Abstract
Animal cell shape is largely determined by the cortex, a thin actin network underlying the plasma membrane in which myosin-driven stresses generate contractile tension. Tension gradients result in local contractions and drive cell deformations. Previous cortical tension regulation studies have focused on myosin motors. Here, we show that cortical actin network architecture is equally important. First, we observe that actin cortex thickness and tension are inversely correlated during cell-cycle progression. We then show that the actin filament length regulators CFL1, CAPZB and DIAPH1 regulate mitotic cortex thickness and find that both increasing and decreasing thickness decreases tension in mitosis. This suggests that the mitotic cortex is poised close to a tension maximum. Finally, using a computational model, we identify a physical mechanism by which maximum tension is achieved at intermediate actin filament lengths. Our results indicate that actin network architecture, alongside myosin activity, is key to cell surface tension regulation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Salbreux, G., Charras, G. & Paluch, E. Actin cortex mechanics and cellular morphogenesis. Trends Cell Biol. 22, 536–545 (2012).
Bray, D. & White, J. G. Cortical flow in animal cells. Science 239, 883–888 (1988).
Clark, A. G. & Paluch, E. in Cell Cycle in Development Vol. 1 (ed. Kubiak, J. Z.) 31–73 (Springer, 2011).
Heisenberg, C. P. & Bellaiche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962 (2013).
Levayer, R. & Lecuit, T. Biomechanical regulation of contractility: spatial control and dynamics. Trends Cell Biol. 22, 61–81 (2012).
Maddox, A. S. & Burridge, K. RhoA is required for cortical retraction and rigidity during mitotic cell rounding. J. Cell Biol. 160, 255–265 (2003).
Matzke, R., Jacobson, K. & Radmacher, M. Direct, high-resolution measurement of furrow stiffening during division of adherent cells. Nat. Cell Biol. 3, 607–610 (2001).
Sedzinski, J. et al. Polar actomyosin contractility destabilizes the position of the cytokinetic furrow. Nature 476, 462–466 (2011).
Stewart, M. P. et al. Hydrostatic pressure and the actomyosin cortex drive mitotic cell rounding. Nature 469, 226–230 (2011).
Bergert, M. et al. Force transmission during adhesion-independent migration. Nat. Cell Biol. 17, 524–529 (2015).
Vicente-Manzanares, M., Ma, X., Adelstein, R. S. & Horwitz, A. R. Non-muscle myosin II takes centre stage in cell adhesion and migration. Nat. Rev. Mol. Cell Biol. 10, 778–790 (2009).
Clark, A. G., Wartlick, O., Salbreux, G. & Paluch, E. K. Stresses at the cell surface during animal cell morphogenesis. Curr. Biol. 24, R484–R494 (2014).
Tinevez, J.-Y. et al. Role of cortical tension in bleb growth. Proc. Natl Acad. Sci. USA 106, 18581–18586 (2009).
Ramanathan, S. P. et al. Cdk1-dependent mitotic enrichment of cortical myosin II promotes cell rounding against confinement. Nat. Cell Biol. 17, 148–159 (2015).
Mayer, M., Depken, M., Bois, J. S., Julicher, F. & Grill, S. W. Anisotropies in cortical tension reveal the physical basis of polarizing cortical flows. Nature 467, 617–621 (2010).
Turlier, H., Audoly, B., Prost, J. & Joanny, J. F. Furrow constriction in animal cell cytokinesis. Biophys. J. 106, 114–123 (2014).
Salbreux, G., Joanny, J. F., Prost, J. & Pullarkat, P. Shape oscillations of non-adhering fibroblast cells. Phys. Biol. 4, 268–284 (2007).
Ennomani, H. et al. Architecture and connectivity govern actin network contractility. Curr. Biol. 26, 616–626 (2016).
Murrell, M. P. & Gardel, M. L. F-actin buckling coordinates contractility and severing in a biomimetic actomyosin cortex. Proc. Natl Acad. Sci. USA 109, 20820–20825 (2012).
Reymann, A.-C. et al. Actin network architecture can determine myosin motor activity. Science 336, 1310–1314 (2012).
Alvarado, J., Sheinman, M., Sharma, A., MacKintosh, F. C. & Koenderink, G. H. Molecular motors robustly drive active gels to a critically connected state. Nat. Phys. 9, 591–597 (2013).
