Abstract
Early diagnosis of tuberculosis can dramatically reduce both its transmission and the associated death rate. The extremely slow growth rate of the causative pathogen, Mycobacterium tuberculosis (Mtb), however, makes this challenging at the point of care, particularly in resource-limited settings. Here we report the use of BlaC (an enzyme naturally expressed/secreted by tubercle bacilli) as a marker and the design of BlaC-specific fluorogenic substrates as probes for Mtb detection. These probes showed an enhancement by 100–200 times in fluorescence emission on BlaC activation and a greater than 1,000-fold selectivity for BlaC over TEM-1 β-lactamase, an important factor in reducing false-positive diagnoses. Insight into the BlaC specificity was revealed by successful co-crystallization of the probe/enzyme mutant complex. A refined green fluorescent probe (CDG-OMe) enabled the successful detection of live pathogen in less than ten minutes, even in unprocessed human sputum. This system offers the opportunity for the rapid, accurate detection of very low numbers of Mtb for the clinical diagnosis of tuberculosis in sputum and other specimens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Dye, C. et al. Measuring tuberculosis burden, trends, and the impact of control programmes. Lancet Infect. Dis. 8, 233–243 (2008).
Dinnes, J. et al. A systematic review of rapid diagnostic tests for the detection of tuberculosis infection. Health Technol. Assess. 11, 1–196 (2007).
Ling, D. I., Flores, L. L., Riley, L. W. & Pai, M. Commercial nucleic-acid amplification tests for diagnosis of pulmonary tuberculosis in respiratory specimens: meta-analysis and meta-regression. PLoS One 3, e1536 (2008).
Greco, S., Girardi, E., Navarra, A. & Saltini, C. Current evidence on diagnostic accuracy of commercially based nucleic acid amplification tests for the diagnosis of pulmonary tuberculosis. Thorax 61, 783–790 (2006).
Boehme, C. C. et al. Rapid molecular detection of tuberculosis and rifampin resistance. N. Engl. J. Med. 363, 1005–1015 (2010).
Hughes, R., Wonderling, D., Li, B. & Higgins, B. The cost effectiveness of nucleic acid amplification techniques for the diagnosis of tuberculosis. Respir. Med. 106, 300–307 (2012).
Flores, A. R., Parsons, L. M. & Pavelka, M. S. Jr Genetic analysis of the beta-lactamases of Mycobacterium tuberculosis and Mycobacterium smegmatis and susceptibility to beta-lactam antibiotics. Microbiology 151, 521–532 (2005).
Hugonnet, J. E., Tremblay, L. W., Boshoff, H. I., Barry, C. E. II & Blanchard, J. S. Meropenem–clavulanate is effective against extensively drug-resistant Mycobacterium tuberculosis. Science 323, 1215–1218 (2009).
Boyd, D. B. & Lunn, W. H. Electronic structures of cephalosporins and penicillins. 9. Departure of a leaving group in cephalosporins. J. Med. Chem. 22, 778–784 (1979).
Faraci, W. S. & Pratt, R. F. Elimination of a good leaving group from the 3′-position of a cephalosporin need not be concerted with beta-lactam ring-opening – TEM-2 beta-lactamase-catalyzed hydrolysis of pyridine-2-azo-4′-(N′,N′-dimethylaniline) cephalosporin (PADAC) and of cephaloridine. J. Am. Chem. Soc. 106, 1489–1490 (1984).
Faraci, W. S. & Pratt, R. F. Mechanism of inhibition of the PC1 beta-lactamase of Staphylococcus aureus by cephalosporins: importance of the 3′-leaving group. Biochemistry 24, 903–910 (1985).
Pratt, R. F. & Faraci, W. S. Direct observation by proton NMR of cephalosporoate intermediates in aqueous solution during the hydrazinolysis and beta-lactamase-catalyzed hydrolysis of cephalosporins with 3′ leaving groups: kinetics and equilibria of the 3′ elimination reaction. J. Am. Chem. Soc. 108, 5328–5333 (1986).
Zlokarnik, G. et al. Quantitation of transcription and clonal selection of single living cells with beta-lactamase as reporter. Science 279, 84–88 (1998).
Gao, W., Xing, B., Tsien, R. Y. & Rao, J. Novel fluorogenic substrates for imaging beta-lactamase gene expression. J. Am. Chem. Soc. 125, 11146–11147 (2003).
Xing, B., Khanamiryan, A. & Rao, J. Cell-permeable near-infrared fluorogenic substrates for imaging beta-lactamase activity. J. Am. Chem. Soc. 127, 4158–4159 (2005).
Yao, H., So, M. K. & Rao, J. A bioluminogenic substrate for in vivo imaging of beta-lactamase activity. Angew. Chem. Int. Ed. 46, 7031–7034 (2007).
Rukavishnikov, A., Gee, K. R., Johnson, I. & Corry, S. Fluorogenic cephalosporin substrates for beta-lactamase TEM-1. Anal. Biochem. 419, 9–16 (2011).
Kong, Y. et al. Imaging tuberculosis with endogenous beta-lactamase reporter enzyme fluorescence in live mice. Proc. Natl Acad. Sci. USA 107, 12239–12244 (2010).
Banerjee, S., Pieper, U., Kapadia, G., Pannell, L. K. & Herzberg, O. Role of the Omega-loop in the activity, substrate specificity, and structure of class A beta-lactamase. Biochemistry 37, 3286–3296 (1998).
