Abstract
The mechanisms by which hypoxic tumours evade immunological pressure and anti-tumour immunity remain elusive. Here, we report that two hypoxia-responsive microRNAs, miR-25 and miR-93, are important for establishing an immunosuppressive tumour microenvironment by downregulating expression of the DNA sensor cGAS. Mechanistically, miR-25/93 targets NCOA3, an epigenetic factor that maintains basal levels of cGAS expression, leading to repression of cGAS during hypoxia. This allows hypoxic tumour cells to escape immunological responses induced by damage-associated molecular pattern molecules, specifically the release of mitochondrial DNA. Moreover, restoring cGAS expression results in an anti-tumour immune response. Clinically, decreased levels of cGAS are associated with poor prognosis for patients with breast cancer harbouring high levels of miR-25/93. Together, these data suggest that inactivation of the cGAS pathway plays a critical role in tumour progression, and reveal a direct link between hypoxia-responsive miRNAs and adaptive immune responses to the hypoxic tumour microenvironment, thus unveiling potential new therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Zitvogel, L., Tesniere, A. & Kroemer, G. Cancer despite immunosurveillance: immunoselection and immunosubversion. Nat. Rev. Immunol. 6, 715–727 (2006).
Joyce, J. A. & Pollard, J. W. Microenvironmental regulation of metastasis. Nat. Rev. Cancer 9, 239–252 (2009).
Palazon, A., Aragones, J., Morales-Kastresana, A., de Landazuri, M. O. & Melero, I. Molecular pathways: hypoxia response in immune cells fighting or promoting cancer. Clin. Cancer Res. 18, 1207–1213 (2012).
Noman, M. Z. et al. Microenvironmental hypoxia orchestrating the cell stroma cross talk, tumor progression and antitumor response. Crit. Rev. Immunol. 31, 357–377 (2011).
Lewis, C. & Murdoch, C. Macrophage responses to hypoxia: implications for tumor progression and anti-cancer therapies. Am. J. Pathol. 167, 627–635 (2005).
Corzo, C. A. et al. HIF-1α regulates function and differentiation of myeloid-derived suppressor cells in the tumor microenvironment. J. Exp. Med. 207, 2439–2453 (2010).
Noman, M. Z. et al. Tumor-promoting effects of myeloid-derived suppressor cells are potentiated by hypoxia-induced expression of miR-210. Cancer Res. 75, 3771–3787 (2015).
Facciabene, A. et al. Tumour hypoxia promotes tolerance and angiogenesis via CCL28 and Treg cells. Nature 475, 226–230 (2011).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
O’Connell, R. M., Rao, D. S., Chaudhuri, A. A. & Baltimore, D. Physiological and pathological roles for microRNAs in the immune system. Nat. Rev. Immunol. 10, 111–122 (2010).
Esquela-Kerscher, A. & Slack, F. J. Oncomirs—microRNAs with a role in cancer. Nat. Rev. Cancer 6, 259–269 (2006).
Shen, J. et al. EGFR modulates microRNA maturation in response to hypoxia through phosphorylation of AGO2. Nature 497, 383–387 (2013).
Kulshreshtha, R., Davuluri, R. V., Calin, G. A. & Ivan, M. A microRNA component of the hypoxic response. Cell Death Differ. 15, 667–671 (2008).
Noman, M. Z. et al. The cooperative induction of hypoxia-inducible factor-1 alpha and STAT3 during hypoxia induced an impairment of tumor susceptibility to CTL-mediated cell lysis. J. Immunol. 182, 3510–3521 (2009).
Vesely, M. D., Kershaw, M. H., Schreiber, R. D. & Smyth, M. J. Natural innate and adaptive immunity to cancer. Annu. Rev. Immunol. 29, 235–271 (2011).
Yang, M. H. et al. Direct regulation of TWIST by HIF-1α promotes metastasis. Nat. Cell Biol. 10, 295–305 (2008).
Clambey, E. T. et al. Hypoxia-inducible factor-1 alpha-dependent induction of FoxP3 drives regulatory T-cell abundance and function during inflammatory hypoxia of the mucosa. Proc. Natl Acad. Sci. USA 109, E2784–E2793 (2012).
Wu, M. Z. et al. Hypoxia drives breast tumor malignancy through a TET-TNFα-p38-MAPK signaling axis. Cancer Res. 75, 3912–3924 (2015).
