Abstract
Recapitulation of lung development from human pluripotent stem cells (hPSCs) in three dimensions (3D) would allow deeper insight into human development, as well as the development of innovative strategies for disease modelling, drug discovery and regenerative medicine1. We report here the generation from hPSCs of lung bud organoids (LBOs) that contain mesoderm and pulmonary endoderm and develop into branching airway and early alveolar structures after xenotransplantation and in Matrigel 3D culture. Expression analysis and structural features indicated that the branching structures reached the second trimester of human gestation. Infection in vitro with respiratory syncytial virus, which causes small airway obstruction and bronchiolitis in infants2, led to swelling, detachment and shedding of infected cells into the organoid lumens, similar to what has been observed in human lungs3. Introduction of mutation in HPS1, which causes an early-onset form of intractable pulmonary fibrosis4,5, led to accumulation of extracellular matrix and mesenchymal cells, suggesting the potential use of this model to recapitulate fibrotic lung disease in vitro. LBOs therefore recapitulate lung development and may provide a useful tool to model lung disease.
Similar content being viewed by others
Main
The respiratory system originates from buds that arise on the ventral aspect of the anterior foregut endoderm (AFE) and develop through a stereotyped branching process into proximal airways and distal alveolar progenitors (pseudoglandular stage). During the canalicular stage, cell cycle activity decreases, and specialization of the airway epithelium occurs in the stalks, with the emergence of basal, goblet, club, ciliated, and other cell types. This stage is followed by the saccular stage, where the canaliculi widen into distal sacculations that will give rise to primitive alveoli6,7. Organoids are in vitro generated 3D structures containing multiple cell types that are organized similar to an organ and recapitulate some specific organ function1. One group reported generation of human lung organoids8,9. However, while these small structures contained cells expressing markers of lung and airway8, and have some airway potential after subcutaneous xenografting in mice9, they do not satisfy the aforementioned criteria for organoids, as neither features of lung development, notably branching morphogenesis and proximodistal specification, nor function were observed.
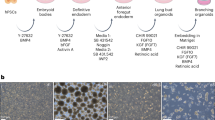
We previously reported a strategy to differentiate hPSCs (embryonic stem cells (ESCs) and induced pluripotent stem cells (iPSCs)) in 2D through sequential developmental steps from definitive endoderm (DE) to AFE, lung field progenitors, and, finally, lung and airway epithelial cells (Supplementary Fig. 1a)10,11,12. Early during induction of a ventral lung fate from AFE, adherent structures formed that detached easily and expanded in suspension culture as clumps of cells (Fig. 1a, b) in the presence of BMP4, FGF10, FGF7, retinoic acid (RA) and the GSK3β antagonist, CHIR99201 (Supplementary Fig. 1b), factors shown previously to be required for lung development6,7. 7.5 × 105 DE cells yielded 2,490 ± 129 clumps (n = 3; RUES2 ESCs). The structures formed folding sheets of EPCAM+KRT8+ECAD+FOXA1/2+ AFE cells (FOXA2: 89.07 ± 3.36%, EPCAM+: 92.08 ± 1.88%, n = 3; RUES2 ESCs) (Fig. 1c). By day 25 51.26 ± 4.37% (n = 3; RUES2 ESCs) of the cells expressed the lung marker NKX2.1+ (Fig. 1c). Except for the epithelial progenitor marker, p63 (18.59 ± 1.49%, n = 3; RUES2 ESCs, Fig. 1c), markers of mature lung and airway cells were absent (not shown). The cells were surrounded by mesodermal PDGFRA+ECAD− cells (Fig. 1d). RNAseq (Supplementary Fig. 1c) confirmed strong enrichment of endoderm/lung genes (FOXA2, SOX2, NKX2.1) in EPCAM+ cells (Fig. 1e). EPCAM− cells expressed mesodermal genes (Fig. 1e), some of which, such as TBX4 and HOX5 paralogs, are expressed in pulmonary mesoderm13,14. Genes expressed in mature lung and airway and in other AFE-derived lineages were nearly undetectable in the EPCAM+ fraction (Supplementary Fig. 1d). Sonic Hedgehog (SHH) was expressed in endodermal cells, and its transcriptional targets15, PTCH1, GLI1 and HHIP in mesoderm (Supplementary Fig. 1d). In situ hybridization confirmed SHH expression in the endodermal fraction at day 15. At day 25, SHH was expressed most strongly in the tips of budding epithelial structures (Supplementary Fig. 1e). These findings are consistent with the developing mouse lung where SHH is expressed throughout the pulmonary endoderm early but is limited to branch tips during branching morphogenesis15,16,17. Because they contain multiple cell types that are spatially organized similar to developing lung buds in vivo, we call these structures lung bud organoids (LBOs).
(a) Development of adherent structures during ventralization of AFE between day 6 and day 8 (see protocol Supplementary Fig. 1b), which could be expanded in suspension culture (day 10, day 20). Representative of >50 independent experiments (ESCs and iPSCs). Scale bars, 250 μm. (b) Cellular expansion during the generation of LBOs (mean ± s.e.m., n = 3 independent experiments in RUES2 ESCs). The source data can be found in Supplementary Table 4. (c) Expression of EPCAM, KRT8, NKX2.1, FOXA1, and P63 in day 25 LBOs. Representative of >10 independent experiments in ESCs and iPSCs. Scale bars, 100 μm. (d) Staining of day 25 LBO for ECAD and PDGFRA. Representative of 3 independent experiments in RUES2 ESCs. Scale bar, 250 μm. (e) Expression of endodermal and mesodermal markers in the EPCAM+ and EPCAM− fraction of day 25 LBOs determined by RNAseq (3 independent biological replicates, RUES2 ESCs).
When transplanted under the kidney capsule of immunodeficient NSG mice, LBOs yielded growths (Fig. 2a) containing tubular structures surrounded by mesenchymal tissue after 1.5 months (Fig. 2b). The tubes were uniformly lined by a FOXA2+NKX2.1+SOX2+ epithelium containing MUC5AC+ (goblet) cells with p63+ cells in the basal layer (Fig. 2c), compatible with airway epithelium. All cells were human (Supplementary Fig. 2a), except for endothelial cells, which were of mouse origin (Supplementary Fig. 2b). After 5 months, branching structures (Fig. 2d and Supplementary Fig. 2c) surrounded by SMA+ mesodermal cells arose (Supplementary Fig. 2c). All epithelial cells were NKX2.1+, while SOX2, a proximal marker later in lung development18,19, was excluded from the branch tips, which expressed SFTPB and SFTPC, markers of surfactant-producing type II alveolar epithelial (ATII) cells (Fig. 2e)20. The stalks and central tubules expressed the proximal (airway) markers FOXJ1 (ciliated cells), CC10 (club cells) and mucins (goblet cells) (Fig. 2e). Haematoxylin–eosin staining showed abundant multiciliated cells (Supplementary Fig. 2d), while live imaging documented beating cilia (Supplementary Video 1). Furthermore, submucosal glands were observed near the larger tubular structures (Supplementary Fig. 2e). Overall, morphology and expression pattern within the growths are consistent with proximodistal specification during lung branching morphogenesis6,7. The fluid in the tubular structures contained all tested secretory products of lung and airway, but was negative for the cell surface mucin21, MUC1, indicating detection of secreted proteins and not proteins associated with sloughed cells, and providing evidence for function (Fig. 2f). After 7 months, dome-shaped groups of CGRP+PGP9.5+ cells, compatible with neuroepithelial bodies22, were present in the airway-like structures (Fig. 2g). Furthermore, areas of the growths developed into a network of thin cell layers (Fig. 2g) containing cells expressing ATII cells markers (SFTPC, ABCA3, HT2-280)23 and cells bearing type I alveolar epithelial cell (ATI) markers (HT1-56, HOPX, PDPN, CAV1, SCNN1A, AKAP5, CLIC5)20, although other markers for mature ATI cells (RAGE, AQP5)20 were not detected (Fig. 2g). However, an alveolar capillary network and bronchoalveolar ducts were not observed. We conclude that, although full phenotypic and architectural alveolar maturation was not achieved, possibly at least in part because of the ectopic location, LBOs recapitulate many essential features of lung development, including branching morphogenesis and proximodistal specification, after xenotransplantation.