Hanakam, F., Albrecht, R., Eckerskorn, C., Matzner, M. & Gerisch, G. Myristoylated and non-myristoylated forms of the pH sensor protein hisactophilin II: intracellular shuttling to plasma membrane and nucleus monitored in real time by a fusion with green fluorescent protein. EMBO J. 15, 2935–2943 (1996).
Charras, G. T., Hu, C.-K., Coughlin, M. & Mitchison, T. J. Reassembly of contractile actin cortex in cell blebs. J. Cell Biol. 175, 477–490 (2006).
Clark, A. G., Dierkes, K. & Paluch, E. K. Monitoring actin cortex thickness in live cells. Biophys. J. 105, 570–580 (2013).
Fischer-Friedrich, E., Hyman, A. A., Jülicher, F., Müller, D. J. & Helenius, J. Quantification of surface tension and internal pressure generated by single mitotic cells. Sci. Rep. 4, 6213 (2014).
Kunda, P., Pelling, A. E., Liu, T. & Baum, B. Moesin controls cortical rigidity, cell rounding, and spindle morphogenesis during mitosis. Curr. Biol. 18, 91–101 (2008).
Matthews, H. K. et al. Changes in ect2 localization couple actomyosin-dependent cell shape changes to mitotic progression. Dev. Cell 23, 371–383 (2012).
Zlotek-Zlotkiewicz, E., Monnier, S., Cappello, G., Le Berre, M. & Piel, M. Optical volume and mass measurements show that mammalian cells swell during mitosis. J. Cell Biol. 211, 765–774 (2015).
Carreno, S. et al. Moesin and its activating kinase Slik are required for cortical stability and microtubule organization in mitotic cells. J. Cell Biol. 180, 739–746 (2008).
Machicoane, M. et al. SLK-dependent activation of ERMs controls LGN-NuMA localization and spindle orientation. J. Cell Biol. 205, 791–799 (2014).
Cooper, J. A. & Sept, D. New insights into mechanism and regulation of actin capping protein. Int. Rev. Cell Mol. Biol. 267, 183–206 (2008).
Carlier, M. F. et al. Actin depolymerizing factor (ADF/cofilin) enhances the rate of filament turnover: implication in actin-based motility. J. Cell Biol. 136, 1307–1322 (1997).
Bamburg, J. R. Proteins of the ADF/cofilin family: essential regulators of actin dynamics. Annu. Rev. Cell Dev. Biol. 15, 185–230 (1999).
Bovellan, M. et al. Cellular control of cortical actin nucleation. Curr. Biol. 24, 1628–1635 (2014).
Rosa, A., Vlassaks, E., Pichaud, F. & Baum, B. Ect2/Pbl acts via Rho and polarity proteins to direct the assembly of an isotropic actomyosin cortex upon mitotic entry. Dev. Cell 32, 604–616 (2015).
Campellone, K. G. & Welch, M. D. A nucleator arms race: cellular control of actin assembly. Nat. Rev. Mol. Cell Biol. 11, 237–251 (2010).
Diz-Munoz, A. et al. Control of directed cell migration in vivo by membrane-to-cortex attachment. PLoS Biol. 8, e1000544 (2010).
Rouven Bruckner, B., Pietuch, A., Nehls, S., Rother, J. & Janshoff, A. Ezrin is a major regulator of membrane tension in epithelial cells. Sci. Rep. 5, 14700 (2015).
Kim, T., Cooper, J. A. & Sept, D. The interaction of capping protein with the barbed end of the actin filament. J. Mol. Biol. 404, 794–802 (2010).
Bombardier, J. P. et al. Single-molecule visualization of a formin-capping protein ‘decision complex’ at the actin filament barbed end. Nat. Commun. 6, 8707 (2015).
Kovar, D. R., Harris, E. S., Mahaffy, R., Higgs, H. N. & Pollard, T. D. Control of the assembly of ATP- and ADP-actin by formins and profilin. Cell 124, 423–435 (2006).
Hiraiwa, T. & Salbreux, G. Role of turnover in active stress generation in a filament network. Phys. Rev. Lett. 116, 188101 (2016).
Dasanayake, N. L., Michalski, P. J. & Carlsson, A. E. General mechanism of actomyosin contractility. Phys. Rev. Lett. 107, 118101 (2011).
Lenz, M., Thoresen, T., Gardel, M. L. & Dinner, A. R. Contractile units in disordered actomyosin bundles arise from F-actin buckling. Phys. Rev. Lett. 108, 238107 (2012).
