Abstract
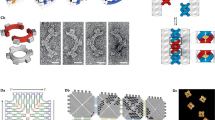
The ability of DNA to store and encode information arises from base pairing of the four-letter nucleobase code to form a double helix. Expanding this DNA ‘alphabet’ by synthetic incorporation of new bases can introduce new functionalities and enable the formation of novel nucleic acid structures. However, reprogramming the self-assembly of existing nucleobases presents an alternative route to expand the structural space and functionality of nucleic acids. Here we report the discovery that a small molecule, cyanuric acid, with three thymine-like faces, reprogrammes the assembly of unmodified poly(adenine) (poly(A)) into stable, long and abundant fibres with a unique internal structure. Poly(A) DNA, RNA and peptide nucleic acid (PNA) all form these assemblies. Our studies are consistent with the association of adenine and cyanuric acid units into a hexameric rosette, which brings together poly(A) triplexes with a subsequent cooperative polymerization. Fundamentally, this study shows that small hydrogen-bonding molecules can be used to induce the assembly of nucleic acids in water, which leads to new structures from inexpensive and readily available materials.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Watson, J. D. & Crick, F. H. Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature 171, 737–738 (1953).
Choi, J. & Majima, T. Conformational changes of non-B DNA. Chem. Soc. Rev. 40, 5893–5909 (2011).
Aldaye, F. A., Palmer, A. L. & Sleiman, H. F. Assembling materials with DNA as the guide. Science 321, 1795–1799 (2008).
Egholm, M., Buchardt, O., Nielsen, P. E. & Berg, R. H. Peptide nucleic-acids (PNA)—oligonucleotide analogs with an achiral peptide backbone. J. Am. Chem. Soc. 114, 1895–1897 (1992).
Egholm, M. et al. PNA hybridizes to complementary oligonucleotides obeying the Watson–Crick hydrogen-bonding rules. Nature 365, 566–568 (1993).
Chen, M. C. et al. Spontaneous prebiotic formation of a β-ribofuranoside that self-assembles with a complementary heterocycle. J. Am. Chem. Soc. 136, 5640–5646 (2014).
Malyshev, D. A. et al. A semi-synthetic organism with an expanded genetic alphabet. Nature 509, 385–388 (2014).
Yang, Z., Chen, F., Chamberlin, S. G. & Benner, S. A. Expanded genetic alphabets in the polymerase chain reaction. Angew. Chem. Int. Ed. 49, 177–180 (2010).
Teo, Y. N. & Kool, E. T. DNA-multichromophore systems. Chem. Rev. 112, 4221–4245 (2012).
Winnacker, M. & Kool, E. T. Artificial genetic sets composed of size-expanded base pairs. Angew. Chem. Int. Ed. 52, 12498–12508 (2013).
Sessler, J. L. & Jayawickramarajah, J. Functionalized base-pairs: versatile scaffolds for self-assembly. Chem. Commun. 1939–1949 (2005).
Woo, S. & Rothemund, P. W. K. Programmable molecular recognition based on the geometry of DNA nanostructures. Nature Chem. 3, 620–627 (2011).
Yang, H. et al. Metal–nucleic acid cages. Nature Chem. 1, 390–396 (2009).
Piccirilli, J. A., Krauch, T., Moroney, S. E. & Benner, S. A. Enzymatic incorporation of a new base pair into DNA and RNA extends the genetic alphabet. Nature 343, 33–37 (1990).
Kimoto, M., Kawai, R., Mitsui, T., Yokoyama, S. & Hirao, I. An unnatural base pair system for efficient PCR amplification and functionalization of DNA molecules. Nucleic Acids Res. 37, e14 (2009).
Ogawa, A. K. et al. Efforts toward the expansion of the genetic alphabet: information storage and replication with unnatural hydrophobic base pairs. J. Am. Chem. Soc. 122, 3274–3287 (2000).
Wu, Y. Q. et al. Efforts toward expansion of the genetic alphabet: optimization of interbase hydrophobic interactions. J. Am. Chem. Soc. 122, 7621–7632 (2000).
Meggers, E., Holland, P. L., Tolman, W. B., Romesberg, F. E. & Schultz, P. G. A novel copper-mediated DNA base pair. J. Am. Chem. Soc. 122, 10714–10715 (2000).
Shionoya, M. & Tanaka, K. Synthetic incorporation of metal complexes into nucleic acids and peptides directed toward functionalized molecules. Bull. Chem. Soc. Jpn 73, 1945–1954 (2000).