(a) Macroscopic aspect of growths 1.5 months after transplantation of 106 LBO cells embedded in Matrigel under the kidney capsule of NSG mice. Scale bar, 1 cm. (b) Haematoxylin–eosin (HE) stain of LBO-derived growth 1.5 months after transplantation. Scale bar, 500 μm. (c) Immunofluorescence for indicated markers in LBO-derived growths 1.5 months after transplantation. Scale bars, 100 μm. (d) HE staining of LBO-derived growths 5 months after transplantation. Scale bars, 250 μm. (e) Immunofluorescence for indicated markers in LBO-derived growths 5 months after transplantation. Scale bars, 250 μm. (f) Dot blots for proteins marked on the left in aspirates from tubules in LBO-derived growths 5 months after transplantation. (g) HE staining and immunofluorescence for indicated markers in LBO-derived growths 7 months after transplantation. Scale bars, 100 μm. All panels used RUES2 ESCs, representative of 4 independent experiments.
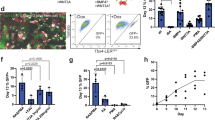
After embedding day 25 LBOs in Matrigel in the presence of CHIR99021, FGF10, FGF7, BMP4 and RA (Supplementary Fig. 1b), more than 95% yielded rapidly expanding branching structures (Fig. 3a and Supplementary Fig. 3a for iPSCs, including C12, a line from a patient with mutations IRF7, causing acute respiratory distress syndrome after influenza infection)24 expressing markers of pulmonary endoderm (FOXA2+: 95.17 ± 1.54%, NKX2.1+: 74.97 ± 4.37%, EPCAM+: 96.83 ± 0.62%, SOX9+: 92.42 ± 3.81% n = 3 at day 70; RUES2 ESCs) (Fig. 3b). Uniform luminal expression of MUC1 demonstrates polarization (Fig. 3b). Cells expressing the ATII markers SFTPC, SFTPB and ABCA3 were present in all structures (Fig. 3b). Airway goblet cells (MUC5B or MUC5AC) were rare while other airway cells (club cells (SCGB3A2), ciliated cells (FOXJ1) and basal cells (KRT5 and P63)) were not detected (not shown). While singly plated LBOs branched randomly in every direction and filled a 6.4 mm well within 90 days, they formed branching trees that occupied a section of the well only when plated together in close proximity (Supplementary Fig. 3b). These findings show that branching architecture can be manipulated in vitro, and that repulsive interactions between branching structures may play a role in determining their architecture. Mesenchymal cells expressing VIMENTIN and CD90 were present surrounding the structures (Fig. 3b). However, their proportion, as determined by flow cytometry for EPCAM− cells, declined during Matrigel culture to less than 2% of the total population (Supplementary Fig. 3c). EPCAM+, but not EPCAM− cells, purified from day 25 LBOs yielded branching colonies after plating in Matrigel, albeit with low cloning efficiency (0.30 ± 0.0316%,) (Supplementary Fig. 3d). These branching colonies displayed a similar pattern of marker expression as Matrigel colonies generated from intact LBOs (Supplementary Fig. 3e). These findings indicate that rare progenitors in the LBOs are capable of generating branching colonies, and that mesenchymal cells are not required for branching in these culture conditions.
(a) Bright-field images of the development of an LBO into a branching structure after plating in Matrigel. RUES2 ESCs. Arrows indicate branching distal tip. Representative of >50 independent experiments. Scale bars, 500 μm. (b) Immunofluorescence staining for indicated markers in day 70 RUES2-derived LBOs plated in Matrigel at day 25. Representative of 4 independent experiments. Scale bars, 250 μm.
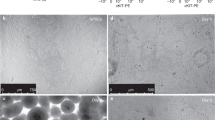
After >170 days, macroscopic tissue (Fig. 4a) consisting of branching tubules with dilated tips, reminiscent of saccules formed during the saccular stage of lung development, had developed (Fig. 4b and Supplementary Fig. 4a and Supplementary Video 2). 84.86 ± 5.21% cells were NKX2.1+, while most cells were SOX9+(76.75 ± 6.89%) and a minority (23.78 ± 5.21%) were SOX2+ (Supplementary Fig. 4b) (n = 4, one ESC and three iPS lines). Most luminal cells expressed HT2-280, MUC1, SFTPB, SFTPC and ABCA3 (RUES2 Fig. 4c, iPSCsSupplementary Fig. 4b), identifying these as ATII cells. Electron microscopy showed large numbers of lamellar bodies (LBs), the organelles where surfactant is stored25 (Fig. 4d and Supplementary Fig. 4c). To examine ATII cell function, we added SFTPB covalently linked to the fluorescent lipid, BODIPY. Within minutes, SFTPB-BODIPY was taken up by the cells and secreted in the lumens (Fig. 4e, f and Supplementary Video 3). Although HOPX, a marker of ATI cells and of putative bipotential alveolar progenitors in the mouse20,26, was widely expressed (Supplementary Fig. 4b), other ATI markers (AQP5, CLIC5, AKAP5, CAV1, AGER) were undetectable. SOX9, a marker for the distal tips that is downregulated as alveoli mature and becomes undetectable postnatally, was mostly expressed at the tips and outer edges of the branching structures in vitro, consistent with mouse development, where SOX9 is a distal lung marker27,28,29 (Supplementary Fig. 4b). Expression of airway markers (MUC5AC, SCGB3A2) in the Matrigel LBO colonies was confined to structures co-expressing SOX2 and SOX9 (Supplementary Fig. 4b), and were therefore probably more proximal. While co-expression of SOX2 and SOX9 is unusual in the mouse19, numerous larger SOX2+SOX9+ structures were identified in the human second trimester fetal distal lung (Supplementary Fig. 4d), suggesting that LBOs recapitulate human lung development. The expression of SOX9, the formation of saccular structures expressing predominantly ATII markers and absence of mature ATI cells are consistent with the canalicular stage of lung development, which occurs at the end of gestation of mice, but during the late second trimester in humans. To further verify the developmental stage of day 170 Matrigel LBO cultures, we performed RNAseq on 12 independent samples from RUES2 cells and from three iPSC lines. Cross-referencing with the KeyGenes database, which contains expression profiles of human organs during first and second trimesters of gestation and adulthood30, showed the best match with second trimester fetal lung, without any match with other organs (Fig. 4g and Supplementary Fig. 4e). Together, the structural, protein expression and transcriptomic data indicate that the day 170 Matrigel LBO organoids reached the late second trimester of human gestation.