Petrocca, F., Vecchione, A. & Croce, C. M. Emerging role of miR-106b-25/miR-17-92 clusters in the control of transforming growth factor beta signaling. Cancer Res. 68, 8191–8194 (2008).
van den Beucken, T. et al. Hypoxia promotes stem cell phenotypes and poor prognosis through epigenetic regulation of DICER. Nat. Commun. 5, 5203 (2014).
Mariani, C. J. et al. TET1-mediated hydroxymethylation facilitates hypoxic gene induction in neuroblastoma. Cell Rep. 7, 1343–1352 (2014).
Gao, D. et al. Cyclic GMP-AMP synthase is an innate immune sensor of HIV and other retroviruses. Science 341, 903–906 (2013).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Matzinger, P. The danger model: a renewed sense of self. Science 296, 301–305 (2002).
Liu, Y. et al. Hypoxia induced HMGB1 and mitochondrial DNA interactions mediate tumor growth in hepatocellular carcinoma through Toll-like receptor 9. J. Hepatol. 63, 114–121 (2015).
Rongvaux, A. et al. Apoptotic caspases prevent the induction of type I interferons by mitochondrial DNA. Cell 159, 1563–1577 (2014).
White, M. J. et al. Apoptotic caspases suppress mtDNA-induced STING-mediated type I IFN production. Cell 159, 1549–1562 (2014).
Liu, L. et al. Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat. Cell Biol. 14, 177–185 (2012).
Kim, H. et al. Fine-tuning of Drp1/Fis1 availability by AKAP121/Siah2 regulates mitochondrial adaptation to hypoxia. Mol. Cell 44, 532–544 (2011).
Deng, L. et al. STING-dependent cytosolic DNA sensing promotes radiation-induced type I interferon-dependent antitumor immunity in immunogenic tumors. Immunity 41, 843–852 (2014).
Yan, J. et al. Steroid receptor coactivator-3 and activator protein-1 coordinately regulate the transcription of components of the insulin-like growth factor/AKT signaling pathway. Cancer Res. 66, 11039–11046 (2006).
Chen, H. et al. Nuclear receptor coactivator ACTR is a novel histone acetyltransferase and forms a multimeric activation complex with P/CAF and CBP/p300. Cell 90, 569–580 (1997).
Percharde, M. et al. Ncoa3 functions as an essential Esrrb coactivator to sustain embryonic stem cell self-renewal and reprogramming. Genes Dev. 26, 2286–2298 (2012).
Kulshreshtha, R. et al. A microRNA signature of hypoxia. Mol. Cell. Biol. 27, 1859–1867 (2007).
Hazarika, S. et al. MicroRNA-93 controls perfusion recovery after hindlimb ischemia by modulating expression of multiple genes in the cell cycle pathway. Circulation 127, 1818–1828 (2013).
Hu, J. et al. MiR-215 is induced post-transcriptionally via HIF-Drosha complex and mediates glioma-initiating cell adaptation to hypoxia by targeting KDM1B. Cancer Cell 29, 49–60 (2016).
Haldar, S., Roy, A. & Banerjee, S. Differential regulation of MCM7 and its intronic miRNA cluster miR-106b-25 during megakaryopoiesis induced polyploidy. RNA Biol. 11, 1137–1147 (2014).
Ramalingam, P. et al. Biogenesis of intronic miRNAs located in clusters by independent transcription and alternative splicing. RNA 20, 76–87 (2014).
Camps, C. et al. Integrated analysis of microRNA and mRNA expression and association with HIF binding reveals the complexity of microRNA expression regulation under hypoxia. Mol. Cancer 13, 28 (2014).
Petrocca, F. et al. E2F1-regulated microRNAs impair TGFβ-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell 13, 272–286 (2008).
Corrales, L. & Gajewski, T. F. Molecular pathways: targeting the stimulator of interferon genes (STING) in the immunotherapy of cancer. Clin. Cancer Res. 21, 4774–4779 (2015).
Matzinger, P. Tolerance, danger, and the extended family. Annu. Rev. Immunol. 12, 991–1045 (1994).
Zhang, Q. et al. Circulating mitochondrial DAMPs cause inflammatory responses to injury. Nature 464, 104–107 (2010).
Wu, M. Z. et al. Interplay between HDAC3 and WDR5 is essential for hypoxia-induced epithelial-mesenchymal transition. Mol. Cell 43, 811–822 (2011).
Lu, L. F. et al. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell 142, 914–929 (2010).