Soares e Silva, M. et al. Active multistage coarsening of actin networks driven by myosin motors. Proc. Natl Acad. Sci. USA 108, 9408–9413 (2011).
Biro, M. et al. Cell cortex composition and homeostasis resolved by integrating proteomics and quantitative imaging. Cytoskeleton 70, 741–754 (2013).
Stam, S., Alberts, J., Gardel, M. L. & Munro, E. Isoforms confer characteristic force generation and mechanosensation by myosin II filaments. Biophys. J. 108, 1997–2006 (2015).
Pagliara, S. et al. Auxetic nuclei in embryonic stem cells exiting pluripotency. Nat. Mater. 13, 638–644 (2014).
Skoufias, D. A. et al. S-trityl-L-cysteine is a reversible, tight binding inhibitor of the human kinesin Eg5 that specifically blocks mitotic progression. J. Biol. Chem. 281, 17559–17569 (2006).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat Methods 9, 676–682 (2012).
Stewart, M. P., Toyoda, Y., Hyman, A. A. & Muller, D. J. Tracking mechanics and volume of globular cells with atomic force microscopy using a constant-height clamp. Nat. Protoc. 7, 143–154 (2012).
Hell, S., Reiner, G., Cremer, C. & Stelzer, E. H. K. Aberrations in confocal fluorescence microscopy induced by mismatches in refractive index. J. Microsc. 169, 391–405 (1993).
Diaspro, A., Federici, F. & Robello, M. Influence of refractive-index mismatch in high-resolution three-dimensional confocal microscopy. Appl. Opt. 41, 685–690 (2002).
Tukey, J. W. Exploratory Data Analysis (Addison-Wesely, 1977).
Iglewicz, B. & Hoaglin, D. C. How to Detect and Handle Outliers (ASQC Quality Press, 1993).
NIST/SEMATECH e-Handbook of Statistical Methods (eds Croarkin, C. & Tobias, P.) (2010); http://www.itl.nist.gov/div898/handbook
Acknowledgements
We thank B. Baum, K. Chalut, A. Jégou, G. Romet-Lemonne and members of the Paluch laboratory, particularly A. Chaigne, M. Serres and B. A. Truong Quang, for discussions and comments on the manuscript. We thank A. Yonis and M. Turmaine for technical help with SEM, R. Henriques for advice on AFM analysis, C. Verstreken (K. Chalut lab) for preparing the mESC cells used for cortex thickness measurements and the MRC-LMCB imaging facility for technical support. We acknowledge support from the Medical Research Council UK (MRC programme award MC_UU_12018/5), the Human Frontier Science Program (Young Investigator Grant to G.C., P.P.R. and E.K.P.), the European Research Council (starting grant 311637-MorphoCorDiv to E.K.P.) and the Polish Ministry of Science and Higher Education (grant 454/N-MPG/2009/0 to E.K.P.). We also acknowledge the International Institute of Molecular and Cell Biology in Warsaw, the Max Planck Society and the Canadian Institutes of Health Research (CIHR Grant MOP123408 to P.P.R.). K.D. has received funding from the Spanish Ministry of Economy and Competitiveness, Plan Nacional, BFU2010-16546 and Centro de Excelencia Severo Ochoa 2013–2017, SEV-2012-0208, as well as the CERCA Programme/Generalitat de Catalunya. P.P.R. holds a Canada Research Chair in Cell Signaling and Proteomics, and a Chercheur-boursier senior award from the Fonds de Recherche Santé Québec (FRQS). G.S. is supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001317), the UK Medical Research Council (FC001317) and the Wellcome Trust (FC001317).
Author information
Authors and Affiliations
Contributions
P.C., A.G.C., M.B.S., G.S. and E.K.P. designed the research and wrote the paper; P.C. carried out most of the experiments and image analysis; A.G.C. carried out some cortical thickness measurements and developed analysis tools; P.C. and A.G.C. designed the experiments and analysed the data; D.A.D.C. carried out the AFM experiments; K.D. designed the image analysis and linescan extraction software; A.R. carried out the myosin western blot experiments; M.B.S. and G.S. designed the computational model; M.B.S. carried out simulations and wrote the SEM picture analysis plugin; G.C. and P.P.R. provided technical support and conceptual advice. All authors discussed the results and manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cortex tension measurements using tipless cantilever atomic force microscopy (AFM).