(a) Macroscopic appearance of day 170 RUES2 LBOs embedded in Matrigel at day 25. Representative of >50 independent experiments. Scale bar, 5 mm. (b) Bright-field images of day 170 RUES2 and C12 LBOs embedded in Matrigel at day 25. Representative of >50 independent experiments. Scale bars, 500 μm. (c) Immunofluorescence for indicated markers in day 170 RUES2 LBOs embedded in Matrigel at day 25. Representative of 3 independent experiments. Scale bars for MUC1 + SFTPB and HT2-280, 100 μm. Scale bar for SFTPC, 10 μm. (d) Electron microscopy of day 170 LBOs embedded in Matrigel at day 25 in RUES2 ESCs and HDF SV iPSCs. Arrows indicate LBs. Representative of 3 independent experiments. (e) Uptake of SFTPB-BODIPY (green) in day 170 LBOs embedded in Matrigel at day 25. Representative of 4 independent experiments. Scale bars, 100 μm. (f) Time-course of uptake of SFTPB-BODIPY in day 170 LBOs embedded in Matrigel at day 25 (mean ± s.e.m., n = 4 independent experiments in RUES2 ESCs). The source data can be found in Supplementary Table 4. (g) Comparison of genome-wide expression in day 170 LBOs derived from hESCs and hiPSCs (12 biologically independent samples) with the KeyGenes database, showing the best match with second trimester human lung.
We next explored whether select infectious and fibrotic lung disease could be recapitulated. We asked whether LBOs infected with respiratory syncytial virus (RSV) display features of human lung infection. RSV is a major cause of lower respiratory tract infection in infants, and causes bronchiolitis with obstruction of small airways2,31. There is no licensed vaccine or effective antiviral drug at this time, and immunity after infection is short-lived32. RSV tropism in humans includes ciliated cells and alveolar epithelial cells2,3. Previous studies in human airway epithelial cell lines showed that cells infected with RSV swell and detach from the epithelium33, a finding consistent with obstruction of small airways by infected cells in archival pathology specimens and with the clinical syndrome of bronchiolitis3. At day 2 after infection of day 170 Matrigel LBO cultures with RSV, confocal microscopy revealed shedding of swollen, infected cells into the lumen of the branching structures (Fig. 5a, arrows, Supplementary Video 4). No shedding was seen at day 1, despite evidence of viral infection. RSV infection in LBOs therefore recapitulates important features of infection in humans.
(a) Confocal images of whole mount day 170 LBOs 1 and 2 days after infection with RSV and stained using anti-RSV (all antigens) antibody. Arrows: infected cells in the lumen. Representative of 3 independent experiments. Scale bars, 100 μm. (b) Bright-field images of day 50 LBO-derived Matrigel colonies from RUES2 and RUES2-HPS1 cells. Representative of six independent experiments. Scale bars, 500 μm. (c) Fraction of EPCAM+ and EPCAM− cells in day 50 LBO-derived colonies in 3D Matrigel cultures of RUES2 and RUES2-HPS1 cells. (n = 6, mean ± s.e.m. of 3 technical replicates from two experiments; ∗P < 0.0001; two-tailed Student’s t-test). The source data can be found in Supplementary Table 4. (d) Immunofluorescence staining for mesenchymal markers and ECM components in 3D Matrigel cultures of RUES2 and RUES2-HPS1 cells. Representative of 3 independent experiments. Scale bars, 500 μm.
Next, we attempted to model pulmonary fibrosis associated with some forms of Hermansky–Pudlak Syndrome (HPS)5. HPS is characterized by pigmentation and bleeding abnormalities caused by abnormal biogenesis and trafficking of lysosome-related organelles (LROs), which include platelet dense granules and melanosomes34. Some forms, in particular HPS1, are associated with early-onset and intractable pulmonary fibrosis (HPS interstitial pneumonia (HPSIP)) that is clinically similar idiopathic pulmonary fibrosis (IPF)5, is characterized by fibrotic obliteration of alveoli and has a median survival of 3–4 years35. The fact that LBs of ATII cells are also LROs34 potentially explains the association of IPF with some mutations causing HPS5. Matrigel colonies derived from LBOs generation from RUES2 cells with CRISPR-CAS9-induced deletion of HPS1 (Supplementary Fig. 5a) exhibited less sharply defined branching structures in Matrigel cultures than the LBOs from parental RUES2 line (Fig. 5b), with an increased fraction of EPCAM− mesenchymal cells (Fig. 5c and Supplementary Fig. 5b), heterogeneously expressing the mesenchymal markers PDGFRA, PDGFRB, SMA, VIMENTIN and CD90 (Fig. 5d, low-magnification tile scans in Supplementary Fig. 5c). The EPCAM−, but not the EPCAM+ population, showed strongly enhanced proliferation in cultures of RUES2-HPS1 cells compared to parental cells (Supplementary Fig. 5d, e), indicating that expansion of mesenchymal cells explains the increased fraction of EPCAM− cells. Surprisingly, however, hyperproliferation of EPCAM− cells was already noticed in RUES2-HPS1 LBOs as early as day 15 of suspension culture, prior to detection of any ATII markers. Furthermore, increased hydroxyproline content (Supplementary Fig. 5f) as well as enhanced extracellular matrix (ECM) autofluorescence (Supplementary Fig. 5c) and immunofluorescent staining for collagens 1 and 3 and fibronectin (Fig. 5d and Supplementary Fig. 5g) in RUES2-HPS1 cells indicated increased ECM deposition. Mixing experiments (Supplementary Fig. 5h–j) were consistent with the notion that the accumulation of mesenchymal cells was driven by mutant epithelial cells, and not a cell intrinsic property of mutant mesenchymal cells, a finding consistent with the notion that HPSIP36 and potentially other forms of IPF4 may be caused by epithelial injury. Together, these findings suggest that it may be possible to model at least some fibrotic pulmonary disease using LBOs.
LBOs and LBO-derived branching colonies in Matrigel in vitro and growths after xenografting fulfil the definition of true organoids1. Previously reported human lung organoids did not show branching in vitro or after xenografting8,9. Furthermore, in contrast to LBOs, these were generated in the presence of serum, but in the absence of BMP4, RA and Wnt agonism, which we have shown to be essential for lung specification in vitro10. Finally, these structures did not develop in vivo after grafting under the kidney capsule of immunodeficient mice, but required preculture on a bioengineered scaffold to generate airway epithelial cells after subcutaneous transplantation9.
We could reproduce the morphological features of RSV infection in the distal lung, for which there is currently no model that reproduces human infection. The LBO model also showed evidence of fibrosis in cells lacking HPS1, mutation of which is the most penetrant for a form of pulmonary fibrosis that is clinically, prognostically, radiologically and pathologically indistinguishable from IPF4,5,36. It is remarkable, however, that while HPSIP typically arises in the third to fourth decade of life, a fibrotic phenotype could be reproduced in vitro within 40 days of directed differentiation. While it cannot be fully excluded that this in vitro model in fact reveals a developmental abnormality not observed in patients, it is possible that stress of in vitro culture recapitulated the changes induced by senescence and led to the very rapid appearance of the phenotype, in particular since age and telomere dysfunction are prime risk factors for IPF35,37.