Acknowledgements
We are grateful to M. C. Ku of the Next Generation Sequencing Core, and C. Benner and M. Shokhirev of the Integrative Genomics and Bioinformatics Core, at Salk Institute for Biological Studies for technical assistance. We would also like to thank D. O’Keefe for his help in preparing the manuscript. This work was supported by The Razavi Newman Integrative Genomics and Bioinformatics Core Facility and the NGS Core Facility of the Salk institute with funding from NIH-NCI CCSG: P30014195, the Chapman Foundation and the Helmsley Charitable Trust. Work in the laboratory of J.C.I.B. was supported by the G. Harold and Leila Y. Mathers Charitable Foundation, The Leona M. and Harry B. Helmsley Charitable Trust (2012-PG-MED002), Fundacion Dr. Pedro Guillen, The Moxie Foundation and UCAM.
Author information
Authors and Affiliations
Contributions
M.-Z.W., L.-F.L. and J.C.I.B. conceived and designed the project. W.-C.C. and K.-J.W. performed TCGA analysis with help from L.M. S.-F.C., S.N. and C.-L.L. performed studies on tissue microarrays of human patient samples with help from Y.-S.L. C.O’C. and C.-J.W. assisted in FACS analysis and samples preparation. W.-W.T. contributed to ChIP experiments as well as discussion and revision of the manuscript. M.-Z.W., L.-F.L. and J.C.I.B. wrote the manuscript with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Immunosuppressive phenotypes in HIF-1α overexpressing tumors.
(a) FACS analysis for MDSCs showed the effect of HIF-1α signaling on accumulation of CD11b+ Ly6G−Ly6Chigh monocytic MDSCs within tumors. E0771 cells expressing HIF-1α or empty vector were used for tumor formation. (b,c) The population of CD11b+ F4/80+ TAMs or CD11b+ Ly6G+ Ly6Clow granulocytic MDSCs was not affected by ectopic expression of HIF-1α. (d,e) The immune cell profile for tumor infiltrating CD4+ T cells or CD8+ T cells in HIF-1α overexpressing tumors vs. control tumors. For control, n = 5 tumors from 5 different animals. For HIF-1α, n = 7 tumors from 7 different animals. Samples were compared using two-tailed Student’s t test. ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001. ns, not significant. All data shown in this figure represent the mean of the above- indicated sample numbers.
Supplementary Figure 2 Identification of hypoxia responsive miRNAs associated with tumor progression.
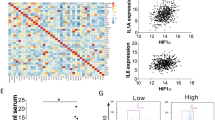
(a) Heat map of small RNA sequencing results from MCF7 cells treated with normoxia or hypoxia, MCF7 cells overexpressing HIF-1α, and MCF7-derived BTIC cells. (b) The correlation of selected hypoxia responsive miRNAs with breast tumor progression. Red, molecule is up-regulated. Green, molecule is down-regulated. The intensity of green and red molecule color indicates the degree of down-regulation and up-regulation, respectively. Font bold in red indicates the focus molecule. (c) The correlation of selected hypoxia responsive miRNAs with certain key cancer signaling pathways. (d) The expression level of miR106b-25 cluster in normoxic, hypoxic, HIF-1α overexpressing, and BTIC cells. MCF7 and MDA-MB-231 cells were used as models, as indicated. (e) Knockdown of HIF-1α abolished hypoxia effect on induction of miR25/93 in either hypoxic MCF7 (left upper) or MDA-MB-231 cells (left lower). The protein levels for HIF-1α were presented in right panel. The cells expressing shRNA targeting EGFP were used as control. All the data in the graphs are presented as means ± SD (n = 3 independent experiments). Samples were compared using two-tailed Student’s t test. Asterisk indicates P < 0.05, compared with controls. The immunoblotting shown in panel e are representative of three independent experiments. Unprocessed scans of western blot analysis is available in Supplementary Figure 9. Statistics source data is available in Supplementary Table 4.
Supplementary Figure 3 The immune cell profile in miR25/93 overexpression tumors.