(a) Representative YZ image of a mitotic adherent HeLa cell with plasma membrane labelled using CellMask Deep Red (magenta) and cantilever visualized by laser reflection (green). Scale bar, 5 μm. (b) Schematic of tipless cantilever AFM assay. rmid: radius of the maximum cross sectional area of the cell; rc: radius of the contact area of the cell with the cantilever; and hcell: height of the cell. (c) AFM force curve from a cell compression experiment. Red arrow indicates the time point after initial force relaxation, at which cell height is recorded. Confocal stack acquisition for cell height recording induces force fluctuations (not displayed), therefore force is measured before the confocal stack is acquired. (d) Calculation of the correction coefficient for refractive index mismatches. Plot of mES cell height values directly measured by AFM versus those extracted from confocal stacks of mESCs with the membrane labelled (peak-to-peak distance). The slope calculated from linear fitting was used to correct for optical aberrations due to refractive index mismatch in cell height measurements, see Methods for details. Points represent individual measurements (n = 30 cells from 3 independent experiments).
Supplementary Figure 2 Extended measurements of cortex properties in interphase and mitosis.
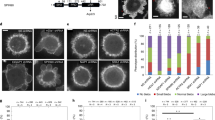
(a,b) Full width at half-maximum (FWHM) of membrane linescans (P = 0.050, 0.16, 0.025, 0.82) (a) and cell radii (P = 9.8 × 10−13, 7.2 × 10−6, 0.0062, 0.048) (b) for different cell lines in interphase and mitosis (n = 41, 100, 47, 27, 11, 13, 22, 13 individual measurements from 2–13 independent experiments). (c) Cortex thickness measurements for interphase HeLa cells detached using trypsin or enzyme-free detachment (Enz.-Free) and immobilized by centrifugation on Poly-L-Lysine (PLL)-coated dishes or by confining the cells in 25 μm microchannels (Channels; n = 41, 7, 8 individual measurements from 1-3 independent experiments; P = 0.11, 0.98). (d) Cortex thickness measurements for HeLa cells blocked in prometaphase with S-Trityl-L-cysteine (STC-Blocked) and unblocked mitotic HeLa cells identified on the basis of morphology and chromosome appearance (n = 100, 16 individual measurements from 13 (STC-blocked) and 1 (unblocked) independent experiments; P = 0.2565). (e) Left: calculation of average cortical actin density using the cortical thickness extraction method (ref. 24 in the main text). Right: calculation of total cortical actin amount using the measured cell radius (r) and cortex thickness (h). (f,g) Relative cortex density (P = 1.2 × 10−6, 0.018, 0.032, 0.0023) (f) and amount (P = 0.47, 0.66, 0.40, 0.53) (g) for different cell lines in interphase and mitosis. Values were normalized by dividing by the median of the interphase value for each cell line (n = 41, 100, 47, 27, 11, 13, 22, 13 individual cell measurements from 2–13 independent experiments). (h) Example segmentation mask (bottom) applied to a scanning electron micrograph of membrane-extracted cortices (top, from Fig. 1g). Images are representative of 17 regions from 8 cells (2 independent experiments). Scale bars, 100 nm. (i) Frequency distribution of pore sizes for adherent Hela and S-HeLa cells in interphase and mitosis (n = 14, 19, 18, 17 regions from 9, 13, 8, 8 cells from 2 independent experiments). Size distributions were comparable between interphase and mitosis, with slightly more large pores in interphase compared to metaphase (inset). (j) Pore coverage (%) in HeLa and S-HeLa cells in interphase and mitosis from regions analysed in (i) (P = 4.3 × 10−6, 0.59). The average pore coverage was unchanged in S-HeLa cells and slightly higher in mitosis in HeLa cells. However, SEM only probes the outer surface of the cortex; it is thus difficult to draw conclusions about overall cortical actin density from this analysis. For all panels, Welch’s t-test P-values: ns P > 0.05, ∗P < 0.05, ∗∗P < 0.01, ∗∗∗P < 0.001.
Supplementary Figure 3 Extended measurements of cortex properties upon depletion of cortex thickness regulators.