However, the LBO model has limitations. After 6 months of culture in Matrigel, the organoids match the second trimester of human gestation in terms of structure, marker expression and genome-wide expression signature. These findings suggest that lung development as modelled in the LBO system keeps pace with human lung development in utero. Full, terminal maturation therefore remains a challenge in the organoid field1. A second limitation is that branching appears random, a finding consistent with a, as yet unproven, ‘space-filling’ model of branching morphogenesis38. However, branching could be directed by plating several LBOs in close proximity to each other in Matrigel, in which case the organoids branch away from each other, suggesting that branching can be manipulated in vitro. A third limitation is that the exact nature and patterning of the mesenchyme present in the LBOs is unclear. In vivo xenografting revealed that LBO-associated mesodermal cells do not have the potential to generate endothelial cells, bone or skeletal muscle, suggesting that the mesenchyme is specified to some extent. The various mesenchymal lineages in the lung and their ontogeny are still poorly characterized6. However, pulmonary vasculature is probably not derived from pulmonary mesenchyme. Proximal pulmonary vessels are derived from a common cardiopulmonary mesenchymal progenitor, while the alveolar capillary network probably arises from VE-cadherin+ progenitors in pre-existing trunk vessels6,39. A fourth limitation is that the in vitro cultures are strongly biased towards distal lung, and, although some areas co-expressing SOX2 and SOX9 expressed more proximal markers for goblet cells and club cell precursors, mature club cells, ciliated cells or basal cells were not observed. We could also not achieve induction of ATI markers in vitro, although ATI potential is present after engraftment in vivo. It is possible that the cultures were not sufficiently mature to allow development of ATI cells. Alternatively, our current culture condition may drive differentiation of distal progenitor towards ATII cells at the expense of ATI cells.
Taken together, this work indicates that, despite certain limitations, LBOs will be a useful tool for the study of human lung development, and possibly for lung disease modelling. □
Methods
Reagents.
Reagents used are listed in Supplementary Table 1.
Human samples.
The use of human fetal tissues procured by the Human Studies Core at Columbia Center for Translational Immunology was approved by the Columbia University Medical Center (CUMC) Human Research Review Committee and the experiments were performed in accordance with the approved protocols.
Media.
hPSC maintenance media consisted of DMEM/F12 (1:1) supplemented with 20% knockout serum replacement, 0.1 mM β-mercaptoethanol, 1 ml Primocin, 5 ml Non-essential amino acids, 5 ml GlutaMax, and 20 ng ml−1 FGF-2. Serum-free differentiation (SFD) media consisted of IMDM/Ham’s F12 (3:1) supplemented with N2, B27, 0.05% bovine serum albumin, 1% penicillin–streptomycin, 50 μg ml−1 ascorbic acid, 2 mM Glutamax, 0.4 μM monothioglycerol and different growth factor cocktails as indicated in Supplementary Table 2.
hPSCs maintenance.
Rockefeller University Embryonic Stem Cell Line 2 (RUES2; A. Brivanlou, Rockefeller University; NIH approval number NIHhESC-09-0013, Registration number 0013, passage 17–28), Sendai Virus and modified mRNA generated hiPSC lines from healthy human dermal fibroblasts7,9 (S. d’Souza, Mount Sinai Stem Cell core; passage 16–25) and IRF7-deficient C12 hiPSC lines28 (L. Notarangelo and S. Volpi) were maintained on mouse embryonic fibroblasts (MEFs) plated at 15,000–18,000 cells cm−2. Cells were cultured in hPSC maintenance media and medium was changed daily. hPSCs were passaged with Accutase/EDTA washed and replated at a dilution of 1:48. Cultures were maintained in a humidified 5% CO2 atmosphere at 37 °C. Lines are karyotyped and verified for mycoplasma contamination using PCR every 6 months.
Endoderm induction.
Induction of endoderm was carried as previous described9. Briefly, MEFs were depleted by passaging onto Matrigel for 24 h supplied with hPSC maintenance media and maintained in a humidified 5% CO2 atmosphere at 37 °C. After MEF depletion, primitive streak and embryoid body induction was performed in embryoid bodies/primitive streak formation media (Supplementary Table 2) in low attachment plates for 12–16 h, followed by switching to endoderm induction media (Supplementary Table 2) for 36–40 h. Embryoid bodies were fed every day and maintained in a humidified 5% CO2/5% O2 atmosphere at 37 °C. Endoderm yield was determined by the expression of CXCR4 and c-KIT. For iPS lines, endodermal cells were purified using human CD184 (CXCR4) MicroBead kit. Cells used in all experiments had >90% endoderm yield.
Anterior foregut endoderm induction.
Anterior foregut endoderm was induced as previous described9. On day 4, embryoid bodies were dissociated with 0.05% Trypsin/EDTA and plated on fibronectin-coated multiple-well plates with a density at 80,000–105,000 cells cm−2. Cells were incubated in Anteriorization media-1 for 24 h, followed by switching to Anteriorization media-2 for another 24 h.
Formation of lung bud organoids.
At the end of anterior foregut endoderm induction, cells were treated with ventralization media (branching media) for 48 h and three-dimensional clump formation was observed. The clumps were then suspended by gently pipetting around the wells. The suspended clumps are called lung bud organoids (LBOs) hereafter. LBOs were maintained in non-tissue culture treated multiple-well plates submerged in branching media and were fed every other day until day 20–day 25.
Branching morphogenesis in Matrigel.
The day 20–day 25 LBOs were embedded in 100% Matrigel in 24-well Transwell inserts and incubated in an incubator until the Matrigel solidified. Branching media were added to the well, after which the Transwell was inserted, branching media added into the Transwell insert as well. Media were changed every other day. A step-by-step protocol describing the generation of LBOs and LBO-derived branching colonies in Matrigel can be found at Nature Protocol Exchange40.
Immunofluorescence staining.
LBOs and branching Matrigel cultures were freshly embedded in Optimal Cutting Temperature (OCT). Samples were sectioned between 5–8 μm, and then air-dried for 2 h. The sections were fixed with 4% paraformaldehyde for 20 min at room temperature (RT) and washed with DPBS for 5 min. The sections were permeabilized with 0.3% Triton X-100/PBS for 30 min followed by blocking in 5% donkey serum for 1 h. Primary antibodies (Supplementary Table 3) were incubated at 4 °C overnight. The next day, sections were washed with DPBS 3 × 5 min followed by secondary antibody (Supplementary Table 3) incubation for 2 h at RT, washed 3 × 10 min with DPBS, then mounted with DAPI contained fluorescent mounting medium. For 3D imaging, D25 LBOs were stained as described above, but were stained as intact organoids.
Isolation of EPCAM+ and EPCAM− population from LBOs.
LBOs were dissociated by 0.05% Trypsin/EDTA. The cells were stained with APC-conjugated EPCAM for 20 min at 4 °C. EPCAM+ and EPCAM− cells were isolated by fluorescence activated cell sorting (FACS) using a BD Influx Cell Sorter.
RNAseq.
Total RNA from LBOs was purified using Direct-zol RNA MicroPrep kit. RNA concentration and RNA integrity number (RIN) were determined using an Agilent microfluidic RNA 6000 Nano Chip kit (Agilent Technologies) on a 2100 Bioanalyzer (Agilent Technologies). Those samples with RIN greater than 9 were used for RNAseq. Poly-A-pull-down was used to enrich mRNAs from total RNA samples. Libraries were prepared using Illumina TruSeq RNA prep kit (Illumina). Libraries were then sequenced using the Illumina HiSeq2000 (Illumina) at the Columbia Genome Center. Samples were multiplexed in each lane, yielding a targeted number of single-end/pair-end 100 bp reads for each sample, as a fraction of 180 million reads for the whole lane. RTA (Illumina) was used for base calling and bc12fastq (version 1.8.4) for converting BCL to fastq format, coupled with adaptor trimming. Reads were mapped to a reference genome (NCBI/build37.2) using Tophat (version 2.0.4) with four mismatches and ten maximum multiple hits. To tackle the mapping of reads that are from exon–exon junctions, Tophat infers novel exon–exon junctions ab initio, and combines them with junctions from known mRNA sequences as the reference annotation. We estimated the relative abundance of genes and splice isoforms using Cufflinks (version 2.0.2) with default settings. We tested for differentially expressed genes under various conditions using DEseq, an R package based on a negative binomial distribution that models the number reads from RNAseq experiments and tests for differential expression.
