Abstract
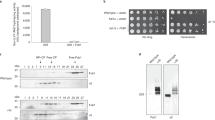
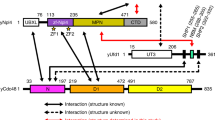
The ubiquitin system recognizes degradation signals of protein substrates through E3–E2 ubiquitin ligases, which produce a substrate-linked multi-ubiquitin chain. Ubiquitinated substrates are degraded by the 26S proteasome, which consists of the 20S protease and two 19S particles. We previously showed that UBR1 and UFD4, two E3 ligases of the yeast Saccharomyces cerevisiae, interact with specific proteasomal subunits. Here we advance this analysis for UFD4 and show that it interacts with RPT4 and RPT6, two subunits of the 19S particle. The 201-residue amino-terminal region of UFD4 is essential for its binding to RPT4 and RPT6. UFD4ΔN, which lacks this N-terminal region, adds ubiquitin to test substrates with apparently wild-type activity, but is impaired in conferring short half-lives on these substrates. We propose that interaction of a targeted substrate with the 26S proteasome involves contacts of specific proteasomal subunits with the substrate-bound ubiquitin ligase, with the substrate-linked multi-ubiquitin chain and with the substrate itself. This multiple-site binding may function to slow down dissociation of the substrate from the proteasome and to facilitate the unfolding of substrate through ATP-dependent movements of the chaperone subunits of the 19S particle.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hershko, A., Ciechanover, A. & Varshavsky, A. Nature Med. 10, 1073–1081 (2000).
Pickart, C. Annu. Rev. Biochem. 70, 503–533 (2001).
Weissman, A. M. Nature Rev. Mol. Cell Biol. 2, 169–178 (2001).
Zwickl, P., Baumeister, W. & Steven, A. Curr. Opin. Struct. Biol. 10, 242–250 (2000).
Russell, S. J., Steger, K. A. & Johnston, S. A. J. Biol. Chem. 274, 21943–21952 (1999).
Rechsteiner, M. in Ubiquitin and the Biology of the Cell (eds Peters, J. M., Harris, J. R. & Finley, D.) 147–189 (Plenum, New York, 1998).
Glickman, M. H., Rubin, D. M., Fried, V. A. & Finley, D. Mol. Cell. Biol. 18, 3149–3162 (1998).
Lam, Y. A., Lawson, T. G., Velayutham, M., Zweier, J. L. & Pickart, C. M. Nature 416, 763–767 (2002).
van Nocker, S. et al. Mol. Cell. Biol. 16, 6020–6028 (1996).
Turner, G. C., Du, F. & Varshavsky, A. Nature 405, 579–583 (2000).
Xie, Y. & Varshavsky, A. Proc. Natl Acad. Sci. USA 97, 2497–2502 (2000).
Kwon, Y. T. et al. Science 297, 96–99 (2002).
Johnson, E. S., Ma, P. C., Ota, I. M. & Varshavsky, A. J. Biol. Chem. 270, 17442–17456 (1995).
Koegl, M. et al. Cell 96, 635–644 (1999).
Verma, R. et al. Mol. Biol. Cell 11, 3425–3439 (2000).
Suzuki, T. & Varshavsky, A. EMBO J. 18, 6017–6026 (1999).
Turner, G. C. & Varshavsky, A. Science 289, 2117–2120 (2000).
Ghislain, M., Dohmen, R. J., Levy, F. & Varshavsky, A. EMBO J. 15, 4884–4899 (1996).
Baker, R. T. & Varshavsky, A. Proc. Natl Acad. Sci. USA 87, 2374–2378 (1991).
Braun, S., Matuschewski, K., Rape, M., Thoms, S. & Jentsch, S. EMBO J. 21, 615–621 (2002).
Meyer, H. H., Shorter, J. G., Seemann, J., Pappin, D. & Warren, G. A. EMBO J. 19, 2181–2192 (2000).
Dai, R. M. & Li, C. C. Nature Cell Biol. 3, 740–744 (2001).
You, J. & Pickart, C. M. J. Biol. Chem. 276, 19871–19878 (2001).
Chen, L. & Madura, K. Mol. Cell. Biol. 22, 4902–4913 (2002).
Bertolaet, B. et al. Nature Struct. Biol. 8, 417–422 (2001).
Hiyama, H. et al. J. Biol. Chem. 274, 28019–28025 (1999).
Kleijnen, M. F. et al. Cell 6, 409–419 (2000).
Farrás, R. et al. EMBO J. 20, 2742–2756 (2001).
Jäger, S., Strayle, J., Heinemeyer, W. & Wolf, D. H. EMBO J. 20, 4423–4431 (2001).
Xie, Y. & Varshavsky, A. Proc. Natl Acad. Sci. USA 98, 3056–3061 (2001).
Acknowledgements
We thank R. Verma and R. Deshaies for the purified 26S proteasome; E. Johnson, D. Botstein and S. Jentsch for S. cerevisiae strains and plasmids; and members of the Varshavsky laboratory, particularly R. Hu and J. Sheng, for assistance and advice. This work was supported by a grant to A.V. from the NIH. Y.X. was supported in part by a postdoctoral fellowship from the NIH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Xie, Y., Varshavsky, A. UFD4 lacking the proteasome-binding region catalyses ubiquitination but is impaired in proteolysis. Nat Cell Biol 4, 1003–1007 (2002). https://doi.org/10.1038/ncb889
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb889
This article is cited by
-
The E3 ubiquitin ligase TRIP12 participates in cell cycle progression and chromosome stability
Scientific Reports (2020)
-
Hul5 HECT ubiquitin ligase plays a major role in the ubiquitylation and turnover of cytosolic misfolded proteins
Nature Cell Biology (2011)
-
Working on a chain: E3s ganging up for ubiquitylation
Nature Cell Biology (2010)
-
The N-end rule pathway is mediated by a complex of the RING-type Ubr1 and HECT-type Ufd4 ubiquitin ligases
Nature Cell Biology (2010)
-
Feedback regulation of proteasome gene expression and its implications in cancer therapy
Cancer and Metastasis Reviews (2010)