Abstract
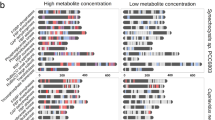
Functional assignment of uncharacterized proteins is a challenge in the era of large-scale genome sequencing. Here, we combine in extracto NMR, proteomics and transcriptomics with a newly developed (knock-out) metabolomics platform to determine a potential physiological role for a ribulose-1,5-bisphosphate carboxylase/oxygenase (RubisCO)-like protein from Rhodospirillum rubrum. Our studies unraveled an unexpected link in bacterial central carbon metabolism between S-adenosylmethionine–dependent polyamine metabolism and isoprenoid biosynthesis and also provide an alternative approach to assign enzyme function at the organismic level.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Phillips, R. & Milo, R. A feeling for the numbers in biology. Proc. Natl. Acad. Sci. USA 106, 21465–21471 (2009).
Tabita, F.R. et al. Function, structure, and evolution of the RubisCO-like proteins and their RubisCO homologs. Microbiol. Mol. Biol. Rev. 71, 576–599 (2007).
Hanson, T.E. & Tabita, F.R. A ribulose-1,5-bisphosphate carboxylase/oxygenase (RubisCO)-like protein from Chlorobium tepidum that is involved with sulfur metabolism and the response to oxidative stress. Proc. Natl. Acad. Sci. USA 98, 4397–4402 (2001).
Ashida, H. et al. A functional link between RuBisCO-like protein of Bacillus and photosynthetic RuBisCO. Science 302, 286–290 (2003).
Imker, H.J., Singh, J., Warlick, B.P., Tabita, F.R. & Gerlt, J.A. Mechanistic diversity in the RuBisCO superfamily: a novel isomerization reaction catalyzed by the RuBisCO-like protein from Rhodospirillum rubrum. Biochemistry 47, 11171–11173 (2008).
Pearce, F.G. Catalytic by-product formation and ligand binding by ribulose bisphosphate carboxylases from different phylogenies. Biochem. J. 399, 525–534 (2006).
Cleland, W.W., Andrews, T.J., Gutteridge, S., Hartman, F.C. & Lorimer, G.H. Mechanism of rubisco: the carbamate as general base. Chem. Rev. 98, 549–562 (1998).
Paech, C., Pierce, J., McCurry, S.D. & Tolbert, N.E. Inhibition of ribulose-1,5-biphosphate carboxylase/oxygenase by ribulose-1,5-bisphosphate epimerization and degradation products. Biochem. Biophys. Res. Commun. 83, 1084–1092 (1978).
Imker, H.J., Fedorov, A.A., Fedorov, E.V., Almo, S.C. & Gerlt, J.A. Mechanistic diversity in the RuBisCO superfamily: the “enolase” in the methionine salvage pathway in Geobacillus kaustophilus. Biochemistry 46, 4077–4089 (2007).
Carré-Mlouka, A. et al. A new rubisco-like protein coexists with a photosynthetic rubisco in the planktonic cyanobacteria. Microcystis. J. Biol. Chem. 281, 24462–24471.
Sekowska, A. et al. Bacterial variations on the methionine salvage pathway. BMC Microbiol. 4, 9 (2004).
Albers, E. Metabolic characteristics and importance of the universal methionine salvage pathway recycling methionine from 5′-methylthioadenosine. IUBMB Life 61, 1132–1142 (2009).
Singh, J. & Tabita, F.R. Roles of RubisCO and the RubisCO-like protein in 5-methylthioadenosine metabolism in the nonsulfur purple bacterium. Rhodospirillum rubrum. J. Bacteriol. 192, 1324–1331 (2010).
Sekowska, A., Mulard, L., Krogh, S., Tse, J.K. & Danchin, A. MtnK, methylthioribose kinase, is a starvation-induced protein in Bacillus subtilis. BMC Microbiol. 1, 15 (2001).
Sekowska, A., Robin, S., Daudin, J.J., Henaut, A. & Danchin, A. Extracting biological information from DNA arrays: an unexpected link between arginine and methionine metabolism in Bacillus subtilis. Genome Biol. 2, RESEARCH0019 (2001).
Sekowska, A. & Danchin, A. The methionine salvage pathway in Bacillus subtilis. BMC Microbiol. 2, 8 (2002).
Ellman, G.L. Tissue sulfhydryl groups. Arch. Biochem. Biophys. 82, 70–77 (1959).
Fagerbakke, K.M., Heldal, M. & Norland, S. Content of carbon, nitrogen, oxygen, sulfur and phosphorus in native aquatic and cultured bacteria. Aquat. Microb. Ecol. 10, 15–27 (1996).
Paulin, L.G., Brander, E.E. & Poso, H.J. Specific inhibition of spermidine synthesis in Mycobacteria spp. by the dextro isomer of ethambutol. Antimicrob. Agents Chemother. 28, 157–159 (1985).
Gopishetty, B. et al. Probing the catalytic mechanism of S-ribosylhomocysteinase (LuxS) with catalytic intermediates and substrate analogues. J. Am. Chem. Soc. 131, 1243–1250 (2009).
Kiene, R.P., Linn, L.J., Gonzalez, J., Moran, M.A. & Bruton, J.A. Dimethylsulfoniopropionate and methanethiol are important precursors of methionine and protein-sulfur in marine bacterioplankton. Appl. Environ. Microbiol. 65, 4549–4558 (1999).
