Abstract
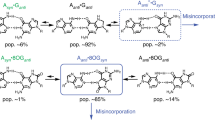
8-Oxopurines (8-oxodG and 8-oxodA) and formamidopyrimidines (FaPydG and FaPydA) are major oxidative DNA lesions involved in cancer development and aging. Their mutagenicity is believed to result from a conformational shift of the N9-C1′ glycosidic bonds from anti to syn, which allows the lesions to form noncanonical Hoogsteen-type base pairs with incoming triphosphates during DNA replication. Here we present biochemical data and what are to our knowledge the first crystal structures of carbocyclic FaPydA and FaPydG containing DNA in complex with a high-fidelity polymerase. Crystallographic snapshots show that the cFaPy lesions keep the anti geometry of the glycosidic bond during error-free and error-prone replication. The observed dG·dC→dT·dA transversion mutations are the result of base shifting and tautomerization.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Finkel, T. & Holbrook, N.J. Oxidants, oxidative stress and the biology of ageing. Nature 408, 239–247 (2000).
Cadet, J., Douki, T. & Ravanat, J.L. Oxidatively generated base damage to cellular DNA. Free Radic. Biol. Med. 49, 9–21 (2010).
Steenken, S. & Jovanovic, S.V. How easily oxidizible is DNA? One-electron reduction potentials of adenosine and guanosine radicals in aqueous solution. J. Am. Chem. Soc. 119, 617–618 (1997).
Cadet, J., Douki, T. & Ravanat, J.L. Oxidatively generated damage to the guanine moiety of DNA: mechanistic aspects and formation in cells. Acc. Chem. Res. 41, 1075–1083 (2008).
Jena, N.R. & Mishra, P.C. Formation of ring-opened and rearranged products of guanine: mechanisms and biological significance. Free Radic. Biol. Med. 53, 81–94 (2012).
Hsu, G.W., Ober, M., Carell, T. & Beese, L.S. Error-prone replication of oxidatively damaged DNA by a high-fidelity DNA polymerase. Nature 431, 217–221 (2004).
Duarte, V., Muller, J.G. & Burrows, C.J. Insertion of dGMP and dAMP during in vitro DNA synthesis opposite an oxidized form of 7,8-dihydro-8-oxoguanine. Nucleic Acids Res. 27, 496–502 (1999).
Brieba, L.G. et al. Structural basis for the dual coding potential of 8-oxoguanosine by a high-fidelity DNA polymerase. EMBO J. 23, 3452–3461 (2004).
Greenberg, M.M. The formamidopyrimidines: purine lesions formed in competition with 8-oxopurines from oxidative stress. Acc. Chem. Res. 45, 588–597 (2012).
Brieba, L.G. & Ellenberger, T. Hold tight (but not too tight) to get it right: accurate bypass of an 8-oxoguanine lesion by DNA polymerase β. Structure 11, 1–2 (2003).
Krahn, J.M., Beard, W.A., Miller, H., Grollman, A.P. & Wilson, S.H. Structure of DNA polymerase β with the mutagenic DNA lesion 8-oxodeoxyguanine reveals structural insights into its coding potential. Structure 11, 121–127 (2003).
Eoff, R.L., Irimia, A., Angel, K.C., Egli, M. & Guengerich, F.P. Hydrogen bonding of 7,8-dihydro-8-oxodeoxyguanosine with a charged residue in the little finger domain determines miscoding events in Sulfolobus solfataricus DNA polymerase Dpo4. J. Biol. Chem. 282, 19831–19843 (2007).
Vasquez-Del Carpio, R. et al. Structure of human DNA polymerase κ inserting dATP opposite an 8-OxoG DNA lesion. PLoS ONE 4, e5766 (2009).
Batra, V.K. et al. Mutagenic conformation of 8-oxo-7,8-dihydro-2′-dGTP in the confines of a DNA polymerase active site. Nat. Struct. Mol. Biol. 17, 889–890 (2010).
Dizdaroglu, M., Kirkali, G. & Jaruga, P. Formamidopyrimidines in DNA: mechanisms of formation, repair, and biological effects. Free Radic. Biol. Med. 45, 1610–1621 (2008).
