Abstract
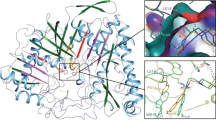
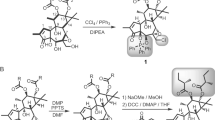
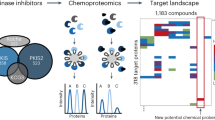
Phospholipase D (PLD) is an essential enzyme responsible for the production of the lipid second messenger phosphatidic acid. Phosphatidic acid participates in both G protein–coupled receptor and receptor tyrosine kinase signal transduction networks. The lack of potent and isoform-selective inhibitors has limited progress in defining the cellular roles of PLD. We used a diversity-oriented synthetic approach and developed a library of PLD inhibitors with considerable pharmacological characterization. Here we report the rigorous evaluation of that library, which contains highly potent inhibitors, including the first isoform-selective PLD inhibitors. Specific members of this series inhibit isoforms with >100-fold selectivity both in vitro and in cells. A subset of inhibitors was shown to block invasiveness in metastatic breast cancer models. These findings demonstrate the power of diversity-oriented synthesis combined with biochemical assays and mass spectrometric lipid profiling of cellular responses to develop the first isoform-selective PLD inhibitors—a new class of antimetastatic agents.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Ponting, C.P. & Kerr, I.D. A novel family of phospholipase D homologues that includes phospholipid synthases and putative endonucleases: identification of duplicated repeats and potential active site residues. Protein Sci. 5, 914–922 (1996).
Xie, Z., Ho, W.T. & Exton, J.H. Association of N- and C-terminal domains of phospholipase D is required for catalytic activity. J. Biol. Chem. 273, 34679–34682 (1998).
Brown, H.A. et al. ADP-ribosylation factor, a small GTP-dependent regulatory protein, stimulates phospholipase D activity. Cell 75, 1137–1144 (1993).
Hammond, S.M. et al. Human ADP-ribosylation factor-activated phosphatidylcholine-specific phospholipase D defines a new and highly conserved gene family. J. Biol. Chem. 270, 29640–29643 (1995).
Park, S.K. et al. Cloning and characterization of phospholipase D from rat brain. J. Biol. Chem. 272, 29263–29271 (1997).
Exton, J.H. Regulation of phospholipase D. FEBS Lett. 531, 58–61 (2002).
Hancock, J.F. PA promoted to manager. Nat. Cell Biol. 9, 615–617 (2007).
Wymann, M.P. & Schneiter, R. Lipid signalling in disease. Nat. Rev. Mol. Cell Biol. 9, 162–176 (2008).
Noh, D.Y. et al. Overexpression of phospholipase D1 in human breast cancer tissues. Cancer Lett. 161, 207–214 (2000).
Uchida, N., Okamura, S., Nagamachi, Y. & Yamashita, S. Increased phospholipase D activity in human breast cancer. J. Cancer Res. Clin. Oncol. 123, 280–285 (1997).
Zhao, Y. et al. Increased activity and intranuclear expression of phospholipase D2 in human renal cancer. Biochem. Biophys. Res. Commun. 278, 140–143 (2000).
Uchida, N., Okamura, S. & Kuwano, H. Phospholipase D activity in human gastric carcinoma. Anticancer Res. 19, 671–675 (1999).
Yamada, Y. et al. Association of a polymorphism of the phospholipase D2 gene with the prevalence of colorectal cancer. J. Mol. Med. 81, 126–131 (2003).
Min, D.S. et al. Neoplastic transformation and tumorigenesis associated with overexpression of phospholipase D isozymes in cultured murine fibroblasts. Carcinogenesis 22, 1641–1647 (2001).
Buchanan, F.G. et al. Requirement of phospholipase D1 activity in H-RasV12-induced transformation. Proc. Natl. Acad. Sci. USA 102, 1638–1642 (2005).
Bocckino, S.B., Wilson, P.B. & Exton, J.H. Ca2+-mobilizing hormones elicit phosphatidylethanol accumulation via phospholipase D activation. FEBS Lett. 225, 201–204 (1987).
Ktistakis, N.T. et al. Evidence that phospholipase D mediates ADP ribosylation factor-dependent formation of Golgi coated vesicles. J. Cell Biol. 134, 295–306 (1996).
Tou, J.S. & Urbizo, C. Diethylstilbestrol inhibits phospholipase D activity and degranulation by stimulated human neutrophils. Steroids 73, 216–221 (2008).