Knox, J. R. Extended-spectrum and inhibitor-resistant TEM-type beta-lactamases – mutations, specificity, and 3-dimensional structure. Antimicrob. Agents Chemother. 39, 2593–2601 (1995).
Albrecht, H. A. et al. Cephalosporin 3′-quinolone esters with a dual mode of action. J. Med. Chem. 33, 77–86 (1990).
Baldwin, J. E., Urban, F. J., Cooper, R. D. G. & Jose, F. L. Direct 6-methoxylation of penicillin derivatives – convenient pathway to substituted beta-lactam antibiotics. J. Am. Chem. Soc. 95, 2401–2403 (1973).
Wang, F., Cassidy, C. & Sacchettini, J. C. Crystal structure and activity studies of the Mycobacterium tuberculosis beta-lactamase reveal its critical role in resistance to beta-lactam antibiotics. Antimicrob. Agents Chemother. 50, 2762–2771 (2006).
Minasov, G., Wang, X. J. & Shoichet, B. K. An ultrahigh resolution structure of TEM-1 beta-lactamase suggests a role for Glu166 as the general base in acylation. J. Am. Chem. Soc. 124, 5333–5340 (2002).
Urano, Y. et al. Evolution of fluorescein as a platform for finely tunable fluorescence probes. J. Am. Chem. Soc. 127, 4888–4894 (2005).
McNerney, R. & Daley, P. Towards a point-of-care test for active tuberculosis: obstacles and opportunities. Nature Rev. Microbiol. 9, 204–213 (2011).
Backus, K. M., et al. Uptake of unnatural trehalose analogs as a reporter for Mycobacterium tuberculosis. Nature Chem. Biol. 7, 228–235 (2011).
Ioerger, T. R., et al. The non-clonality of drug resistance in Beijing-genotype isolates of Mycobacterium tuberculosis from the Western Cape of South Africa. BMC Genomics 11, 670 (2010).
Kwon, H. H., Tomioka, H. & Saito, H. Distribution and characterization of beta-lactamases of mycobacteria and related organisms. Tuber. Lung Dis. 76, 141–148 (1995).
Majiduddin, F. K., Materon, I. C. & Palzkill, T. G. Molecular analysis of beta-lactamase structure and function. Int. J. Med. Microbiol. 292, 127–137 (2002).
Petrosino, J., Cantu, C. III & Palzkill, T. β-Lactamases: protein evolution in real time. Trends Microbiol. 6, 323–327 (1998).
Hugonnet, J. E. & Blanchard, J. S. Irreversible inhibition of the Mycobacterium tuberculosis beta-lactamase by clavulanate. Biochemistry 46, 11998–12004 (2007).
Tremblay, L. W., Fan, F. & Blanchard, J. S. Biochemical and structural characterization of Mycobacterium tuberculosis beta-lactamase with the carbapenems ertapenem and doripenem. Biochemistry 49, 3766–3773 (2010).
Mitchell, R. S., Kumar, V., Robbins, S. L., Abbas, A. K. & Fausto, N. Robbins Basic Pathology (Saunders/Elsevier, 2007).
McPherson, A. Preparation and Analysis of Protein Crystals (Waverly, 1982).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution – from diffraction images to an initial model in minutes. Acta Crystallogr. D Biol. Crystallogr. 62, 859–866 (2006).
Mccoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Bailey, S. The CCP4 suite – programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
This work was supported by grant 48523 from the Bill and Melinda Gates Foundation, the Welch Foundation grant no. A-0015 and NIH P01 68135 for TB Structural Genomics. We thank Bob Fader in the Scott & White Memorial Hospital (Temple, Texas) for providing sputum samples from cystic fibrosis patients.
Author information
Authors and Affiliations
Contributions
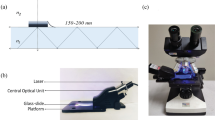
H.X. performed all the compound syntheses and characterizations, collected enzymatic kinetics and carried out the E. coli imaging. J.M. performed the crystallization and structural studies and analysed the data. Y.K., M.H.C. and H.A.H. performed the testing with BCG in human sputum. C.N.T. contributed the imaging box used for cellular phone imaging. H.X., J.M., Y.K., J.C.S., J.D.C. and J.R. conceived and designed the experiments. All authors discussed the results and commented on the manuscript. H.X., J.M., Y.K., J.C.S., J.D.C. and J.R. co-wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 1979 kb)
Rights and permissions
About this article
Cite this article
Xie, H., Mire, J., Kong, Y. et al. Rapid point-of-care detection of the tuberculosis pathogen using a BlaC-specific fluorogenic probe. Nature Chem 4, 802–809 (2012). https://doi.org/10.1038/nchem.1435
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1435
This article is cited by
-
Heterogeneity in M. tuberculosis β-lactamase inhibition by Sulbactam
Nature Communications (2023)
-
Spacer-free BODIPY fluorogens in antimicrobial peptides for direct imaging of fungal infection in human tissue
Nature Communications (2016)
-
Tuberculosis
Nature Reviews Disease Primers (2016)
-
Rapid absolute determination platform of nucleic acid for point-of-care testing
Chemical Research in Chinese Universities (2015)
-
Review and analysis of performance metrics of droplet microfluidics systems
Microfluidics and Nanofluidics (2014)