Büsch, F. et al. Dissecting the differences between the α and β anomers of the oxidative DNA lesion FaPydG. Chemistry 14, 2125–2132 (2008).
Graziewicz, M.A. et al. Fapyadenine is a moderately efficient chain terminator for prokaryotic DNA polymerases. Free Radic. Biol. Med. 28, 75–83 (2000).
Delaney, M.O., Wiederholt, C.J. & Greenberg, M.M. Fapy.dA induces nucleotide misincorporation translesionally by a DNA polymerase. Angew. Chem. Int. Ed. Engl. 41, 771–773 (2002).
Wiederholt, C.J. & Greenberg, M.M. Fapy.dG instructs Klenow exo− to misincorporate deoxyadenosine. J. Am. Chem. Soc. 124, 7278–7279 (2002).
Ober, M., Müller, H., Pieck, C., Gierlich, J. & Carell, T. Base pairing and replicative processing of the formamidopyrimidine-dG DNA lesion. J. Am. Chem. Soc. 127, 18143–18149 (2005).
Tudek, B. et al. Mutagenic specificity of imidazole ring-opened 7-methylpurines in M13mp18 phage DNA. Acta Biochim. Pol. 46, 785–799 (1999).
Kalam, M.A. et al. Genetic effects of oxidative DNA damages: comparative mutagenesis of the imidazole ring-opened formamidopyrimidines (Fapy lesions) and 8-oxo-purines in simian kidney cells. Nucleic Acids Res. 34, 2305–2315 (2006).
Patro, J.N. et al. Studies on the replication of the ring opened formamidopyrimidine, Fapy.dG in Escherichia coli. Biochemistry 46, 10202–10212 (2007).
Asagoshi, K., Terato, H., Ohyama, Y. & Ide, H. Effects of a guanine-derived formamidopyrimidine lesion on DNA replication: translesion DNA synthesis, nucleotide insertion, and extension kinetics. J. Biol. Chem. 277, 14589–14597 (2002).
Christov, P.P., Angel, K.C., Guengerich, F.P. & Rizzo, C.J. Replication past the N5-methyl-formamidopyrimidine lesion of deoxyguanosine by DNA polymerases and an improved procedure for sequence analysis of in vitro bypass products by mass spectrometry. Chem. Res. Toxicol. 22, 1086–1095 (2009).
Ober, M., Linne, U., Gierlich, J. & Carell, T. The two main DNA lesions 8-oxo-7,8-dihydroguanine and 2,6-diamino-5-formamido-4-hydroxypyrimidine exhibit strongly different pairing properties. Angew. Chem. Int. Ed. Engl. 42, 4947–4951 (2003).
Ober, M., Marsch, M., Harms, K. & Carell, T. A carbocyclic analogue of a protected β-d-2-deoxyribosylamine. Acta Crystallogr. Sect. E Struct. Rep. Online 60, o1191–o1192 (2004).
Patro, J.N., Haraguchi, K., Delaney, M.O. & Greenberg, M.M. Probing the configurations of formamidopyrimidine lesions Fapy·dA and Fapy·dG in DNA using endonuclease IV. Biochemistry 43, 13397–13403 (2004).
Kiefer, J.R. et al. Crystal structure of a thermostable Bacillus DNA polymerase I large fragment at 2.1 Å resolution. Structure 5, 95–108 (1997).
Raoul, S., Bardet, M. & Cadet, J. Gamma irradiation of 2′-deoxyadenosine in oxygen-free aqueous solutions: identification and conformational features of formamidopyrimidine nucleoside derivatives. Chem. Res. Toxicol. 8, 924–933 (1995).
Lukin, M. et al. Novel post-synthetic generation, isomeric resolution, and characterization of Fapy-dG within oligodeoxynucleotides: differential anomeric impacts on DNA duplex properties. Nucleic Acids Res. 39, 5776–5789 (2011).