Bolten, C.J., Schroder, H., Dickschat, J. & Wittmann, C. Towards methionine overproduction in Corynebacterium glutamicum—methanethiol and dimethyldisulfide as reduced sulfur sources. J. Microbiol. Biotechnol. 20, 1196–1203 (2010).
Rohmer, M., Knani, M., Simonin, P., Sutter, B. & Sahm, H. Isoprenoid biosynthesis in bacteria—a novel pathway for the early steps leading to isopentenyl diphosphate. Biochem. J. 295, 517–524 (1993).
Eisenreich, W., Bacher, A., Arigoni, D. & Rohdich, F. Biosynthesis of isoprenoids via the non-mevalonate pathway. Cell. Mol. Life Sci. 61, 1401–1426 (2004).
Hamana, K., Kamekura, M., Onishi, H., Akazawa, T. & Matsuzaki, S. Polyamines in photosynthetic eubacteria and extreme-halophilic archaebacteria. J. Biochem. 97, 1653–1658 (1985).
Hamana, K. & Matsuzaki, S. Polyamines as a chemotaxonomic marker in bacterial systematics. Crit. Rev. Microbiol. 18, 261–283 (1992).
Salim, H.M., Negritto, M.C. & Cavalcanti, A.R. 1+1 = 3: a fusion of 2 enzymes in the methionine salvage pathway of Tetrahymena thermophila creates a trifunctional enzyme that catalyzes 3 steps in the pathway. PLoS Genet. 5, e1000701 (2009).
Ashida, H. et al. RuBisCO-like proteins as the enolase enzyme in the methionine salvage pathway: functional and evolutionary relationships between RuBisCO-like proteins and photosynthetic RuBisCO. J. Exp. Bot. 59, 1543–1554 (2008).
Tabita, F.R., Hanson, T.E., Satagopan, S., Witte, B.H. & Kreel, N.E. Phylogenetic and evolutionary relationships of RubisCO and the RubisCO-like proteins and the functional lessons provided by diverse molecular forms. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 363, 2629–2640 (2008).
Tabita, F.R., Satagopan, S., Hanson, T.E., Kreel, N.E. & Scott, S.S. Distinct form I, II, III, and IV Rubisco proteins from the three kingdoms of life provide clues about Rubisco evolution and structure/function relationships. J. Exp. Bot. 59, 1515–1524 (2008).
Ashida, H., Danchin, A. & Yokota, A. Was photosynthetic RuBisCO recruited by acquisitive evolution from RuBisCO-like proteins involved in sulfur metabolism? Res. Microbiol. 156, 611–618 (2005).
Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Smith, C.A., Want, E.J., O'Maille, G., Abagyan, R. & Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Kanehisa, M. et al. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 34, D354–D357 (2006).
Smith, C.A. et al. METLIN: a metabolite mass spectral database. Ther. Drug Monit. 27, 747–751 (2005).
Wishart, D.S. et al. HMDB: a knowledgebase for the human metabolome. Nucleic Acids Res. 37, D603–D610 (2009).
Acknowledgements
This research was supported by a fellowship to T.J.E. from the Deutsche Forschungsgemeinschaft (ER 646/1-1) and research grants from the US National Institutes of Health (R01GM065155, R01GM095742, P01GM071790 and U54GM093342). We would like to thank J.E. Cronan for critical comments and C. Deane and T. Miyairi for excellent technical assistance. We also acknowledge S.C. Almo and B.S. Hillerich (New York Structural Genomics Research Consortium) for providing expression plasmids and purified cupin for in vitro assays.
Author information
Authors and Affiliations
Contributions
T.J.E., B.S.E. and J.A.G. conceived and designed the experiments. T.J.E. performed the experiments, with the exception of metabolome analyses, which were acquired by B.S.E., K.C. and B.M.W. and biochemical characterization of the cupin protein in vitro, which was performed by B.S.E and B.P.W. B.S.E., K.C., B.M.W. and J.V.S. developed the mass spectrometric platform and evaluated MS data. J.S. and F.R.T. created and provided the rlp mutant strain and the cupin mutant strain of R. rubrum. B.P.W. and H.J.I. provided chemicals and recombinant enzymes for the synthesis of chemicals. T.J.E. and J.A.G. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 7127 kb)
Rights and permissions
About this article
Cite this article
Erb, T., Evans, B., Cho, K. et al. A RubisCO-like protein links SAM metabolism with isoprenoid biosynthesis. Nat Chem Biol 8, 926–932 (2012). https://doi.org/10.1038/nchembio.1087
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1087
This article is cited by
-
Multilevel interactions between native and ectopic isoprenoid pathways affect global metabolism in rice
Transgenic Research (2022)
-
Towards efficient terpenoid biosynthesis: manipulating IPP and DMAPP supply
Bioresources and Bioprocessing (2019)
-
Functional assignment of multiple catabolic pathways for d-apiose
Nature Chemical Biology (2018)
-
Comparative genomic inference suggests mixotrophic lifestyle for Thorarchaeota
The ISME Journal (2018)
-
Discovery of a RuBisCO-like Protein that Functions as an Oxygenase in the Novel d-Hamamelose Pathway
Biotechnology and Bioprocess Engineering (2018)