(a) Cortex thickness measurements for mitotic HeLa cells treated with the myosin II inhibitor Blebbistatin (Blebb.), the Arp2/3 inhibitor CK-666 and corresponding DMSO controls (n = 12, 16, 27, 33 individual measurements from two or three independent experiments; P = 0.078, 0.47). (b) Cortex tension measurements following treatment with the actin stabilizing drug jasplakinolide (Jas.), Blebbistatin (Blebb.) and corresponding DMSO controls (n = 13, 16, 21, 38 individual measurements from 3 independent experiments; P = 8.1 × 10−5, 3.4 × 10−7). (c,d) Box plot comparing the full width at half maximum (FWHM) of the membrane linescan (P = 0.94, 0.20, 0.0065) (c) and cell radii (P = 0.19, 0.084, 0.31) (d) in mitotic HeLa cells following siRNA against CAPZB, CFL1and DIAPH1 (n = 38, 17, 40, 44, 20, 20 individual measurements from 3–4 independent experiments). The membrane FWHM was ∼10–15% smaller in DIAPH1 knockdown cells, not enough to account for the ∼60% reduction in cortex thickness. (e) Scatter plot of cortex thickness versus cell radius for all measured conditions in mitotic HeLa cells (untreated, target knock-downs and corresponding scrambled controls [Scr.]). (f,g) Relative cortex density (P = 0.23, 0.0008, 0.059) (f) and amount (P = 1.4 × 10−5, 3.6 × 10−7, 0.016) (g) in mitotic HeLa cells treated with siRNA against CAPZB, CFL1and DIAPH1 (Target) and corresponding scrambled controls (Scr.). Values were normalized by dividing by the median of the corresponding Scr. control (n = 38, 17, 40, 44, 20, 20 individual measurements from 3-4 independent experiments). Cortex density and amount were calculated as described in Supplementary Fig. 2e. (h) Western blot showing the levels of active myosin light chain (pMLC2-Ser19) and total myosin light chain (MLC2) in whole cell lysates of mitotic HeLa cells treated with siRNA against CAPZB, DIAPH1, or CFL1, and corresponding scrambled (Scr.) controls. GAPDH, β-Actin and α-Tubulin were used as loading controls. Blots are representative of at least 3 independent experiments. Uncropped Western blots are provided in Supplementary Fig. 7. (i) Quantification of the Western blot shown in panel h, showing the mitotic levels of MLC2 and pMLC2-Ser19 in each knock-down relative to its Scr. siRNA control. For all panels, Welch’s t-test P-values: ns P > 0.05, ∗P < 0.05, ∗∗P < 0.01, ∗∗∗P < 0.001.
Supplementary Figure 4 Depletion efficiency for the targeted siRNA screen for cortex thickness regulators.
The bar plot shows the mRNA expression levels (RQ) in adherent HeLa cells treated with siRNA against various ABPs normalized to mRNA levels in control non-silencing pools (RQ (Norm.)). RQ levels were calculated by normalizing mRNA levels of queried genes to the averaged levels of GAPDH and ACTB. Relative mRNA abundance (RQ (Norm.)) was then calculated by dividing by the RQ value for control pools. Differences were considered significant if RQ (Norm.) was reduced by at least 40%. qPCR analysis was performed once for all siRNA conditions. For hits from the targeted ABP screen, Western blots were performed (Fig. 3b).
Supplementary Figure 5 Computer simulations of tension generation in a crosslinked actomyosin cortex.
(a) Actin filament representation (left): r is the centre of mass, s describes a position along the filament and the unit vector n is oriented towards the plus end of the filament. Myosin minifilament representation (right) with the centre of mass (r), orientation vector (n) and the two connecting heads. (b) Initialized network with a seeding thickness h0 and width W. Grayed filaments cross periodic boundaries in the x and y directions. (c) Myosin head attachment (top) and force balance on the myosin-actin attachment point (bottom). vs is the velocity of the myosin, αs is a friction coefficient, f0 is the myosin stall force and fms is the spring force of the myosin head. (d) Schematic of actin filament i with three springs connected, force balance on the filament, and sliced filament for surface tension measurement. The resulting force f ⊥ i along the normal of the slicing plane is used for calculating the total force. (e) Tension as a function of mean filament length for simulations where filament lengths were chosen according to a Gaussian distribution around a mean length. The standard deviation of the Gaussian distribution were 10% (blue), 20% (red), and 30% (green) of the mean length. (f) Tension as a function of the seeding thickness at constant filament length, density and number of motors. Seeding thickness is varied by changing the number of filaments. (g) Tension as a function of filament length at constant seeding thickness and density. The total number of filaments is adjusted to keep the total amount (length) of actin, La = Nfla, constant. For (e–g), each point represents the mean of 9 simulations, ±s.d. Tensions are normalized to T0 = 230 pN/μm.