Martin-Ortiz, M., Gomez-Gallego, M., Ramirez de Arellano, C. & Sierra, M. A. The selective synthesis of metallanucleosides and metallanucleotides: a new tool for the functionalization of nucleic acids. Chem. Eur. J. 18, 12603–12608 (2012).
Chen, H., Meena & McLaughlin, L. W. A Janus-wedge DNA triplex with A-W1-T and G-W2-C base triplets. J. Am. Chem. Soc. 130, 13190–13191 (2008).
Shin, D. & Tor, Y. Bifacial nucleoside as a surrogate for both T and A in duplex DNA. J. Am. Chem. Soc. 133, 6926–6929 (2011).
Yang, H. Z., Pan, M. Y., Jiang, D. W. & He, Y. Synthesis of Janus type nucleoside analogues and their preliminary bioactivity. Org. Biomol. Chem. 9, 1516–1522 (2011).
Pan, M.-Y. et al. Janus-type AT nucleosides: synthesis, solid and solution state structures. Org. Biomol. Chem. 9, 5692–5702 (2011).
Largy, E., Liu, W., Hasan, A. & Perrin, D. M. Base-pairing behavior of a carbocyclic Janus–AT nucleoside analogue capable of recognizing A and T within a DNA duplex. ChemBioChem 14, 2199–2208 (2013).
Marsh, A., Silvestri, M. & Lehn, J. M. Self-complementary hydrogen bonding heterocycles designed for the enforced self-assembly into supramolecular macrocycles. Chem. Commun. 1527–1528 (1996).
Zeng, Y., Pratumyot, Y., Piao, X. & Bong, D. Discrete assembly of synthetic peptide–DNA triplex structures from polyvalent melamine–thymine bifacial recognition. J. Am. Chem. Soc. 134, 832–835 (2012).
Chaput, J. C. & Switzer, C. A DNA pentaplex incorporating nucleobase quintets. Proc. Natl Acad. Sci. USA 96, 10614–10619 (1999).
Jiang, D. & Seela, F. Oligonucleotide duplexes and multistrand assemblies with 8-aza-2′-deoxyisoguanosine: a fluorescent isoG(d) shape mimic expanding the genetic alphabet and forming ionophores. J. Am. Chem. Soc. 132, 4016–4024 (2010).
Polak, M. & Hud, N. V. Complete disproportionation of duplex poly(dT)·poly(dA) into triplex poly(dT)·poly(dA)·poly(dT) and poly(dA) by coralyne. Nucleic Acids Res. 30, 983–992 (2002).
Persil, Ö., Santai, C. T., Jain, S. S. & Hud, N. V. Assembly of an antiparallel homo-adenine DNA duplex by small-molecule binding. J. Am. Chem. Soc. 126, 8644–8645 (2004).
Jain, S. S., Polak, M. & Hud, N. V. Controlling nucleic acid secondary structure by intercalation: effects of DNA strand length on coralyne-driven duplex disproportionation. Nucleic Acids Res. 31, 4608–4615 (2003).
Çetinkol, O. P. & Hud, N. V. Molecular recognition of poly(A) by small ligands: an alternative method of analysis reveals nanomolar, cooperative and shape-selective binding. Nucleic Acids Res. 37, 611–621 (2009).
Park, K. S. & Park, H. G. A DNA-templated silver nanocluster probe for label-free, turn-on fluorescence-based screening of homo-adenine binding molecules. Biosens. Bioelectron. 64, 618–624 (2015).
Balasubramanian, S., Hurley, L. H. & Neidle, S. Targeting G-quadruplexes in gene promoters: a novel anticancer strategy? Nature Rev. Drug Discov. 10, 261–275 (2011).
Rich, A., Davies, D. R., Crick, F. H. & Watson, J. D. The molecular structure of polyadenylic acid. J. Mol. Biol. 3, 71–86 (1961).
Chakraborty, S., Sharma, S., Maiti, P. K. & Krishnan, Y. The poly dA helix: a new structural motif for high performance DNA-based molecular switches. Nucleic Acids Res. 37, 2810–2817 (2009).
Zerkowski, J. A., Seto, C. T. & Whitesides, G. M. Solid-state structures of rosette and crinkled tape motifs derived from the cyanuric acid melamine lattice. J. Am. Chem. Soc. 114, 5473–5475 (1992).
Rakotondradany, F. et al. Hydrogen-bond self-assembly of DNA-analogues into hexameric rosettes. Chem. Commun. 5441–5443 (2005).
