Abstract
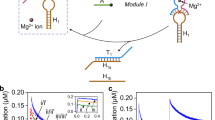
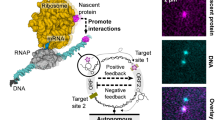
DNA performs diverse functional roles in biology, nanotechnology and biotechnology, but current methods for autonomously synthesizing arbitrary single-stranded DNA are limited. Here, we introduce the concept of primer exchange reaction (PER) cascades, which grow nascent single-stranded DNA with user-specified sequences following prescribed reaction pathways. PER synthesis happens in a programmable, autonomous, in situ and environmentally responsive fashion, providing a platform for engineering molecular circuits and devices with a wide range of sensing, monitoring, recording, signal-processing and actuation capabilities. We experimentally demonstrate a nanodevice that transduces the detection of a trigger RNA into the production of a DNAzyme that degrades an independent RNA substrate, a signal amplifier that conditionally synthesizes long fluorescent strands only in the presence of a particular RNA signal, molecular computing circuits that evaluate logic (AND, OR, NOT) combinations of RNA inputs, and a temporal molecular event recorder that records in the PER transcript the order in which distinct RNA inputs are sequentially detected.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Zheng, J. et al. From molecular to macroscopic via the rational design of a self-assembled 3D DNA crystal. Nature 461, 74–77 (2009).
Wei, B., Dai, M. & Yin, P. Complex shapes self-assembled from single-stranded DNA tiles. Nature 485, 623–626 (2012).
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Three-dimensional structures self-assembled from DNA bricks. Science 338, 1177–1183 (2012).
Han, D. et al. DNA gridiron nanostructures based on four-arm junctions. Science 339, 1412–1415 (2013).
Gerling, T., Wagenbauer, K. F., Neuner, A. M. & Dietz, H. Dynamic DNA devices and assemblies formed by shape-complementary, non-base pairing 3D components. Science 347, 1446–1452 (2015).
Benson, E. et al. DNA rendering of polyhedral meshes at the nanoscale. Nature 523, 441–444 (2015).
Dunn, K. E. et al. Guiding the folding pathway of DNA origami. Nature 525, 82–86 (2015).
Veneziano, R. et al. Designer nanoscale DNA assemblies programmed from the top down. Science 352, 1534 (2016).
Seelig, G., Soloveichik, D., Zhang, D. Y. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Zhang, D. Y., Turberfield, A. J., Yurke, B. & Winfree, E. Engineering entropy-driven reactions and networks catalyzed by DNA. Science 318, 1121–1125 (2007).
Yin, P., Choi, H. M. T., Calvert, C. R. & Pierce, N. A. Programming biomolecular self-assembly pathways. Nature 451, 318–322 (2008).
Omabegho, T., Sha, R. & Seeman, N. C. A bipedal DNA Brownian motor with coordinated legs. Science 324, 67–71 (2009).
Qian, L. & Winfree, E. Scaling up digital circuit computation with DNA strand displacement cascades. Science 332, 1196–1201 (2011).
Chirieleison, S. M., Allen, P. B., Simpson, Z. B., Ellington, A. D. & Chen, X. Pattern transformation with DNA circuits. Nat. Chem. 5, 1000–1005 (2013).
Weitz, M. et al. Diversity in the dynamical behaviour of a compartmentalized programmable biochemical oscillator. Nat. Chem. 6, 295–302 (2014).
Karzbrun, E., Tayar, A. M., Noireaux, V. & Bar-Ziv, R. H. Programmable on-chip DNA compartments as artificial cells. Science 345, 829–832 (2014).
Mohammed, A. M., Šulc, P., Zenk, J. & Schulman, R. Self-assembling DNA nanotubes to connect molecular landmarks. Nat. Nanotech. 12, 312–316 (2016).
Choi, H. M. T. et al. Programmable in situ amplification for multiplexed imaging of mRNA expression. Nat. Biotechnol. 28, 1208–1212 (2010).
Zhang, D. Y., Chen, S. X. & Yin, P. Optimizing the specificity of nucleic acid hybridization. Nat. Chem. 4, 208–214 (2012).
Douglas, S. M., Bachelet, I. & Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science 335, 831–834 (2012).
Kuzyk, A. et al. DNA-based self-assembly of chiral plasmonic nanostructures with tailored optical response. Nature 483, 311–314 (2012).