Münzel, M. et al. Improved synthesis and mutagenicity of oligonucleotides containing 5-hydroxymethylcytosine, 5-formylcytosine and 5-carboxylcytosine. Chemistry 17, 13782–13788 (2011).
Stathis, D., Lischke, U., Koch, S.C., Deiml, C.A. & Carell, T. Discovery and mutagenicity of a guanidinoformimine lesion as a new intermediate of the oxidative deoxyguanosine degradation pathway. J. Am. Chem. Soc. 134, 4925–4930 (2012).
Johnson, S.J., Taylor, J.S. & Beese, L.S. Processive DNA synthesis observed in a polymerase crystal suggests a mechanism for the prevention of frameshift mutations. Proc. Natl. Acad. Sci. USA 100, 3895–3900 (2003).
Topal, M.D. & Fresco, J.R. Complementary base pairing and the origin of substitution mutations. Nature 263, 285–289 (1976).
Wang, W., Hellinga, H.W. & Beese, L.S. Structural evidence for the rare tautomer hypothesis of spontaneous mutagenesis. Proc. Natl. Acad. Sci. USA 108, 17644–17648 (2011).
Johnson, S.J. & Beese, L.S. Structures of mismatch replication errors observed in a DNA polymerase. Cell 116, 803–816 (2004).
Creighton, S., Bloom, L.B. & Goodman, M.F. Gel fidelity assay measuring nucleotide misinsertion, exonucleolytic proofreading, and lesion bypass efficiencies. Methods Enzymol. 262, 232–256 (1995).
Münzel, M., Lercher, L., Müller, M. & Carell, T. Chemical discrimination between dC and 5MedC via their hydroxylamine adducts. Nucleic Acids Res. 38, e192 (2010).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
CCP4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Evans, P. Joint CCP4 and ESF-EACMB. Newsletter Prot. Crystallogr. 33, 22–24 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
Winn, M.D., Murshudov, G.N. & Papiz, M.Z. Macromolecular TLS refinement in REFMAC at moderate resolutions. Methods Enzymol. 374, 300–321 (2003).
Murshudov, G.N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Olson, W.K. et al. A standard reference frame for the description of nucleic acid base-pair geometry. J. Mol. Biol. 313, 229–237 (2001).
Lavery, R., Moakher, M., Maddocks, J.H., Petkeviciute, D. & Zakrzewska, K. Conformational analysis of nucleic acids revisited: Curves+. Nucleic Acids Res. 37, 5917–5929 (2009).
Acknowledgements
We are grateful to the beamline scientists at the Swiss Light Source and European Synchrotron Radiation Facility for setting up the beamlines. This research project was supported by the Deutsche Forschungsgemeinschaft through SFB 646 and SFB 749. Further support was obtained by the Volkswagen Foundation and in particular by the Excellence Cluster CiPSM. We thank K. Karaghiosoff and K. Lux for solving the crystal structures of the small molecules. We thank M. Müller for critical reading of the manuscript and many helpful discussions.
Author information
Authors and Affiliations
Contributions
T.C. conceived and directed the study. He wrote the manuscript and designed experiments. T.H.G. and U.L. designed experiments. T.H.G. performed the synthesis of the lesions and of the DNA strands. U.L. and T.H.G. performed the biochemical experiments. U.L. purified the protein. K.L.G. performed the synthesis of cdG. S.A. developed the synthesis of cFaPydA. H.C.M. developed the synthesis of cdG. S.S. conducted crystallographic data collection and solved the crystal structures. H.Z. performed the theoretical studies. D.S.S. performed the NMR studies.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Notes 1 and 2 (PDF 2994 kb)
Rights and permissions
About this article
Cite this article
Gehrke, T., Lischke, U., Gasteiger, K. et al. Unexpected non-Hoogsteen–based mutagenicity mechanism of FaPy-DNA lesions. Nat Chem Biol 9, 455–461 (2013). https://doi.org/10.1038/nchembio.1254
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1254
This article is cited by
-
Role of oxidative stress in carbon nanotube-generated health effects
Archives of Toxicology (2014)
-
FaPy lesions and DNA mutations
Nature Chemical Biology (2013)