In situ hybridization.
In situ hybridization was performed on frozen sections (5–8 μm) using digoxigenin (DIG)-UTP-labelled SHH riboprobes. Briefly, human adult lung tissue cDNA was used as template to generate SHH PCR products containing either T7 or T3 promoter sequences (Forward: AATTAACCCTCACTAAAGGGACAGCTCGGAAGTCATCAGTT; Reverse: TAATACGACTCACTATAGGGG CCTCTGAGTGGTGGCCATCTT). The PCR products were used as templates to generate SHH riboprobes using T7 MAXIscript kit (Ambion) followed by RNeasy micro kit (Qiagen) to clean up the riboprobes. Different stages of the LBOs freshly embedded in OCT. Samples were sectioned between 5–8 μm, followed by fixation with 4% paraformaldehyde for 20 min RT. The sections were washed with DEPC-DPBS for 3 × 5 min and acetylated in acetylation buffer (584 μl of triethanolamine/50 ml of DEPC-H2O/125 μl acetic anhydride) for 10 min. Permeabilization was carried in 0.1% Triton X-100/PBS for 30 min at RT followed by washed with DEPC-DPBS 3 × 5 min. The sections were incubated with hybridization buffer (5% dextran sulfate/4× SSC/50% formamide/1× Denhardt’s/5% fish sperm DNA) for at least 2 h at RT then overnight with 200 ng ml−1 of DIG-labelled SHH probe in hybridization buffer at 72 °C. The next day, sections were incubated with 0.2× SSC pre-warmed to 72 °C for 2 h, followed by cool down to RT for 30 min. The sections were washed with fresh 0.2× SSC for 5 min then PBS for another 5 min. The sections were incubated with blocking solution (2% sheep serum/TBST) for 1 h followed by anti-DIG-AP Ig overnight at 4 °C. The sections were washed with TBST 3 × 10 min and rinsed in colour reaction buffer (100 mM Tris, pH 9.5/0.1% Tween-20/100 mM NaCl/50 mM MgCl2) for 10 min. Colour was developed by incubating the section with BM-purple.
Mouse kidney capsule transplantation.
The NOD.Cg-Prkdcscid.Il2rgtm1Wjl/SzJ (NSG) mice were housed in a specific pathogen-free mouse facility. All the mice used at 10–13 weeks of age and not selected for gender. The experiment was set up to use 5–7 mice per time point. No statistical method was used to predetermine sample size. The experiments were not randomized. Experiments and animal care were performed in accordance with the protocols approved by The Columbia University Institutional Animal Care And Use Committee. One million day 20–day 25 LBO cells were mixed with 5 μl Matrigel prior to surgery and implanted under the kidney capsule. Outgrowths were excised, embedded freshly in OCT for immunofluorescence or fixed in 4% paraformaldehyde for paraffin embedding. Histology was analysed using haematoxylin/eosin staining.
Dot blots.
Three microlitres of fluid aspirated from the tubular structures of 5 month grafts was deposited onto a nitrocellulose blotting membrane (GE Healthcare Life Sciences). The dot-blot membrane was air-dried for 5 min, and blocked in 5% milk/PBS for 1 h and then probed with the indicated primary antibodies (Supplementary Table 3) overnight at 4 °C. HRP-conjugated secondary antibodies was applied to the membranes, followed by signal detection with ECL Western Blotting Detection Reagents and exposure to X-ray film.
Imaging.
Samples were imaged using motorized Leica DMI6000 B (Leica Microsystems) or DMi8 (Leica Microsystems) inverted microscopes or a two-photon confocal laser scanning microscope Leica TCS SP8 (Leica Microsystems). Macroscopic images (Figs 3a and 5a) were taken using an iPhone 6 (Model: MG5A2LL/A, Apple).
Quantification of immunofluorescence.
Images for each nuclear marker were quantified using ImageJ. Briefly, images were converted to 8-bit images and the threshold was adjusted to correspond with the nuclear stain, which allows for measurement of total area. The total area was analysed by the ‘Analyze Particles’ function of ImageJ. Percentage of positive cells were calculated by dividing the total area of positive cells over the total area of DAPI. For extracellular matrix quantification, fluorescence intensity was quantified using Leica Application Suite X. The values were normalized to the RUES2 control for each individual experiment before statistical analysis.
Transmission electron microscopy.
Transmission electron microscopy (TEM) was performed at the NYU Langone Medical Center Microscopy Core. LBOs were fixed with 2.5% glutaraldehyde in 0.1M sodium cacodylate buffer (pH 7.2) for 2 h and post-fixed with 1% osmium tetroxide for 1.5 h at room temperature, then processed in a standard manner and embedded in EMbed 812 (Electron Microscopy Sciences). Semi-thin sections were cut at 1 mm and stained with 1% toluidine blue to evaluate the quality of preservation and find the area of interest. Ultrathin sections (60 nm) were cut, mounted on copper grids and stained with uranyl acetate and lead citrate by standard methods. Stained grids were examined under a Philips CM-12 electron microscope and photographed with a Gatan (4 k × 2.7 k) digital camera (Gatan).
Generation of RUES2-HPS1 line.
The RUES2-HPS1 line was generated at the Stem Cell Core Facility at Columbia University Medical Center. Briefly, RUES2 cells (passage 25) were cultured in six-well plates coated with Matrigel to 70–80% confluence. Cells were electroporated with 7.5 μg of HPS1 guide RNA plasmid plus 2.5 μg of Cas9mCherry per well of a six-well plate using Nucleofector 4D. Cas9mCherry-derived mCherry was used as a fluorescent marker to sort transfected cells. Twenty-four hours posttransfection, cells were sorted using FACS with a Bio-Rad S3e cell sorter and seeded at ∼2,000 cells/6 cm dish on MEF feeders. Colonies were picked 7–10 days post sorting. Genomic DNAs from individual clones were isolated and genotyping was done using HPS1-specific PCR primers (HPS1-F-1 (5′-GTAGAGGCAGCAGATCCAAGAGG-3′) and HPS1-R-1 (5′-GAACAAGGTGGTCCACACA-3′). 420 bp band to be expected). The PCR products were cloned into a plasmid for proper sequence using In-Fusion reaction (Clontech). Sequencing revealed premature stop codons in each allele (Supplementary Fig. 5).
Uptake of SPB-BODIPY in live LBOs and quantification.
Day 170 LBOs were stained with CellMask Deep Red Plasma membrane Stain for 10 min and washed five times, followed by imaging prior loading SPB-BODIPY to obtained background fluorescence levels (0 min). The cultures then were loaded with 20 ng ml−1 purified human SPB-BODIPY protein (10 ng in total per culture) directly on top of the Matrigel. Images were taken every 2 min using a two-photon confocal laser scanning microscope (Leica TCS SP8) and the fluorescent intensities were quantified using Leica Application Suite X. The background fluorescence values were subtracted from all measurements before statistical analysis.
Hydroxyproline content.
Hydroxyproline content was measured following the manufacturer’s protocol (Sigma, MAK008-1KT). Briefly, samples from RUES2 or RUES2-HPS1 cultures were homogenized by tissue glass Teflon dounce homogenizer (10 mg samples in 100 μl of water) and transferred to a pressure-tight vial, followed by adding 100 μl of concentrated hydrochloric acid (∼12 M) per 10 mg of sample. The mixtures were hydrolysed at 120 °C for 3 h. Samples were dried in a 96-well plate at 60 °C followed by Chloramine T/Oxidation Buffer Mixture for 5 min at RT and DMAB reagent for another 90 min at 60 °C. Hydroxyproline content were measured at 560 nm. The same amount of Matrigel was used as control.