(a) The levels for miR25/93 in either control tumors or miR25/93 overexpressing tumors. (b) FACS analysis showed accumulation of CD11b+ Ly6G− Ly6Chigh monocytic MDSC in miR25/93 overexpressing tumors, as compared with control tumors. E0771 cells were used for generating miR25/93 overexpressing tumors. (c and d) The percentage of CD11b+ F4/80+ TAM or CD11b+ Ly6G+ Ly6Clow granulocytic MDSC inmiR25/93 overexpressing tumors vs. control tumors.(e and f)The immune cell profile for tumor infiltrating CD4+ T cells or CD8+ T cells inmiR25/93 overexpressing tumors vs. control tumors. (g) Gene expression analysis in whole tumor specimen for selected genes responsible for anti-tumor immunity. (h) The elevated levels of tumor promoting factors in miR25/93 overexpressing tumors as compared to control tumors. For panel a, the data are presented as means ± SD (n = 3 independent experiments). Asterisk indicates P < 0.05, compared with controls. For panels b–d, control: n = 4 tumors from 4 different animals, miR25/93: n = 4 tumors from 4 different animals. For panels e,f, control: n = 10 tumors from 10 different animals, miR25/93: n = 13 tumors from 13 different animals. For panels g,h, n = 8 tumors from 8 different animals. Data shown in b–h represent the mean of the above-indicated sample numbers. Samples were compared using two-tailed Student’s t test. ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, ns not significant.
Supplementary Figure 4 The regulation of miR25 and miR93 by TET1 protein-mediated epigenetic change in response to hypoxia.
(a) The binding of HIF-1α to the VEGF gene promoter and miR25/93genome regions. VEGF gene promoter was used a positive control for HIF-1α binding upon hypoxia. Four sets of primers targeting genome regions of miR25/93 were used, as indicated. MCF7 and MDA-MB-231 cells were used, as indicated. (b) Levels of 5hmC were locally increased in miR25/93 genomic regions in response to hypoxia. (c) hMeDIP-qPCR validation of the 5hmC change in the miR25/93 genomic regions upon hypoxia. (d) ChIP analysis showed an increase in binding of TET1, but not TET3, to the miR25/93 genomic regions in response to hypoxia (left). TNF genome region was used as a positive control for either TET1 or TET3 (right). (e) qPCR analysis of TET1 knockdown cells for the expression of miR25-3p, miR93-5p, and miR93-3p upon hypoxia. (f) Western bolt analysis for TET1 level in TET1 knockdown cells. (g) The deficiency of TET3 did not alter the ability of hypoxia to induce miR25 and miR93 expression. (h) Western bolt analysis for TET3 in TET3 knockdown cells (i) In TET1 knockdown cells, hMeDIP-qPCR analysis showed that 5hmC levelson the miR25/93 genomic regions were restored in response to hypoxia by re-expressing wild type TET1, but not a TET1 inactive mutant. As representation, the third primer set out of four primer sets was used in qPCR analysis. The cells expressing shRNA targeting EGFP were used as control in panels d to h. N: normoxia H: hypoxia. All data are presented as means ± SD (n = 3 independent experiments). Samples were compared using two-tailed Student’s t test. Asterisk indicates P < 0.05, compared with controls. The image shown in panel f,h is representative of three independent experiments. Unprocessed scans of western blot analysis is available in Supplementary Figure 9. Statistics source data is available in Supplementary Table 4.
Supplementary Figure 5 Hypoxia reduces immune response to cytosolic mtDNA.
(a) Hypoxia increased the level of cytosolic mtDNA. Upper: Western blot analysis for specific markers in cytosolic (β-actin), nucleus (Histone H3), and mitochondria (cytochrome c) fractions. Nucleus is referred to the fraction containing both nucleus and mitochondria fraction. Lower: qPCR analysis of cytosolic mtDNA in either hypoxic MCF7 or MDA-MB-231 cells using three sets of primers targeting different mitochondria DNA regions, as indicated. (b) Hypoxia induced mitophagy, as shown by accumulated level of LC-II and decreased level of p62. (c) Hypoxia suppressed mtDNA-induced immune response in either MCF7 (upper) or MDA-MB-231 cells (lower). The IFN-β expression was used as an indicator for mtDNA-induced immune response. (d,e) Loss of function of miR25/93 restored immune response in the presence of cytosolic mtDNA upon hypoxia. (f,g) The analysis of IFN-β expression in cGAS deficient cells indicated cGAS as a major mediator in mtDNA-induced immune response. The cells expressing shRNA targeting EGFP were used as control. N: normoxia H: hypoxia. All the data in graphs are presented as means ± SD (n = 3 independent experiments). Samples were compared using two-tailed Student’s t test. Asterisk indicates P < 0.05. The image shown in panel a,b is representative of three independent experiments. Unprocessed scans of western blot analysis is available in Supplementary Figure 9. Statistics source data is available in Supplementary Table 4.
