress protein of Escherichia coli. Proc. Natl. Acad. Sci. USA 87, 862–866 (1990).
Rakonjac, J., Jovanovic, G. & Model, P. Filamentous phage infection-mediated gene expression: construction and propagation of the gIII deletion mutant helper phage R408d3. Gene 198, 99–103 (1997).
Bonner, G., Lafer, E.M. & Sousa, R. Characterization of a set of T7 RNA polymerase active site mutants. J. Biol. Chem. 269, 25120–25128 (1994).
Rakonjac, J., Feng, J. & Model, P. Filamentous phage are released from the bacterial membrane by a two-step mechanism involving a short C-terminal fragment of pIII. J. Mol. Biol. 289, 1253–1265 (1999).
Bennett, N.J. & Rakonjac, J. Unlocking of the filamentous bacteriophage virion during infection is mediated by the C domain of pIII. J. Mol. Biol. 356, 266–273 (2006).
Grant, R.A., Lin, T.C., Konigsberg, W. & Webster, R.E. Structure of the filamentous bacteriophage fl. Location of the A, C, and D minor coat proteins. J. Biol. Chem. 256, 539–546 (1981).
Lopez, J. & Webster, R.E. Minor coat protein composition and location of the A protein in bacteriophage f1 spheroids and I-forms. J. Virol. 42, 1099–1107 (1982).
Lynch, S.A. & Gallivan, J.P. A flow cytometry–based screen for synthetic riboswitches. Nucleic Acids Res. 37, 184–192 (2009).
Raskin, C.A., Diaz, G., Joho, K. & McAllister, W.T. Substitution of a single bacteriophage T3 residue in bacteriophage T7 RNA polymerase at position 748 results in a switch in promoter specificity. J. Mol. Biol. 228, 506–515 (1992).
Cheetham, G.M., Jeruzalmi, D. & Steitz, T.A. Structural basis for initiation of transcription from an RNA polymerase–promoter complex. Nature 399, 80–83 (1999).
Imburgio, D., Rong, M., Ma, K. & McAllister, W.T. Studies of promoter recognition and start site selection by T7 RNA polymerase using a comprehensive collection of promoter variants. Biochemistry 39, 10419–10430 (2000).
Datsenko, K.A. & Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640–6645 (2000).
Dove, S.L. & Hochschild, A. Conversion of the ω subunit of Escherichia coli RNA polymerase into a transcriptional activator or an activation target. Genes Dev. 12, 745–754 (1998).
Xi, L., Cho, K.W. & Tu, S.C. Cloning and nucleotide sequences of lux genes and characterization of luciferase of Xenorhabdus luminescens from a human wound. J. Bacteriol. 173, 1399–1405 (1991).
Hammar, M., Arnqvist, A., Bian, Z., Olsen, A. & Normark, S. Expression of two csg operons is required for production of fibronectin- and Congo Red–binding curli polymers in Escherichia coli K-12. Mol. Microbiol. 18, 661–670 (1995).
Danese, P.N., Pratt, L.A., Dove, S.L. & Kolter, R. The outer membrane protein, antigen 43, mediates cell-to-cell interactions within Escherichia coli biofilms. Mol. Microbiol. 37, 424–432 (2000).
Wang, X., Preston, J.F. III & Romeo, T. The pgaABCD locus of Escherichia coli promotes the synthesis of a polysaccharide adhesin required for biofilm formation. J. Bacteriol. 186, 2724–2734 (2004).
Khlebnikov, A., Datsenko, K.A., Skaug, T., Wanner, B.L. & Keasling, J.D. Homogeneous expression of the PBAD promoter in Escherichia coli by constitutive expression of the low-affinity high-capacity AraE transporter. Microbiology 147, 3241–3247 (2001).
Acknowledgements
The authors thank K. Esvelt, D. Thompson, B. Dorr and K. Davis for helpful discussions. This research was supported by DARPA HR0011-11-2-0003, DARPA N66001-12-C-4207 and the Howard Hughes Medical Institute. J.C.C. was supported by the Harvard Chemical Biology Graduate Program. A.H.B. was supported by a National Science Foundation Graduate Research Fellowship. D.A.G.-N. was supported by the Harvard Biophysics Graduate Program.
Author information
Authors and Affiliations
Contributions
J.C.C. designed the research, prepared materials and performed experiments. A.H.B. prepared materials and performed experiments. D.A.G.-N. designed and constructed the real-time luminescence monitor. D.R.L. designed and supervised the research. All of the authors contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have filed a provisional patent application on PACE and related improvements.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–5 and Supplementary Table 1. (PDF 5132 kb)
Rights and permissions
About this article
Cite this article
Carlson, J., Badran, A., Guggiana-Nilo, D. et al. Negative selection and stringency modulation in phage-assisted continuous evolution. Nat Chem Biol 10, 216–222 (2014). https://doi.org/10.1038/nchembio.1453
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1453
This article is cited by
-
High-throughput continuous evolution of compact Cas9 variants targeting single-nucleotide-pyrimidine PAMs
Nature Biotechnology (2023)
-
Mobile CRISPR-Cas9 based anti-phage system in E. coli
Frontiers of Chemical Science and Engineering (2022)
-
Orthogonal translation enables heterologous ribosome engineering in E. coli
Nature Communications (2021)
-
Engineered dual selection for directed evolution of SpCas9 PAM specificity
Nature Communications (2021)
-
Multiplex suppression of four quadruplet codons via tRNA directed evolution
Nature Communications (2021)