Comparative analysis using KeyGenes.
RNAseq data obtained from day 170 LBOs from RUES2, C12, HDF SV and HDF mRNA lines was compared to different first and second trimesters and adult organs, including the lungs, using KeyGenes. Hierarchical clustering of 12 samples of the day 170 LBOs and 75 samples from 19 organs from the second trimester was performed using Cluster 3.0 and viewed by TreeView. The 87 classifier genes were calculated by KeyGenes.
Respiratory syncytial virus preparation and infection.
Recombinant red fluorescent protein (RFP)-expressing RSV A2 (rrRSV) was generated from the full-length RSV plasmid, MP224 by replacing the enhanced green fluorescent protein gene with the wild-type Discosoma RFP gene from pDsRed. For cell maintenance, HEp-2 cells (ATCC no. CCL-23) and Vero cells (ATCC no. CCL-81) were grown in monolayer culture and maintained in DMEM supplemented with 10% fetal calf serum (FCS) and 2 mM L-glutamine in a humidified atmosphere with 5%CO2 at 37 °C. Viral stocks were prepared in HEp-2 cells (ATCC no. CCL-23). Briefly, HEp-2 cells were grown overnight, washed with OptiMEM, and inoculated with rrRSV. After a 2.5-h adsorption period the cells were incubated for 3 days in DMEM supplemented with 1% FCS. Virus was harvested by one freeze-thaw cycle followed by a clarifying centrifugation at 3,500 r.p.m. and stored at − 80 °C. Viral titres were determined by plaque assay in Vero cells using a 2% methyl cellulose overlay, 5% (v/v) formaldehyde fixation, and crystal violet staining (0.015% w/v) at 5 days. For RSV infection of day 170 LBOs, 107 plaque-forming units (PFU) of RSV in 1 ml was directly added onto each Matrigel culture in wells and incubated for 3 h at 37 °C. The RSV inocula were then removed and the cultures were washed with SFD media five times for 5 min and maintained in branching media. The cultures were collected at indicated time points for whole mount staining using anti-RSV (all antigens) antibody (Meridian Life Science, B65890G). Images were taking using inverted microscopes or a two-photon confocal laser scanning microscope Leica TCS SP8 (Leica Microsystems).
Statistics and reproducibility.
Statistical analysis was done using unpaired two-tailed Student’s t-test or one-way ANOVA where appropriate using Prism 7. Results were shown mean ± s.e.m.; P values < 0.05 were considered statistically significant. N-value refers to biologically independent replicates, unless noted otherwise. The investigators were not blinded to allocation during experiments and outcome assessment in animal studies, as no statistics were performed.
Data availability.
The RNA sequencing data sets that support the findings of this study are available from the Sequence Read Archive (SRA). The SRA accession number for day 25 LBOs sequencing is SRP073749 and SRR4295269 for day 170 LBOs. All source data supporting the findings of this study are provided in Supplementary Table 4. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
Additional Information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Lancaster, M. A. & Knoblich, J. A. Organogenesis in a dish: modeling development and disease using organoid technologies. Science 345, 1247125 (2014).
Collins, P. L., Fearns, R. & Graham, B. S. Respiratory syncytial virus: virology, reverse genetics, and pathogenesis of disease. Curr. Top. Microbiol. Immunol. 372, 3–38 (2013).
Johnson, J. E., Gonzales, R. A., Olson, S. J., Wright, P. F. & Graham, B. S. The histopathology of fatal untreated human respiratory syncytial virus infection. Mod. Pathol. 20, 108–119 (2007).
Mulugeta, S., Nureki, S. & Beers, M. F. Lost after translation: insights from pulmonary surfactant for understanding the role of alveolar epithelial dysfunction and cellular quality control in fibrotic lung disease. Am. J. Physiol. Lung Cell. Mol. Physiol. 309, L507–L525 (2015).
Vicary, G. W., Vergne, Y., Santiago-Cornier, A., Young, L. R. & Roman, J. Pulmonary fibrosis in Hermansky–Pudlak syndrome. Ann. Am. Thorac. Soc. 13, 1839–1846 (2016).
Herriges, M. & Morrisey, E. E. Lung development: orchestrating the generation and regeneration of a complex organ. Development 141, 502–513 (2014).
Morrisey, E. E. & Hogan, B. L. Preparing for the first breath: genetic and cellular mechanisms in lung development. Dev. Cell 18, 8–23 (2010).
Dye, B. R. et al. In vitro generation of human pluripotent stem cell derived lung organoids. eLife 4, http://dx.doi.org/10.7554/eLife.05098 (2015).
Dye, B. R. et al. A bioengineered niche promotes in vivo engraftment and maturation of pluripotent stem cell derived human lung organoids. eLife 5, http://dx.doi.org/10.7554/eLife.19732 (2016).
Huang, S. X. et al. Efficient generation of lung and airway epithelial cells from human pluripotent stem cells. Nat. Biotechnol. 32, 84–91 (2014).
Green, M. D. et al. Generation of anterior foregut endoderm from human embryonic and induced pluripotent stem cells. Nat. Biotechnol. 29, 267–272 (2011).
Huang, S. X. et al. The in vitro generation of lung and airway progenitor cells from human pluripotent stem cells. Nat. Protoc. 10, 413–425 (2015).
Kumar, M. E. et al. Mesenchymal cells. Defining a mesenchymal progenitor niche at single-cell resolution. Science 346, 1258810 (2014).
Hrycaj, S. M. et al. Hox5 genes regulate the Wnt2/2b-Bmp4-signaling axis during lung development. Cell Rep. 12, 903–912 (2015).
Liu, L. et al. Hedgehog signaling in neonatal and adult lung. Am. J. Respir. Cell Mol. Biol. 48, 703–710 (2013).
Bellusci, S. et al. Involvement of Sonic hedgehog (Shh) in mouse embryonic lung growth and morphogenesis. Development 124, 53–63 (1997).
Pepicelli, C. V., Lewis, P. M. & McMahon, A. P. Sonic hedgehog regulates branching morphogenesis in the mammalian lung. Curr. Biol. 8, 1083–1086 (1998).
Que, J. et al. Multiple dose-dependent roles for Sox2 in the patterning and differentiation of anterior foregut endoderm. Development 134, 2521–2531 (2007).
Alanis, D. M., Chang, D. R., Akiyama, H., Krasnow, M. A. & Chen, J. Two nested developmental waves demarcate a compartment boundary in the mouse lung. Nat. Commun. 5, 3923 (2014).
Treutlein, B. et al. Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375 (2014).
Sakurai, J. et al. Differential expression of the glycosylated forms of MUC1 during lung development. Eur. J. Histochem. 51, 95–102 (2007).
Cutz, E., Pan, J., Yeger, H., Domnik, N. J. & Fisher, J. T. Recent advances and contraversies on the role of pulmonary neuroepithelial bodies as airway sensors. Semin. Cell Dev. Biol. 24, 40–50 (2013).
Ban, N. et al. ABCA3 as a lipid transporter in pulmonary surfactant biogenesis. J. Biol. Chem. 282, 9628–9634 (2007).
Ciancanelli, M. J. et al. Life-threatening influenza and impaired interferon amplification in human IRF7 deficiency. Science 348, 448–453 (2015).