Supplementary Figure 6 Effect of mitotic thickness regulators on the interphase cortex.
Box plots comparing (a) cortex thickness (P = 0.0016, 0.79, 0.16), (b) cortex tension (P = 0.018, 0.0025, 0.48), (c) relative cortex density (P = 0.0002, 6.3 × 10−5, 0.031), and (d) relative actin amount (P = 1.5 × 10−7, 0.023, 0.014) between interphase HeLa cells treated with siRNA targeted against CAPZB, CFL1 or DIAPH1 (Target, blue) or the corresponding scrambled siRNAs (Scr., black). Relative values were normalized to the median of the corresponding Scr. control. Points represent individual measurements (n = 33, 33, 31, 36, 26, 20 cells from 3 independent experiments for thickness, density and amount measurements; n = 22, 37, 20, 24, 23, 36 cells from 3-4 independent experiments for cortex tension measurements). Cortex thickness increased upon depletion of CAPZB, but remained unchanged upon CFL1 and DIAPH1 knock-down. Cortex tension increased upon depletion of CAPZB and CFL1 (as previously reported for CFL1, ref. 13), but remained unchanged upon DIAPH1 depletion. These results could suggest a mitosis-specific role of DIAPH1 in thickness/tension regulation. (e) Western blots showing the levels of CAPZB, CFL1, and DIAPH1 in whole cell lysates of interphase (Thymidine-blocked) and mitotic (STLC-blocked) HeLa cells. GAPDH was used as a loading control. Blots are representative from 7 independent experiments. (f) Box plots comparing the interphase/mitosis ratio of normalized CAPZB, CFL1, and DIAPH1 expression from 7 independent experiments. The whole cell levels of both CAPZB and CFL1 were higher in interphase than in mitosis, while DIAPH1 levels did not change. This suggests that the hits identified in our screen do not directly control the cortical thinning between interphase and mitosis. However, it is important to note that Western blot analysis reflects whole cell levels and may not reflect changes in cortical localization. Furthermore, these experiments do not take into account any possible post-translational modification of these proteins. Dotted line (ratio = 1) signifies no change in levels between interphase and mitosis. Uncropped western blots are provided in Supplementary Fig. 7. Welch’s t-test P-values: ns P > 0.05, ∗P < 0.05, ∗∗P < 0.01, ∗∗∗P < 0.001.
Supplementary Figure 7 Uncropped Western blots for Fig. 3b, Supplementary Figs 3h and 6e.
Red dotted lines indicate regions that were cropped for the figures.
Supplementary information
Supplementary Information
Supplementary Information (PDF 10685 kb)
Supplementary Table 1
Supplementary Information (XLSX 33 kb)
Supplementary Table 2
Supplementary Information (XLSX 45 kb)
Supplementary Table 3
Supplementary Information (XLSX 44 kb)
Supplementary Table 4
Supplementary Information (XLSX 41 kb)
Supplementary Table 5
Supplementary Information (XLSX 22 kb)
Simulation of a 3D cortex with 200 nm long actin filaments.
Left: xy and xz views of the simulated cortex with la = 200 nm. Right: Evolution of tension over time during the simulation run. (MOV 9991 kb)
Simulation of a 3D cortex with 500 nm long actin filaments.
Left: xy and xz views of the simulated cortex with la = 500 nm. Right: Evolution of tension over time during the simulation run. (MOV 6092 kb)
Simulation of a 3D cortex with 740 nm long actin filaments.
Left: xy and xz views of the simulated cortex with la = 740 nm. Right: Evolution of tension over time during the simulation run. (MOV 4935 kb)
Rights and permissions
About this article
Cite this article
Chugh, P., Clark, A., Smith, M. et al. Actin cortex architecture regulates cell surface tension. Nat Cell Biol 19, 689–697 (2017). https://doi.org/10.1038/ncb3525
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3525
This article is cited by
-
SimuCell3D: three-dimensional simulation of tissue mechanics with cell polarization
Nature Computational Science (2024)
-
Intracellular tension sensor reveals mechanical anisotropy of the actin cytoskeleton
Nature Communications (2023)
-
TRIM40 is a pathogenic driver of inflammatory bowel disease subverting intestinal barrier integrity
Nature Communications (2023)
-
Membrane tension induces F-actin reorganization and flow in a biomimetic model cortex
Communications Biology (2023)
-
The effects of internal forces and membrane heterogeneity on three-dimensional cell shapes
Journal of Mathematical Biology (2023)