Tou, J. & Urbizo, C. Resveratrol inhibits the formation of phosphatidic acid and diglyceride in chemotactic peptide- or phorbol ester-stimulated human neutrophils. Cell. Signal. 13, 191–197 (2001).
Garcia, A. et al. Honokiol suppresses survival signals mediated by Ras-dependent phopholipase D activity in human cancer cells. Clin. Cancer Res. 14, 4267–4274 (2008).
Puar, M.S., Barrabee, E., Hallade, M. & Patel, M. Sch 420789: a novel fungal metabolite with phospholipase D inhibitory activity. J. Antibiot. (Tokyo) 53, 837–838 (2000).
McDonald, L.A. et al. 07H239-A, a new cytotoxic eremophilane sesquiterpene from the marine-derived Xylariaceous fungus LL-07H239. J. Nat. Prod. 67, 1565–1567 (2004).
Levy, B.D. et al. Novel polyisoprenyl phosphates block phospholipase D and human neutrophil activation in vitro and murine peritoneal inflammation in vivo. Br. J. Pharmacol. 146, 344–351 (2005).
Eisen, S.F. & Brown, H.A. Selective estrogen receptor (ER) modulators differentially regulate phospholipase D catalytic activity in ER-negative breast cancer cells. Mol. Pharmacol. 62, 911–920 (2002).
Monovich, L. et al. Optimization of halopemide for phospholipase D2 inhibition. Bioorg. Med. Chem. Lett. 17, 2310–2311 (2007).
Brown, H.A. et al. Biochemical analysis of phospholipase D. Methods Enzymol. 434, 49–87 (2007).
Henage, L.G., Exton, J.H. & Brown, H.A. Kinetic analysis of a mammalian phospholipase D: allosteric modulation by monomeric GTPases, protein kinase C, and polyphosphoinositides. J. Biol. Chem. 281, 3408–3417 (2006).
Yang, H. & Roberts, M.F. Phosphohydrolase and transphosphatidylation reactions of two Streptomyces phospholipase D enzymes: covalent versus noncovalent catalysis. Protein Sci. 12, 2087–2098 (2003).
Billah, M.M. et al. Regulation of phospholipase D in HL-60 granulocytes. Activation by phorbol esters, diglyceride, and calcium ionophore via protein kinase-independent mechanisms. J. Biol. Chem. 264, 9069–9076 (1989).
Colley, W.C. et al. Phospholipase D2, a distinct phospholipase D isoform with novel regulatory properties that provokes cytoskeletal reorganization. Curr. Biol. 7, 191–201 (1997).
Aguirre Ghiso, J.A. et al. A phospholipase D and protein kinase C inhibitor blocks the spreading of murine mammary adenocarcinoma cells altering f-actin and β1-integrin point contact distribution. Int. J. Cancer 71, 881–890 (1997).
Shen, Y., Zheng, Y. & Foster, D.A. Phospholipase D2 stimulates cell protrusion in v-Src-transformed cells. Biochem. Biophys. Res. Commun. 293, 201–206 (2002).
Imamura, F. et al. Induction of in vitro tumor cell invasion of cellular monolayers by lysophosphatidic acid or phospholipase D. Biochem. Biophys. Res. Commun. 193, 497–503 (1993).
Pai, J.K., Frank, E.A., Blood, C. & Chu, M. Novel ketoepoxides block phospholipase D activation and tumor cell invasion. Anticancer Drug Des. 9, 363–372 (1994).
Guy, C.T., Cardiff, R.D. & Muller, W.J. Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol. Cell. Biol. 12, 954–961 (1992).
Bi, K., Roth, M.G. & Ktistakis, N.T. Phosphatidic acid formation by phospholipase D is required for transport from the endoplasmic reticulum to the Golgi complex. Curr. Biol. 7, 301–307 (1997).
Rizzo, M.A. et al. Phospholipase D and its product, phosphatidic acid, mediate agonist-dependent raf-1 translocation to the plasma membrane and the activation of the mitogen-activated protein kinase pathway. J. Biol. Chem. 274, 1131–1139 (1999).
Zhao, C. et al. Phospholipase D2-generated phosphatidic acid couples EGFR stimulation to Ras activation by Sos. Nat. Cell Biol. 9, 706–712 (2007).
Fang, Y. et al. Phosphatidic acid-mediated mitogenic activation of mTOR signaling. Science 294, 1942–1945 (2001).