Whitsett, J. A., Wert, S. E. & Weaver, T. E. Diseases of pulmonary surfactant homeostasis. Annu. Rev. Pathol. 10, 371–393 (2015).
Jain, R. et al. Plasticity of Hopx+ type I alveolar cells to regenerate type II cells in the lung. Nat. Commun. 6, 6727 (2015).
Liu, Y. & Hogan, B. L. Differential gene expression in the distal tip endoderm of the embryonic mouse lung. Expr. Patterns 2, 229–233 (2002).
Rockich, B. E. et al. Sox9 plays multiple roles in the lung epithelium during branching morphogenesis. Proc. Natl Acad. Sci. USA 110, E4456–E4464 (2013).
Perl, A. K., Kist, R., Shan, Z., Scherer, G. & Whitsett, J. A. Normal lung development and function after Sox9 inactivation in the respiratory epithelium. Genesis 41, 23–32 (2005).
Roost, M. S. et al. KeyGenes, a tool to probe tissue differentiation using a human fetal transcriptional atlas. Stem Cell Rep. 4, 1112–1124 (2015).
Florin, T. A., Plint, A. C. & Zorc, J. J. Viral bronchiolitis. Lancet 389, 211–224 (2016).
Simoes, E. A. et al. Challenges and opportunities in developing respiratory syncytial virus therapeutics. J. Infect. Dis. 211 (suppl. 1), S1–S20 (2015).
Liesman, R. M. et al. RSV-encoded NS2 promotes epithelial cell shedding and distal airway obstruction. J. Clin. Invest. 124, 2219–2233 (2014).
Huizing, M., Helip-Wooley, A., Westbroek, W., Gunay-Aygun, M. & Gahl, W. A. Disorders of lysosome-related organelle biogenesis: clinical and molecular genetics. Annu. Rev. Genomics Hum. Genet. 9, 359–386 (2008).
Ryu, J. H. et al. Idiopathic pulmonary fibrosis: evolving concepts. Mayo Clin. Proc. 89, 1130–1142 (2014).
Young, L. R. et al. The alveolar epithelium determines susceptibility to lung fibrosis in Hermansky–Pudlak syndrome. Am. J. Respir. Crit. Care Med. 186, 1014–1024 (2012).
Armanios, M. Telomerase and idiopathic pulmonary fibrosis. Mutat. Res. 730, 52–58 (2012).
Short, K., Hodson, M. & Smyth, I. Spatial mapping and quantification of developmental branching morphogenesis. Development 140, 471–478 (2013).
Peng, T. et al. Coordination of heart and lung co-development by a multipotent cardiopulmonary progenitor. Nature 500, 589–592 (2013).
Chen, Y. W., Ahmed, A. & Snoeck, H. W. Generation of three-dimensional lung bud organoid and its derived branching colonies. Protoc. Exch. http://dx.doi.org/10.1038/protex.2017.027 (2017).
Acknowledgements
This work was supported by grant NIH HL120046-01 (H.-W.S.), 1U01HL134760-01 (H.-W.S.) RO1 AI031971 (A.M.), and RO1 AI114736 (A.M.), as well as a sponsored research and agreement from Northern Biologics Inc. (H.-W.S.), and funding from the Thomas R Kully IPF Research Fund (H.-W.S.). RUES2-HPS1 cells were generated by the Columbia Stem Cell Core Facility. We thank NYULMC OCS Microscopy core C. Petzold and K. Dancel for their assistance with transmission electron microscopy. We thank M. Peeples (Ohio State University) for providing the original recombinant RSV. Flow cytometry was performed in the CCTI Flow Cytometry Core, supported in part by the Office of the Director, National Institutes of Health under awards S10RR027050 and S10OD020056.
Author information
Authors and Affiliations
Contributions
Y.-W.C. designed and performed most experiments, contributed to the concept, and co-wrote the manuscript with H.-W.S. S.X.H. and A.L.R.T.d.C. assisted Y.-W.C. A.F.L. performed transmission electron microscopy. S.-H.H. provided assistance with flow cytometry. J.-L.C. and M.C. provided patient material. L.D.N. and S.V. generated the C12 IRF7-deficient iPS line. M.N.I. and J.B. provided SFTPB-BODIPY. A.M., M.P. and L.M.P. generated and provided virology reagents, and provided design and instruction for experiments involving RSV. H.-W.S. provided concept and guidance, and co-wrote with Y.-W.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of lung bud organoids.
(a)Published 2D directed differentiation protocol for the generation of lung and airway epithelial cells1,2. (b) Schematic overview of the protocol for generating and differentiating LBOs. (c) Unsupervised clustering of RNAseq data generated from EPCAM+ and EPCAM− cells isolated from d25 RUES2 LBOs (3 independent biological replicates). (d) Expression SHH and of its transcriptional targets, GLI1, PTCH and HHIP, of genes expressed in AFE, in lung and airway, and in other AFE-derived lineages in d25LBOs (extracted from the RNAseq data shown in Supplementary Fig. 1c; mean ± s.e.m.,n = 3 independent experiments in RUES2 ESCs).The source data can be found in Supplementary Table 4. (e) ISH for SHH in LBOs at d15, d20 and d25. Representative of 3 independent experiments, RUES2 ESCs. Scale bars 250 μm.
Supplementary Figure 2 Potential of LBOs in vivo.
(a) Staining for human nuclei of RUES2 ESC LBO-derived growths 1.5 months after transplantation under the kidney capsule of NSG mice. Scale bars 500 μm. (b) Staining of LBO-derived growths 5 months after transplantation for murine CD31 (mCD31). Scale bars 50 μm. (c) Staining of LBO-derived growths 5 months after transplantation for SMA and EPCAM. Scale bars 500 μm. (d) Hematoxyline-eosine stain of LBO-derived growths showing ciliated cells 5 months after transplantation under the kidney capsule of NSG mice. Scale bars 25 μm. (e) Hematoxyline-eosine stain of LBO-derived growths showing submucosal glands 5 months after transplantation under the kidney capsule of NSG mice. Scale bars 100 μm. All panels used RUES2 ESCs, representative of 4 independent experiments.
Supplementary Figure 3 Branching in iPSC and ESC-derived LBOs and mesoderm requirement for branching.
(a) Branching colonies in d70 cultures of LBOs derived from RUES2 and three different iPS lines plated at d25 in Matrigel 3D culture in the presence of CHIR99021, BMP4, FGF7, FGF10, and RA. Representative of >10 independent experiments. Scale bar 100 μm. (b) Branching colonies 90 days after plating RUES2 LBOs in Matrigel at 1 (top) or 4 LBOs (bottom) per 6.4 μm well. Scale bars 2.5 μm. All images are representative of >10 independent experiments. (c) Fraction of EPCAM− cells in LBOs (mean ± s.e.m.,n = 3 independent experiments, RUES2 ESCs). The source data can be found in Supplementary Table 4. (d) Colonies from single EPCAM+ and EPCAM− cells isolated from LBOs. Representative of 5 experiments. Scale bar 500 μm. (e) IF of colonies generated from single cells derived from LBOs in Matrigel 3D culture. Representative of 5 experiments. Scale bars 500 μm, 25 μM for SFTPB and SFTPC.
Supplementary Figure 4 LBO maturation in Matrigel at d170.