Derr, N. D. et al. Tug-of-war in motor protein ensembles revealed with a programmable DNA origami scaffold. Science 338, 662–665 (2012).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and exchange-PAINT. Nat. Methods 11, 313–318 (2014).
Sun, W. et al. Casting inorganic structures with DNA molds. Science 346, 1258361 (2014).
Wang, J. S. & Zhang, D. Y. Simulation-guided DNA probe design for consistently ultraspecific hybridization. Nat. Chem. 7, 545–553 (2015).
Gopinath, A., Miyazono, E., Faraon, A. & Rothemund, P. W. K. Engineering and mapping nanocavity emission via precision placement of DNA origami. Nature 535, 401–405 (2016).
Bhatia, D. et al. Quantum dot-loaded monofunctionalized DNA icosahedra for single-particle tracking of endocytic pathways. Nat. Nanotech. 11, 1112–1119 (2016).
Kilchherr, F. et al. Single-molecule dissection of stacking forces in DNA. Science 353, aaf5508 (2016).
Nickels, P. C. et al. Molecular force spectroscopy with a DNA origami-based nanoscopic force clamp. Science 354, 305–307 (2016).
Walker, G. T. et al. Strand displacement amplification—an isothermal, in vitro DNA amplification technique. Nucleic Acids Res. 20, 1691–1696 (1992).
Lizardi, P. M. et al. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat. Genet. 19, 225–232 (1998).
Notomi, T. et al. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 28, e63 (2000).
Wang, H. H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Du, Y. & Dong, S. Nucleic acid biosensors: recent advances and perspectives. Anal. Chem. 89, 189–215 (2017).
Church, G. M., Gao, Y. & Kosuri, S. Next-generation digital information storage in DNA. Science 337, 1628 (2012).
Kosuri, S. & Church, G. M. Large-scale de novo DNA synthesis: technologies and applications. Nat. Methods 11, 499–507 (2014).
Saiki, R. K. et al. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239, 487–491 (1988).
Montagne, K., Plasson, R., Sakai, Y., Fujii, T. & Rondelez, Y. Programming an in vitro DNA oscillator using a molecular networking strategy. Mol. Syst. Biol. 7, 466 (2011).
Baccouche, A., Montagne, K., Padirac, A., Fujii, T. & Rondelez, Y. Dynamic DNA-toolbox reaction circuits: a walkthrough. Methods 67, 234–249 (2014).
Lee, C. S., Davis, R. W. & Davidson, N. A physical study by electron microscopy of the terminally repetitious, circularly permuted DNA from the coliphage particles of Escherichia coli 15. J. Mol. Biol. 48, 1–22 (1970).
Schaus, T. E., Woo, S., Xuan, F., Chen, X. & Yin, P. A DNA nanoscope via auto-cycling proximity recording. Nat. Commun. 8, 696 (2017).
Newton, C. R. et al. The production of PCR products with 5′ single-stranded tails using primers that incorporate novel phosphoramidite intermediates. Nucleic Acids Res. 21, 1155–1162 (1993).
Sakamoto, K. et al. State transitions by molecules. Biosystems 52, 81–91 (1999).
Whitcombe, D., Theaker, J., Guy, S. P., Brown, T. & Little, S. Detection of PCR products using self-probing amplicons and fluorescence. Nat. Biotechnol. 17, 804–807 (1999).
Aubert, N., Rondelez, Y., Fujii, T. & Hagiya, M. Enforcing logical delays in DNA computing systems. Nat. Comput. 13, 559–572 (2014).
Grillari, J., Hackl, M. & Grillari-Voglauer, R. miR-17-92 cluster: ups and downs in cancer and aging. Biogerontology 11, 501–506 (2010).
Hjiantoniou, E., Iseki, S., Uney, J. B. & Phylactou, L. A. DNAzyme-mediated cleavage of twist transcripts and increase in cellular apoptosis. Biochem. Biophys. Res. Commun. 300, 178–181 (2003).
Santoro, S. W. & Joyce, G. F. A general purpose RNA-cleaving DNA enzyme. Proc. Natl Acad. Sci. USA 94, 4262–4266 (1997).
Mohanty, J. et al. Thioflavin T as an efficient inducer and selective fluorescent sensor for the human telomeric G-quadruplex DNA. J. Am. Chem. Soc. 135, 367–376 (2012).
Yurke, B., Turberfield, A. J., Mills, A. P., Simmel, F. C. & Neumann, J. L. A DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nat. Chem. 3, 103–113 (2011).