Chen, Y., Zheng, Y. & Foster, D.A. Phospholipase D confers rapamycin resistance in human breast cancer cells. Oncogene 22, 3937–3942 (2003).
Williger, B.T., Ho, W.T. & Exton, J.H. Phospholipase D mediates matrix metalloproteinase-9 secretion in phorbol ester-stimulated human fibrosarcoma cells. J. Biol. Chem. 274, 735–738 (1999).
Kato, Y. et al. Acidic extracellular pH induces matrix metalloproteinase-9 expression in mouse metastatic melanoma cells through the phospholipase D-mitogen-activated protein kinase signaling. J. Biol. Chem. 280, 10938–10944 (2005).
Kang, D.W. et al. Phorbol ester up-regulates phospholipase D1 but not phospholipase D2 expression through a PKC/Ras/ERK/NFkappaB-dependent pathway and enhances matrix metalloproteinase-9 secretion in colon cancer cells. J. Biol. Chem. 283, 4094–4104 (2008).
Reich, R., Blumenthal, M. & Liscovitch, M. Role of phospholipase D in laminin-induced production of gelatinase A (MMP-2) in metastatic cells. Clin. Exp. Metastasis 13, 134–140 (1995).
Williger, B.T. et al. Release of gelatinase A (matrix metalloproteinase 2) induced by photolysis of caged phosphatidic acid in HT 1080 metastatic fibrosarcoma cells. J. Biol. Chem. 270, 29656–29659 (1995).
Wang, L. et al. Involvement of phospholipases D1 and D2 in sphingosine 1-phosphate-induced ERK (extracellular-signal-regulated kinase) activation and interleukin-8 secretion in human bronchial epithelial cells. Biochem. J. 367, 751–760 (2002).
Walker, S.J., Wu, W.J., Cerione, R.A. & Brown, H.A. Activation of phospholipase D1 by Cdc42 requires the Rho insert region. J. Biol. Chem. 275, 15665–15668 (2000).
Acknowledgements
The authors acknowledge outstanding support from A. Goodman (mass spectrometry) and D. Myers (data analysis and statistics) as well as R. Bruntz (Vanderbilt University) for assisting with the generation of the HEK293-gfpPLD2 stable cell line. We thank C. Rouzer for helpful discussions. We gratefully acknowledge partial support for this project from the A.B. Hancock Jr. Memorial Laboratory for Cancer Research and the Vanderbilt Institute for Chemical Biology (to H.A.B. and C.W.L.). P.E.S. was supported in part by a US National Research Service Award Training Fellowship (T32 GM007628). T.L.C. was partially supported by the US National Cancer Institute T32 grant CA09592 and the Breast Cancer Specialized Program of Research Excellence (P50-CA98131 to C.L.A.).
Author information
Authors and Affiliations
Contributions
S.A.S. carried out the cell-based inhibitor screening. P.E.S. and M.D.A. carried out in vitro inhibitor assays and protein purification. J.R.B. and A.L.T. performed chemical synthesis and characterization. H.P.C. was responsible for siRNA, RT-PCR and western blots. T.L.C. performed transwell experiments. C.L.A., C.W.L. and H.A.B. were responsible for coordination, planning and data interpretation of different aspects of the project.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Schemes 1–3 and Supplementary Methods (PDF 465 kb)
Rights and permissions
About this article
Cite this article
Scott, S., Selvy, P., Buck, J. et al. Design of isoform-selective phospholipase D inhibitors that modulate cancer cell invasiveness. Nat Chem Biol 5, 108–117 (2009). https://doi.org/10.1038/nchembio.140
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.140
This article is cited by
-
Inhibition of phospholipase D1 induces immunogenic cell death and potentiates cancer immunotherapy in colorectal cancer
Experimental & Molecular Medicine (2022)
-
Diacylglycerol kinase and phospholipase D inhibitors alter the cellular lipidome and endosomal sorting towards the Golgi apparatus
Cellular and Molecular Life Sciences (2021)
-
Inhibition of phospholipase D2 augments histone deacetylase inhibitor-induced cell death in breast cancer cells
Biological Research (2020)
-
Crystal structure of human PLD1 provides insight into activation by PI(4,5)P2 and RhoA
Nature Chemical Biology (2020)
-
Crystal structure of plant PLDα1 reveals catalytic and regulatory mechanisms of eukaryotic phospholipase D
Cell Research (2020)