(a) Morphology of d170 cultures of LBOs derived from three iPS lines plated at d25 in Matrigel in the presence of CHIR99021, BMP4, FGF7, FGF10, and RA. Representative of >10 independent experiments. Scale bar 250 μm. (b) Low-magnification tile scan immunofluorescence images after staining for indicated markers. Staining performed on serial sections of a d170 culture of LBOs derived from C12 iPS line plated at d25 in Matrigel in the presence of CHIR99021, BMP4, FGF7, FGF10, and RA. Representative of 4 independent experiments. Scale bars 1 μm. (c) Electron microscopy of d170 LBOs embedded in Matrigel at d25 in HDF mRNA iPSCs. Arrows indicate LBs. Representative of 3 independent experiments. Scale bar 1 μm. (d) Hematoxylin-Eosin stain (left) and expression of SOX2 and SOX9 in week 14 distal human fetal lung (HFL). Note tubes that co-express SOX2 and SOX9 (arrows). Representative of 3 independent experiments. Scale bar 250 μm. (e) Hierarchical clustering of the genome-wide expression profile in d170 LBOs with genome-wide expression profiles of 2nd trimester human organs and tissues from the KeyGenes database3.
Supplementary Figure 5 Modeling of HPS-associate interstitial pneumonia (HPSIP).
(a) Schematic representation of the HPS1 gene, and location of sequence complementary to the gRNA (upper). Nucleotide sequence of wild type alleles in RUES2 and of both targeted alleles in RUES2-HPS1 cells in the region targeted by the gRNA (middle). Nucleotide and amino acid sequence of exons 15 and 16 of HPS1, showing deletions and premature stop codons in the targeted alleles of RUES2-HPS1 cells (lower). (b) Representative example (of two biological replicates each consisting of three technical replicates) of flow cytometric analysis of EPCAM+ and EPCAM− cells in d50 LBO-derived colonies in 3D Matrigel cultures of RUES2 and RUES2-HPS1 cells. (c) Tile scan images of immunofluorescence staining for EPCAM and PDGFRA of LBO-derived branching colonies in 3D Matrigel cultures generated from parental RUES2 cells and from RUES2-HPS1 cells. Representative of five independent experiments. Scale bars 1 μm. (d) Representative example of the expression of the proliferation antigen Ki67 in EPCAM+ and EPCAM− cells from d40 LBOs derived from parental RUES2 and mutant RUES2-HPS1 cells. Representative of three independent experiments. (e) Fraction of Ki67+ (proliferating) cells in EPCAM+ and EPCAM− cells from LBO in suspension (d15, d25) and in 3D Matrigel cultures (d40) of RUES2 and RUES2-HPS1 cells (mean ± s.e.m., n = 4 for d15 and d25, n = 3 for d40 independent experiments; ∗P < 0.0001 compared to RUES2; two-tailed Student’s t-test). The source data can be found in Supplementary Table 4. (f) Hydroxyproline content in LBO-derived colonies in 3D Matrigel cultures of RUES2 and RUES2-HPS1 cells (mean ± s.e.m.,n = 3 independent experiments; ∗P < 0.05; two-tailed Student’s t-test). The source data can be found in Supplementary Table 4. (g) Quantification of collagens I and III and fibronectin in 3D Matrigel cultures of RUES2 and RUES2-HPS1 cells using immunofluorescence intensity relative to DAPI (mean ± s.e.m., n = 3 independent experiments; ∗P < 0.05 for fibronectin, ∗P < 0.01 for collagens; two-tailed Student’s t-test after normalizing RUES2 controls to 1 in each experiment). The source data can be found in Supplementary Table 4. (h–j) Mixing experiments suggest that epithelial cells drive the accumulation of mesenchymal cells. It is still debated to what extent the pathogenetic origin IPF is epithelial, mesenchymal or both4,5,6, although most evidence points towards a prime role for epithelial injury, in particular ATII injury7. We devised a mixing strategy to address this issue without disrupting the LBO structures. When definitive endoderm cells derived from ZsGreen- or mCherry-expressing RUES2 cells were mixed at d4 of the protocol (see Supplementary Fig. 1b), after dissociation of embryoid bodies and before replating in 2D to generate anterior foregut cells, the resulting LBOs contained homogenously distributed red and green cells (h, left panel). However, when red and green LBOs were first generated separately and then grown together in suspension from d10 on, fusion between LBOs occurred such that large sections were either green or red (h, right panel). When ZsGreen+ RUES2 cells were mixed with non-fluorescent mutant RUES2-HPS1 cells at d4, the fraction of both wild type and mutant mesenchymal cells was increased (i, blue bars). As both genotypes were equally affected, these data indicate that a primary mesenchymal cause is unlikely. When LBOs were fused at d10, only sections of the resulting branching colonies were ZsGreen+ after plating in Matrigel, as expected (j). In these cultures, only accumulation of mutant mesenchymal cells could be demonstrated (i, orange bars). These data are consistent with the notion that fibrosis in HPS1 is driven by mutant epithelial cells7,8, and suggest that the interaction between mutant epithelial cells and mesenchymal cells occurs over a short range. Mean ± s.e.m., n = 3 independent experiments; ∗P < 0.01 compared to parental RUES2 and to parental RUES2 mixed at d10 with RUES2-HPS1; one way ANOVA. The source data can be found in Supplementary Table 4.
Supplementary information
Supplementary Information
Supplementary Information (PDF 127790 kb)
Supplementary Table 1
Supplementary Information (XLSX 12 kb)
Supplementary Table 2
Supplementary Information (XLSX 9 kb)
Supplementary Table 3
Supplementary Information (XLSX 12 kb)
Supplementary Table 4
Supplementary Information (XLSX 633 kb)
Beating cilia in the LBO-derived growth.
Bright field movie showing beating cilia in LBO-derived growth (RUES2) 5 months after engraftment. Section was made using a vibratome. Scale bar 75 μm. (MOV 29282 kb)
Morphology of d170 LBOs.
Bright field movie showing the connected dilated saccules and structures resembling pulmonary acini in C12 LBOs. Scale bar 100 μm. (MOV 46931 kb)
Uptake of SPB-BODIPY in d170 LBOs.
Time lapse movie of several distal buds of different d170 LBOs (RUES2) taken at 2 min intervals. The movie shows uptake of SPB-BODIPY over time in the cells and secretion into the lumen. Green: SPB-BODIPY. Scale bar 50 μm. (MOV 18049 kb)
Modeling respiratory syncytial virus infection in d170 LBOs.
Confocal microscopy movie of a distal bud of a d170 LBO (RUES2) showing infected cells in the lumen (RSV antigen, green; DAPI, blue). Scale bar 50 μm. (MOV 2960 kb)
Rights and permissions
About this article
Cite this article
Chen, YW., Huang, S., de Carvalho, A. et al. A three-dimensional model of human lung development and disease from pluripotent stem cells. Nat Cell Biol 19, 542–549 (2017). https://doi.org/10.1038/ncb3510
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3510
This article is cited by
-
Lung Organoids: Systematic Review of Recent Advancements and its Future Perspectives
Tissue Engineering and Regenerative Medicine (2024)
-
Insights on Three Dimensional Organoid Studies for Stem Cell Therapy in Regenerative Medicine
Stem Cell Reviews and Reports (2024)
-
Alternatives of Animal Models for Biomedical Research: a Comprehensive Review of Modern Approaches
Stem Cell Reviews and Reports (2024)
-
MSCs alleviate LPS-induced acute lung injury by inhibiting the proinflammatory function of macrophages in mouse lung organoid–macrophage model
Cellular and Molecular Life Sciences (2024)
-
Innovative explorations: unveiling the potential of organoids for investigating environmental pollutant exposure
Environmental Science and Pollution Research (2024)