Roy, B., Bairi, P. & Nandi, A. K. Supramolecular assembly of melamine and its derivatives: nanostructures to functional materials. RSC Adv. 4, 1708–1734 (2014).
Berova, N., Nakanishi, K. & Woody, R. Circular Dichroism: Principles and Applications (Wiley-VCH, 2000).
De Greef, T. F. et al. Supramolecular polymerization. Chem. Rev. 109, 5687–5754 (2009).
Safaee, N. et al. Structure of the parallel duplex of poly(A) RNA: evaluation of a 50 year-old prediction. Angew. Chem. Int. Ed. 52, 10370–10373 (2013).
Krishnan, Y. & Simmel, F. C. Nucleic acid based molecular devices. Angew. Chem. Int. Ed. 50, 3124–3156 (2011).
Ma, M. & Bong, D. Determinants of cyanuric acid and melamine assembly in water. Langmuir 27, 8841–8853 (2011).
Cafferty, B. J. et al. Efficient self-assembly in water of long noncovalent polymers by nucleobase analogues. J. Am. Chem. Soc. 135, 2447–2450 (2013).
Whitesides, G. M. et al. Noncovalent synthesis: using physical–organic chemistry to make aggregates. Acc. Chem. Res. 28, 37–44 (1995).
Timmerman, P. et al. NMR diffusion spectroscopy for the characterization of multicomponent hydrogen-bonded assemblies in solution. J. Chem. Soc. Perkin Trans. 2, 2077–2089 (2000).
Davies, R. J. & Davidson, N. Base pairing equilibria between polynucleotides and complementary monomers. Biopolymers 10, 1455–1479 (1971).
Davies, R. J. Complexes of poly(adenylic acid) with complementary monomers. Eur. J. Biochem. 61, 225–236 (1976).
Pyle, A. M. et al. Mixed-ligand complexes of ruthenium(II): factors governing binding to DNA. J. Am. Chem. Soc. 111, 3051–3058 (1989).
Chworos, A. et al. Building programmable jigsaw puzzles with RNA. Science 306, 2068–2072 (2004).
Hao, C. et al. Construction of RNA nanocages by re-engineering the packaging RNA of Phi29 bacteriophage. Nature Commun. 5, 3890 (2014).
Geary, C., Rothemund, P. W. K. & Andersen, E. S. A single-stranded architecture for cotranscriptional folding of RNA nanostructures. Science 345, 799–804 (2014).
Weill, L., Belloc, E., Bava, F.-A. & Mendez, R. Translational control by changes in poly(A) tail length: recycling mRNAs. Nature Struct. Mol. Biol. 19, 577–585 (2012).
Di Giammartino, D. C., Nishida, K. & Manley, J. L. Mechanisms and consequences of alternative polyadenylation. Mol. Cell 43, 853–866 (2011).
Acknowledgements
We acknowledge the Natural Sciences and Engineering Research Council of Canada (NSERC), the Canada Research Chairs Program, the Canada Foundation for Innovation, the Centre for Self-Assembled Chemical Structures and the Canadian Institute for Advanced Research for financial support. N.A. and A.G. thank NSERC and Fonds Québécois de la Recherche sur la Nature et les Technologies for doctoral fellowships. C.J.S. thanks NSERC for a Banting Postdoctoral Fellowship. H.F.S. is a Cottrell Scholar of the Research Corporation.
Author information
Authors and Affiliations
Contributions
H.F.S. and F.A. conceived the study. N.A., F.A. and H.F.S. designed the experiments. N.A. performed the experimental studies with assistance from A.A.G. (DNA synthesis), C.J.S. (adenosine/CA crystal growth and X-ray analysis), A.P. (VPO) and V.T. (synthesis of TIPDS-dA and hex-CA). N.A. and H.F.S analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 3101 kb)
Rights and permissions
About this article
Cite this article
Avakyan, N., Greschner, A., Aldaye, F. et al. Reprogramming the assembly of unmodified DNA with a small molecule. Nature Chem 8, 368–376 (2016). https://doi.org/10.1038/nchem.2451
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2451
This article is cited by
-
Prebiotic synthesis of noncanonical nucleobases under plausible alkaline hydrothermal conditions
Scientific Reports (2022)
-
Dissipative DNA fibres
Nature Chemistry (2021)
-
A dissipative pathway for the structural evolution of DNA fibres
Nature Chemistry (2021)
-
A poly(thymine)–melamine duplex for the assembly of DNA nanomaterials
Nature Materials (2020)
-
Modular self-assembly of gamma-modified peptide nucleic acids in organic solvent mixtures
Nature Communications (2020)