Hagiya, M., Arita, M., Kiga, D., Sakamoto, K. & Yokoyama, S. in DNA Based Computers III Vol. 48 (eds Rubin, H. & Wood, D. H.) 57–72 (DIMACS Series in Discrete Mathematics and Theoretical Computer Science, American Mathematical Society, 1999).
Winfree, E. Whiplash PCR for O(1) Computing Technical Report 1998.23 (Caltech, 1998).
Rose, J. A., Deaton, R. J., Hagiya, M. & Suyama, A. in DNA Computing (Jonoska, N. & Seeman, N. C.) 104–116 (Springer, 2002).
Komiya, K., Yamamura, M. & Rose, J. A. in International Workshop on DNA-Based Computers 1–10 (Springer, 2008).
Reif, J. H. & Majumder, U. Isothermal reactivating whiplash PCR for locally programmable molecular computation. Nat. Comput. 9, 183–206 (2010).
Fujii, T. & Rondelez, Y. Predator–prey molecular ecosystems. ACS Nano 7, 27–34 (2012).
Padirac, A., Fujii, T., Estévez-Torres, A. & Rondelez, Y. Spatial waves in synthetic biochemical networks. J. Am. Chem. Soc. 135, 14586–14592 (2013).
Zadorin, A. S., Rondelez, Y., Galas, J.-C. & Estevez-Torres, A. Synthesis of programmable reaction-diffusion fronts using DNA catalyzers. Phys. Rev. Lett. 114, 068301 (2015).
Dirks, R. M. & Pierce, N. A. Triggered amplification by hybridization chain reaction. Proc. Natl Acad. Sci. USA 101, 15275–15278 (2004).
Green, A. A., Silver, P. A., Collins, J. J. & Yin, P. Toehold switches: de-novo-designed regulators of gene expression. Cell 159, 925–939 (2014).
Dirks, R. M., Bois, J. S., Schaeffer, J. M., Winfree, E. & Pierce, N. A. Thermodynamic analysis of interacting nucleic acid strands. SIAM Rev. 49, 65–88 (2007).
Zadeh, J. N. et al. NUPACK: analysis and design of nucleic acid systems. J. Comput. Chem. 32, 170–173 (2011).
Wolfe, B. R. & Pierce, N. A. Sequence design for a test tube of interacting nucleic acid strands. ACS Synth. Biol. 4, 1086–1100 (2014).
Acknowledgements
The authors thank W. Shih, J. Kim, X. Chen, N. Hanikel, E. Winfree, B. Beliveau and N. Liu for their discussions and comments. This work was supported by the Office on Naval Research (grants N000141310593, N000141410610, N000141612182 and N000141612410), the National Science Foundation (grants CCF1317291, CMMI1334109 and 1540214), the National Institutes of Health (grant 1R01EB01865901) and the Wyss Institute's Molecular Robotics Initiative. J. Kishi was supported by an NSF graduate research fellowship and T. Schaus was supported by the Jane Coffin Childs Postdoctoral Fellowship.
Author information
Authors and Affiliations
Contributions
J.Y.K. conceived and designed the study, designed and performed the experiments, analysed the data and wrote the manuscript. T.E.S. designed and performed the experiments and analysed the data. N.G. designed and performed the experiments and analysed the data. F.X. designed and performed the experiments and analysed the data. P.Y. conceived and supervised the study, interpreted the data and wrote the manuscript. All authors reviewed, edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
A provisional US patent has been filed based on this work. P.Y. is co-founder of Ultivue Inc. and NuProbe Global.
Supplementary information
Supplementary information
Supplementary information (PDF 21412 kb)
Rights and permissions
About this article
Cite this article
Kishi, J., Schaus, T., Gopalkrishnan, N. et al. Programmable autonomous synthesis of single-stranded DNA. Nature Chem 10, 155–164 (2018). https://doi.org/10.1038/nchem.2872
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2872
This article is cited by
-
Highly sensitive SERS platform for pathogen analysis by cyclic DNA nanostructure@AuNP tags and cascade primer exchange reaction
Journal of Nanobiotechnology (2024)
-
DNA as a universal chemical substrate for computing and data storage
Nature Reviews Chemistry (2024)
-
Molecular robotic agents that survey molecular landscapes for information retrieval
Nature Communications (2024)
-
A simple and available measurement of onco-sEV dsDNA to protein ratio as a potential tumor marker
BMC Cancer (2023)
-
DNA-based programmable gate arrays for general-purpose DNA computing
Nature (2023)