Supplementary Figure 6 Inhibition of miR25/93 or cGAS is unable to affect cytoplasmic level of mtDNA as a response to hypoxia.
(a) qPCR analysis for cytosolic mtDNA level in MCF7 or MDA-MB-231stable transfectants expressing shRNA against miR25/93 vs. control cells. (b) Knockdown of cGAS was unable to affect the hypoxic effect on increasing the level of cytosolic mtDNA. The cells expressing shRNA targeting EGFP were used as control. N: normoxia H: hypoxia. All the data are presented as means ± SD (n = 3 independent experiments). Samples were compared using two-tailed Student’s t test. Asterisk indicates P < 0.05, compared with controls.
Supplementary Figure 7 Down-regulation of cGAS leads to establishment of immunosuppressive phenotypes.
(a) Tumor growth analysis showed increased proliferation rate of cGAS deficient tumors compared to control tumors. E0771 cells were used for generating tumors with cGAS deficiency. (b–d) FACS analysis for CD11b+ Ly6G− Ly6Chigh monocytic MDSC, CD11b+ F4/80+ TAM, and CD11b+ Ly6G+ Ly6Clow granulocytic MDSC in cGAS down-regulated tumors vs. control tumors. (e,f) The immune cell profile for tumor infiltrating CD4+ T cells or CD8+ T cells in cGAS deficient tumors vs. control tumors. (g) Knockdown of cGAS decreased the percentage of effector CD8+ cytotoxic T cells. Changes in effector CD8+ cytotoxic T cells normalized to total CD8 + T cell proportions. (h) Increased level of Treg cells in cGAS deficient tumors, as compared to control tumors. (i) Gene expression analysis of cGAS deficient tumors showed significantly lower level of anti-tumor immunity factors than control tumors. (j) Increased level of tumor promoting factors in cGAS down-regulated tumors vs. control tumors. (k) cGAS expression in tumor biopsies derived from cGAS deficient cells or control cells. The cells expressing shRNA targeting EGFP were used as control in this figure. For panels b–d, n = 4 tumors from 4 different animals. For panels e–h, control: n = 7 tumors from 7 different animals, cGAS-si: n = 10 tumors from 10 different animals. For panels i–k, n = 8 tumors from 8 different animals. Data shown in panel a are means ± SD of n = 8 tumors from 8 different animals mice for each group. For panel g, data is presented as mean ± SD of indicated sample numbers. For panels b–f and h–k, data represent the mean of the above-indicated sample numbers. Samples were compared using two-tailed Student’s t test. ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001. ns, not significant.
Supplementary Figure 8 TCGA analysis for Bim and PTEN in breast cancer patients harboring high level of miR93.
(a,b) The levels of Bim and PTEN were not correlated with overall survival of breast cancer patients bearing high level of miR93. Sample number and p value were indicated in the panels. The prognostic significance was assessed by Kaplan–Meier analysis. (c) The expression of MCM7 gene was down-regulated by hypoxia. Knocking down HIF-1α partially rescued MCM7 level in hypoxia. The data is presented as means ± SD (n = 3 independent experiments). Samples were compared using two-tailed Student’s t test. Asterisk indicates P < 0.05.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1099 kb)
Supplementary Table 1
Supplementary Information (XLSX 42 kb)
Supplementary Table 2
Supplementary Information (XLSX 623 kb)
Supplementary Table 3
Supplementary Information (XLSX 12 kb)
Supplementary Table 4
Supplementary Information (XLSX 73 kb)
Rights and permissions
About this article
Cite this article
Wu, MZ., Cheng, WC., Chen, SF. et al. miR-25/93 mediates hypoxia-induced immunosuppression by repressing cGAS. Nat Cell Biol 19, 1286–1296 (2017). https://doi.org/10.1038/ncb3615
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3615
This article is cited by
-
Hypoxic glioblastoma-cell-derived extracellular vesicles impair cGAS-STING activity in macrophages
Cell Communication and Signaling (2024)
-
cGAS and cancer therapy: a double-edged sword
Acta Pharmacologica Sinica (2022)
-
miR-25 and miR-92b regulate insulin biosynthesis and pancreatic β-cell apoptosis
Endocrine (2022)
-
Cytosolic DNA sensing by cGAS: regulation, function, and human diseases
Signal Transduction and Targeted Therapy (2021)
-
miR-23a/b suppress cGAS-mediated innate and autoimmunity
Cellular & Molecular Immunology (2021)